Seymour Knowles-Barley
A Multi-Pass Approach to Large-Scale Connectomics
Dec 07, 2016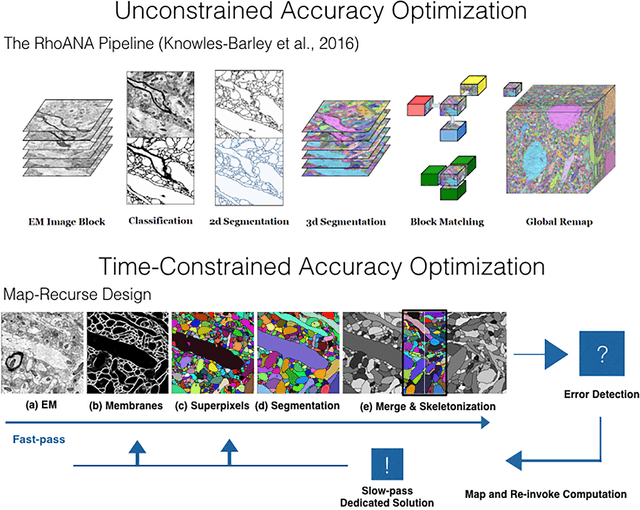
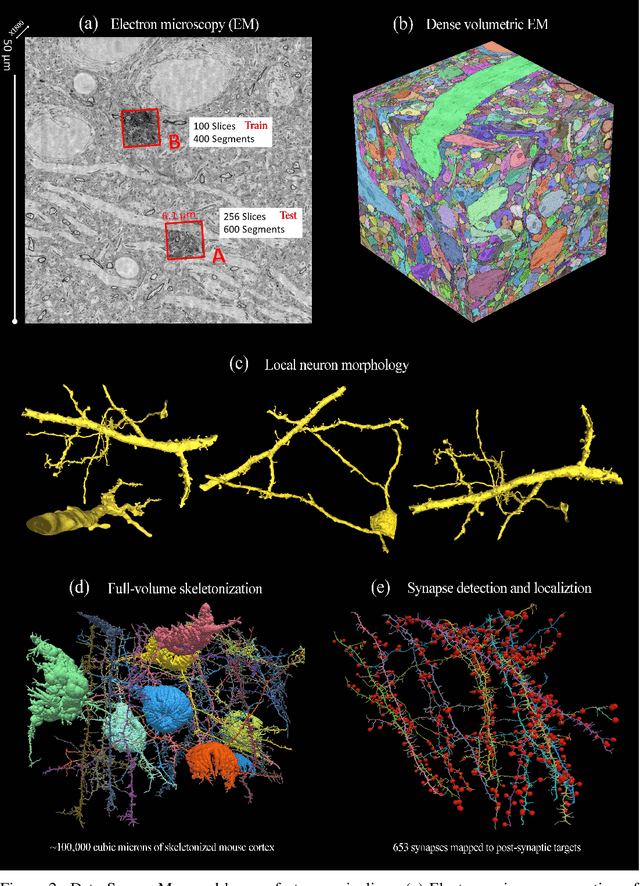
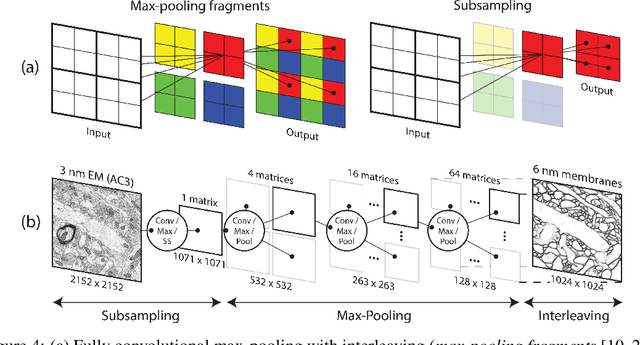
Abstract:The field of connectomics faces unprecedented "big data" challenges. To reconstruct neuronal connectivity, automated pixel-level segmentation is required for petabytes of streaming electron microscopy data. Existing algorithms provide relatively good accuracy but are unacceptably slow, and would require years to extract connectivity graphs from even a single cubic millimeter of neural tissue. Here we present a viable real-time solution, a multi-pass pipeline optimized for shared-memory multicore systems, capable of processing data at near the terabyte-per-hour pace of multi-beam electron microscopes. The pipeline makes an initial fast-pass over the data, and then makes a second slow-pass to iteratively correct errors in the output of the fast-pass. We demonstrate the accuracy of a sparse slow-pass reconstruction algorithm and suggest new methods for detecting morphological errors. Our fast-pass approach provided many algorithmic challenges, including the design and implementation of novel shallow convolutional neural nets and the parallelization of watershed and object-merging techniques. We use it to reconstruct, from image stack to skeletons, the full dataset of Kasthuri et al. (463 GB capturing 120,000 cubic microns) in a matter of hours on a single multicore machine rather than the weeks it has taken in the past on much larger distributed systems.
RhoanaNet Pipeline: Dense Automatic Neural Annotation
Nov 21, 2016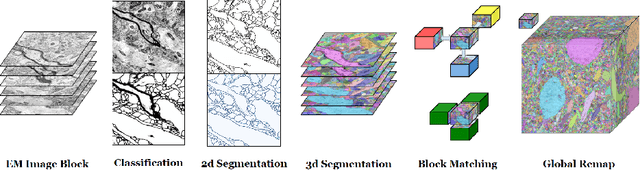
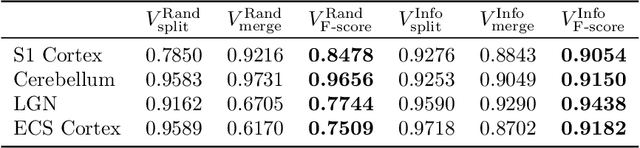
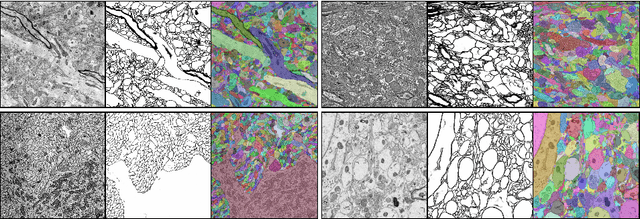
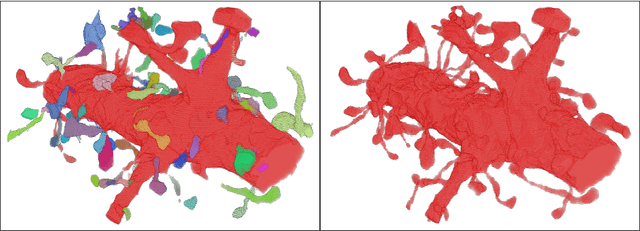
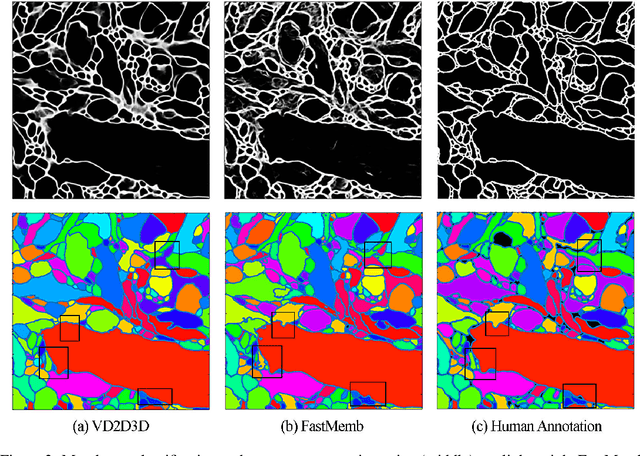
Abstract:Reconstructing a synaptic wiring diagram, or connectome, from electron microscopy (EM) images of brain tissue currently requires many hours of manual annotation or proofreading (Kasthuri and Lichtman, 2010; Lichtman and Sanes, 2008; Seung, 2009). The desire to reconstruct ever larger and more complex networks has pushed the collection of ever larger EM datasets. A cubic millimeter of raw imaging data would take up 1 PB of storage and present an annotation project that would be impractical without relying heavily on automatic segmentation methods. The RhoanaNet image processing pipeline was developed to automatically segment large volumes of EM data and ease the burden of manual proofreading and annotation. Based on (Kaynig et al., 2015), we updated every stage of the software pipeline to provide better throughput performance and higher quality segmentation results. We used state of the art deep learning techniques to generate improved membrane probability maps, and Gala (Nunez-Iglesias et al., 2014) was used to agglomerate 2D segments into 3D objects. We applied the RhoanaNet pipeline to four densely annotated EM datasets, two from mouse cortex, one from cerebellum and one from mouse lateral geniculate nucleus (LGN). All training and test data is made available for benchmark comparisons. The best segmentation results obtained gave $V^\text{Info}_\text{F-score}$ scores of 0.9054 and 09182 for the cortex datasets, 0.9438 for LGN, and 0.9150 for Cerebellum. The RhoanaNet pipeline is open source software. All source code, training data, test data, and annotations for all four benchmark datasets are available at www.rhoana.org.
Large-Scale Automatic Reconstruction of Neuronal Processes from Electron Microscopy Images
Mar 28, 2013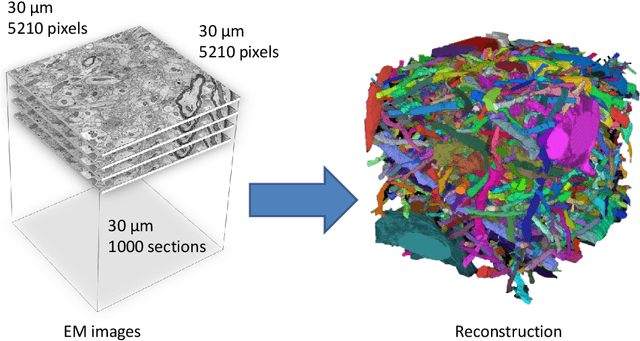
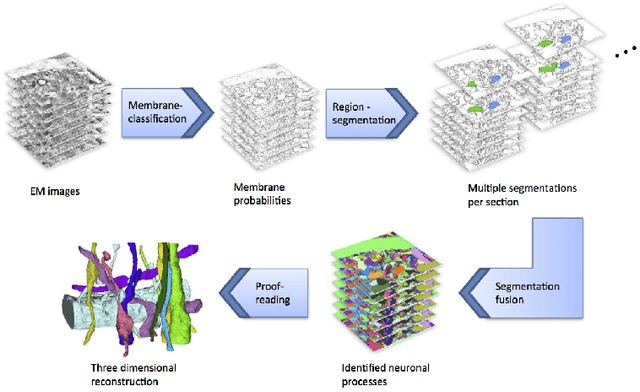
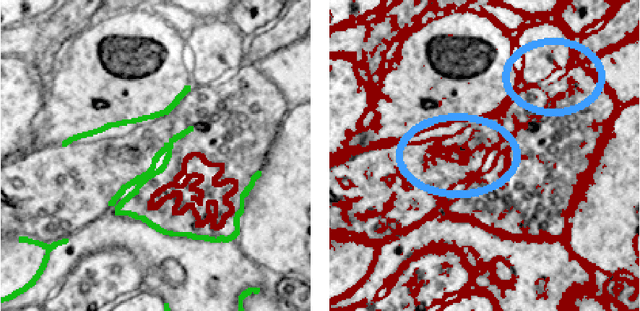
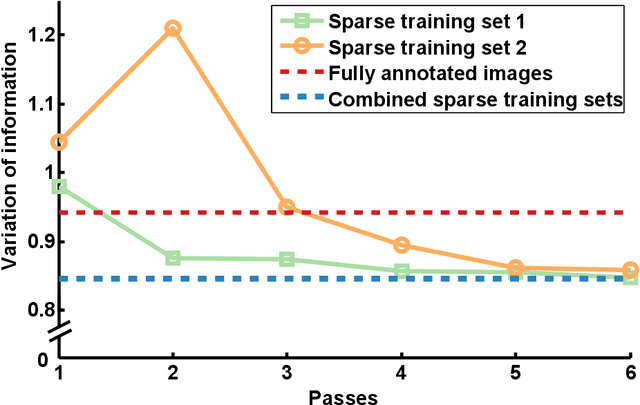
Abstract:Automated sample preparation and electron microscopy enables acquisition of very large image data sets. These technical advances are of special importance to the field of neuroanatomy, as 3D reconstructions of neuronal processes at the nm scale can provide new insight into the fine grained structure of the brain. Segmentation of large-scale electron microscopy data is the main bottleneck in the analysis of these data sets. In this paper we present a pipeline that provides state-of-the art reconstruction performance while scaling to data sets in the GB-TB range. First, we train a random forest classifier on interactive sparse user annotations. The classifier output is combined with an anisotropic smoothing prior in a Conditional Random Field framework to generate multiple segmentation hypotheses per image. These segmentations are then combined into geometrically consistent 3D objects by segmentation fusion. We provide qualitative and quantitative evaluation of the automatic segmentation and demonstrate large-scale 3D reconstructions of neuronal processes from a $\mathbf{27,000}$ $\mathbf{\mu m^3}$ volume of brain tissue over a cube of $\mathbf{30 \; \mu m}$ in each dimension corresponding to 1000 consecutive image sections. We also introduce Mojo, a proofreading tool including semi-automated correction of merge errors based on sparse user scribbles.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge