Yaron Meirovitch
X-Ray2EM: Uncertainty-Aware Cross-Modality Image Reconstruction from X-Ray to Electron Microscopy in Connectomics
Mar 02, 2023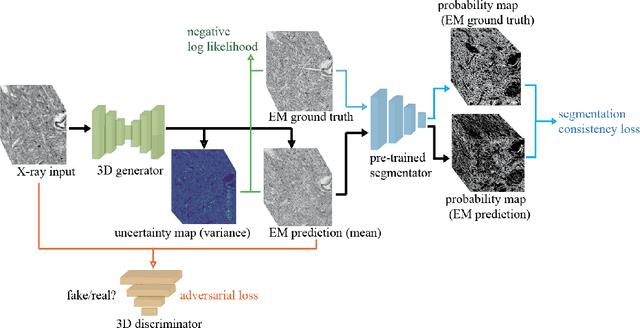
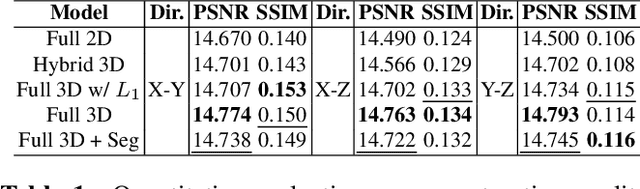
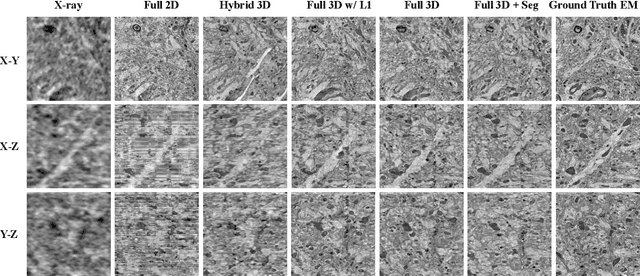
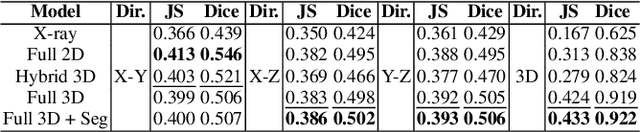
Abstract:Comprehensive, synapse-resolution imaging of the brain will be crucial for understanding neuronal computations and function. In connectomics, this has been the sole purview of volume electron microscopy (EM), which entails an excruciatingly difficult process because it requires cutting tissue into many thin, fragile slices that then need to be imaged, aligned, and reconstructed. Unlike EM, hard X-ray imaging is compatible with thick tissues, eliminating the need for thin sectioning, and delivering fast acquisition, intrinsic alignment, and isotropic resolution. Unfortunately, current state-of-the-art X-ray microscopy provides much lower resolution, to the extent that segmenting membranes is very challenging. We propose an uncertainty-aware 3D reconstruction model that translates X-ray images to EM-like images with enhanced membrane segmentation quality, showing its potential for developing simpler, faster, and more accurate X-ray based connectomics pipelines.
Learning Guided Electron Microscopy with Active Acquisition
Jan 07, 2021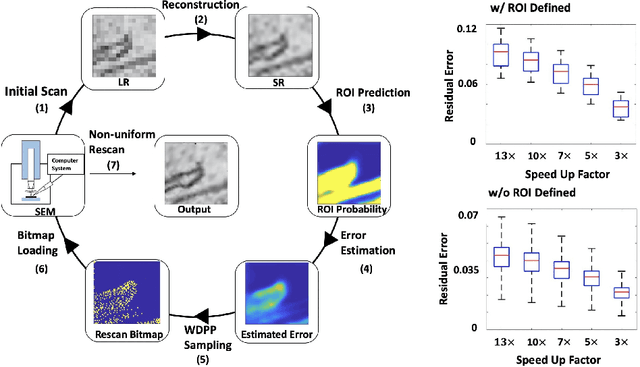
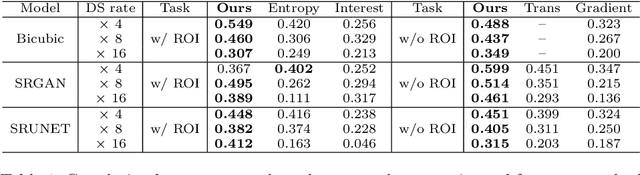
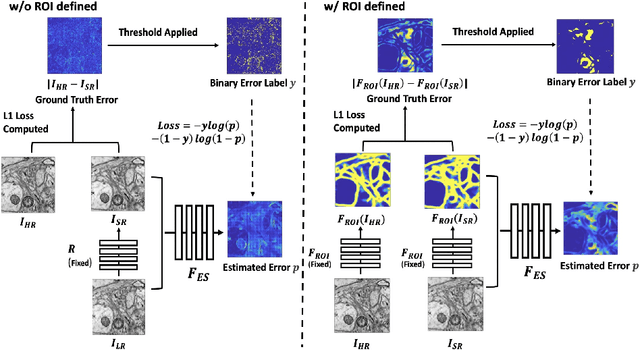
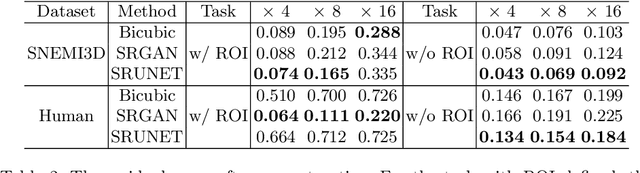
Abstract:Single-beam scanning electron microscopes (SEM) are widely used to acquire massive data sets for biomedical study, material analysis, and fabrication inspection. Datasets are typically acquired with uniform acquisition: applying the electron beam with the same power and duration to all image pixels, even if there is great variety in the pixels' importance for eventual use. Many SEMs are now able to move the beam to any pixel in the field of view without delay, enabling them, in principle, to invest their time budget more effectively with non-uniform imaging. In this paper, we show how to use deep learning to accelerate and optimize single-beam SEM acquisition of images. Our algorithm rapidly collects an information-lossy image (e.g. low resolution) and then applies a novel learning method to identify a small subset of pixels to be collected at higher resolution based on a trade-off between the saliency and spatial diversity. We demonstrate the efficacy of this novel technique for active acquisition by speeding up the task of collecting connectomic datasets for neurobiology by up to an order of magnitude.
Cross-Classification Clustering: An Efficient Multi-Object Tracking Technique for 3-D Instance Segmentation in Connectomics
Dec 04, 2018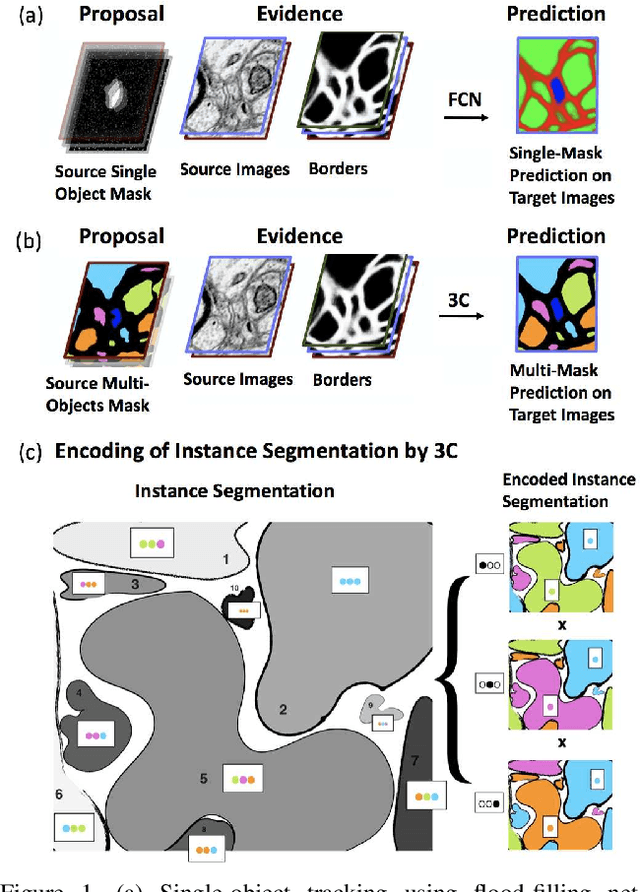
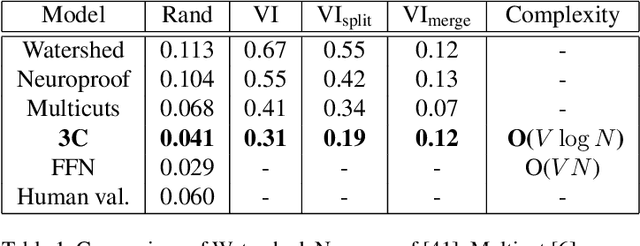
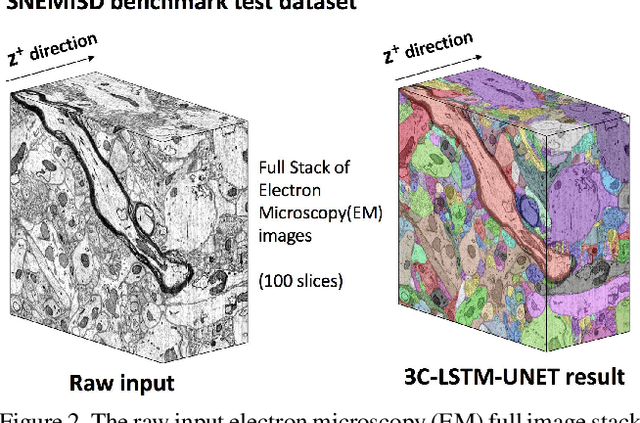
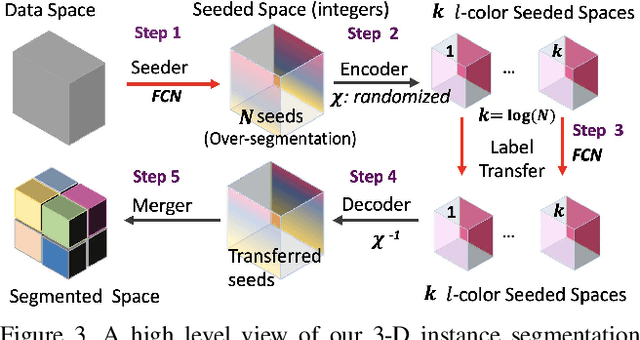
Abstract:Pixel-accurate tracking of objects is a key element in many computer vision applications, often solved by iterated individual object tracking or instance segmentation followed by object matching. Here we introduce cross-classification clustering (3C), a new technique that simultaneously tracks all objects in an image stack. The key idea in cross-classification is to efficiently turn a clustering problem into a classification problem by running a logarithmic number of independent classifications, letting the cross-labeling of these classifications uniquely classify each pixel to the object labels. We apply the 3C mechanism to achieve state-of-the-art accuracy in connectomics - nanoscale mapping of the brain from electron microscopy volumes. Our reconstruction system introduces an order of magnitude scalability improvement over the best current methods for neuronal reconstruction, and can be seamlessly integrated within existing single-object tracking methods like Google's flood-filling networks to improve their performance. This scalability is crucial for the real-world deployment of connectomics pipelines, as the best performing existing techniques require computing infrastructures that are beyond the reach of most labs. We believe 3C has valuable scalability implications in other domains that require pixel-accurate tracking of multiple objects in image stacks or video.
Morphological Error Detection in 3D Segmentations
May 30, 2017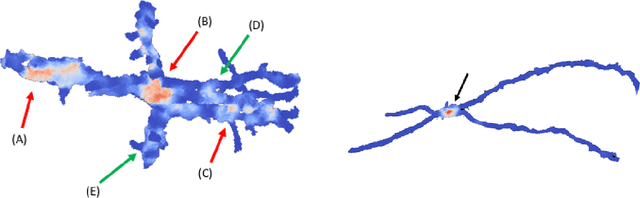
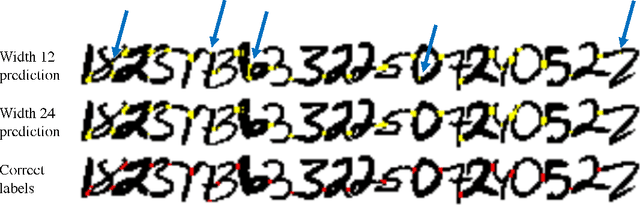
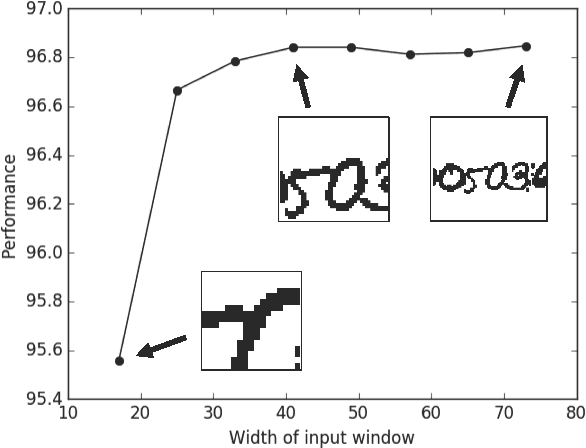
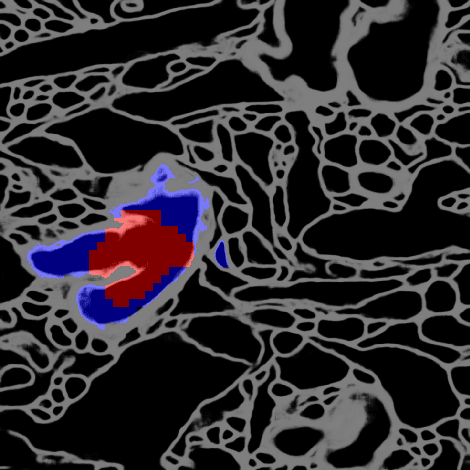
Abstract:Deep learning algorithms for connectomics rely upon localized classification, rather than overall morphology. This leads to a high incidence of erroneously merged objects. Humans, by contrast, can easily detect such errors by acquiring intuition for the correct morphology of objects. Biological neurons have complicated and variable shapes, which are challenging to learn, and merge errors take a multitude of different forms. We present an algorithm, MergeNet, that shows 3D ConvNets can, in fact, detect merge errors from high-level neuronal morphology. MergeNet follows unsupervised training and operates across datasets. We demonstrate the performance of MergeNet both on a variety of connectomics data and on a dataset created from merged MNIST images.
Toward Streaming Synapse Detection with Compositional ConvNets
Feb 23, 2017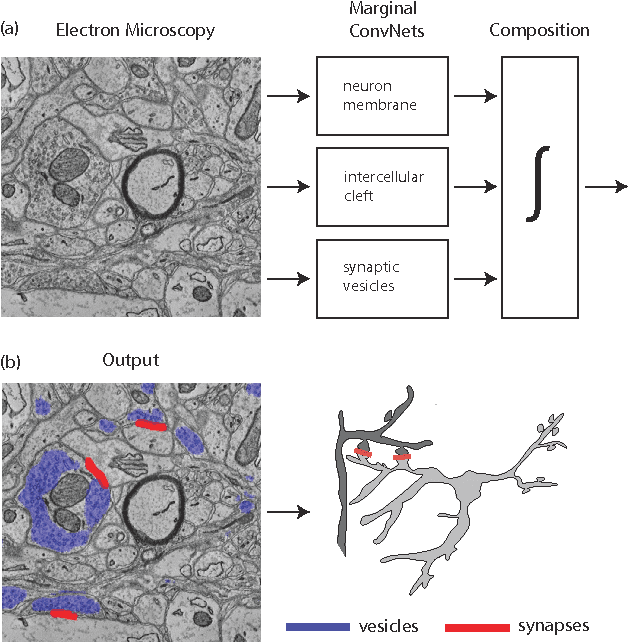
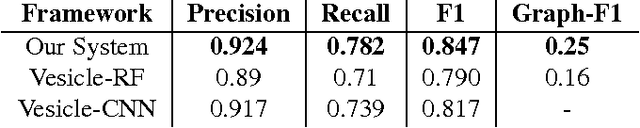
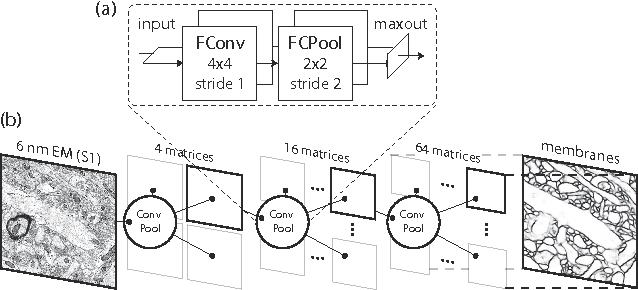
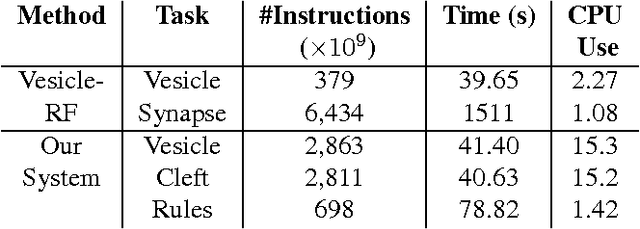
Abstract:Connectomics is an emerging field in neuroscience that aims to reconstruct the 3-dimensional morphology of neurons from electron microscopy (EM) images. Recent studies have successfully demonstrated the use of convolutional neural networks (ConvNets) for segmenting cell membranes to individuate neurons. However, there has been comparatively little success in high-throughput identification of the intercellular synaptic connections required for deriving connectivity graphs. In this study, we take a compositional approach to segmenting synapses, modeling them explicitly as an intercellular cleft co-located with an asymmetric vesicle density along a cell membrane. Instead of requiring a deep network to learn all natural combinations of this compositionality, we train lighter networks to model the simpler marginal distributions of membranes, clefts and vesicles from just 100 electron microscopy samples. These feature maps are then combined with simple rules-based heuristics derived from prior biological knowledge. Our approach to synapse detection is both more accurate than previous state-of-the-art (7% higher recall and 5% higher F1-score) and yields a 20-fold speed-up compared to the previous fastest implementations. We demonstrate by reconstructing the first complete, directed connectome from the largest available anisotropic microscopy dataset (245 GB) of mouse somatosensory cortex (S1) in just 9.7 hours on a single shared-memory CPU system. We believe that this work marks an important step toward the goal of a microscope-pace streaming connectomics pipeline.
A Multi-Pass Approach to Large-Scale Connectomics
Dec 07, 2016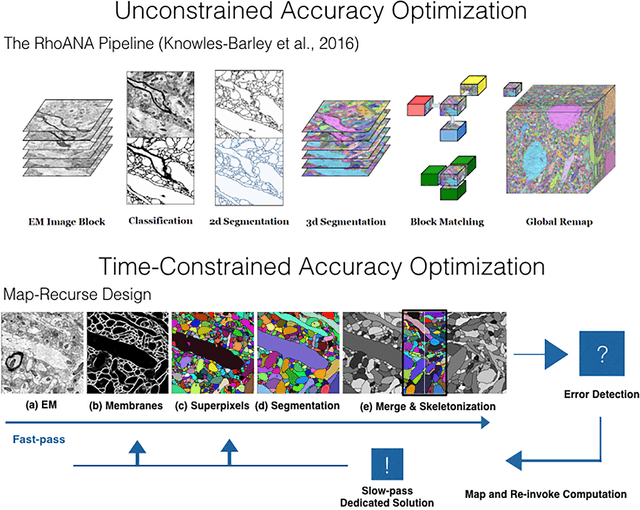
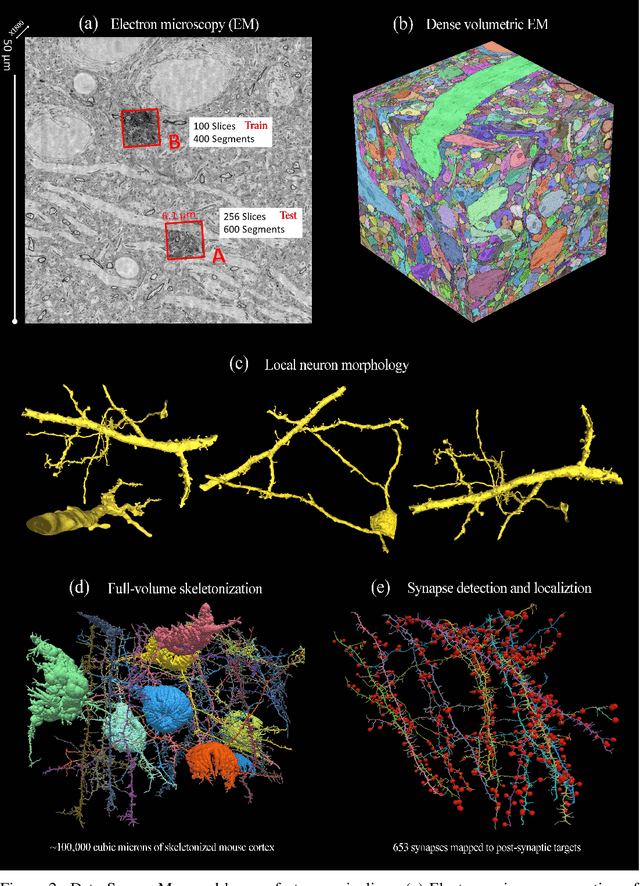
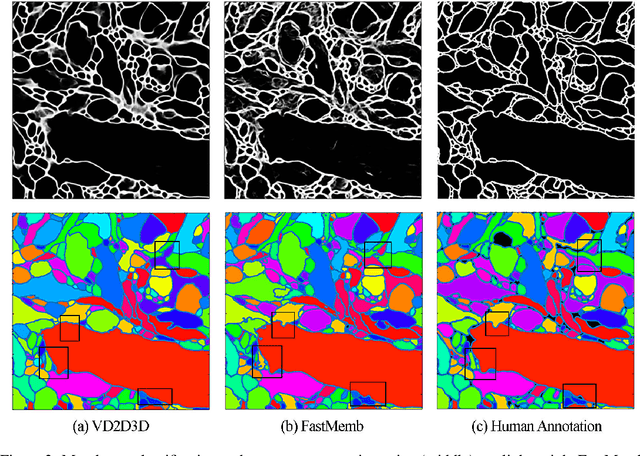
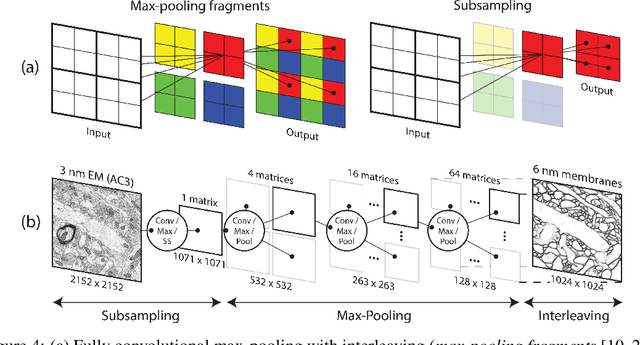
Abstract:The field of connectomics faces unprecedented "big data" challenges. To reconstruct neuronal connectivity, automated pixel-level segmentation is required for petabytes of streaming electron microscopy data. Existing algorithms provide relatively good accuracy but are unacceptably slow, and would require years to extract connectivity graphs from even a single cubic millimeter of neural tissue. Here we present a viable real-time solution, a multi-pass pipeline optimized for shared-memory multicore systems, capable of processing data at near the terabyte-per-hour pace of multi-beam electron microscopes. The pipeline makes an initial fast-pass over the data, and then makes a second slow-pass to iteratively correct errors in the output of the fast-pass. We demonstrate the accuracy of a sparse slow-pass reconstruction algorithm and suggest new methods for detecting morphological errors. Our fast-pass approach provided many algorithmic challenges, including the design and implementation of novel shallow convolutional neural nets and the parallelization of watershed and object-merging techniques. We use it to reconstruct, from image stack to skeletons, the full dataset of Kasthuri et al. (463 GB capturing 120,000 cubic microns) in a matter of hours on a single multicore machine rather than the weeks it has taken in the past on much larger distributed systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge