Daniel Berger
Transported Memory Networks accelerating Computational Fluid Dynamics
Feb 25, 2025


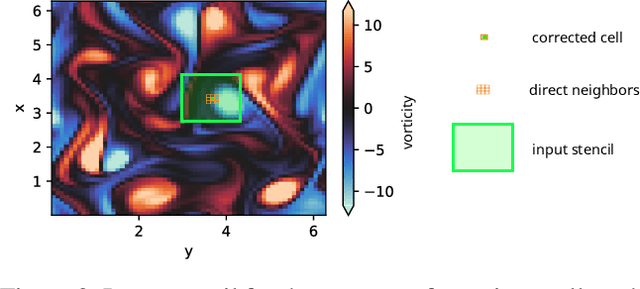
Abstract:In recent years, augmentation of differentiable PDE solvers with neural networks has shown promising results, particularly in fluid simulations. However, most approaches rely on convolutional neural networks and custom solvers operating on Cartesian grids with efficient access to cell data. This particular choice poses challenges for industrial-grade solvers that operate on unstructured meshes, where access is restricted to neighboring cells only. In this work, we address this limitation using a novel architecture, named Transported Memory Networks. The architecture draws inspiration from both traditional turbulence models and recurrent neural networks, and it is fully compatible with generic discretizations. Our results show that it is point-wise and statistically comparable to, or improves upon, previous methods in terms of both accuracy and computational efficiency.
FreSeg: Frenet-Frame-based Part Segmentation for 3D Curvilinear Structures
Apr 19, 2024



Abstract:Part segmentation is a crucial task for 3D curvilinear structures like neuron dendrites and blood vessels, enabling the analysis of dendritic spines and aneurysms with scientific and clinical significance. However, their diversely winded morphology poses a generalization challenge to existing deep learning methods, which leads to labor-intensive manual correction. In this work, we propose FreSeg, a framework of part segmentation tasks for 3D curvilinear structures. With Frenet-Frame-based point cloud transformation, it enables the models to learn more generalizable features and have significant performance improvements on tasks involving elongated and curvy geometries. We evaluate FreSeg on 2 datasets: 1) DenSpineEM, an in-house dataset for dendritic spine segmentation, and 2) IntrA, a public 3D dataset for intracranial aneurysm segmentation. Further, we will release the DenSpineEM dataset, which includes roughly 6,000 spines from 69 dendrites from 3 public electron microscopy (EM) datasets, to foster the development of effective dendritic spine instance extraction methods and, consequently, large-scale connectivity analysis to better understand mammalian brains.
Detecting Synapse Location and Connectivity by Signed Proximity Estimation and Pruning with Deep Nets
Oct 25, 2018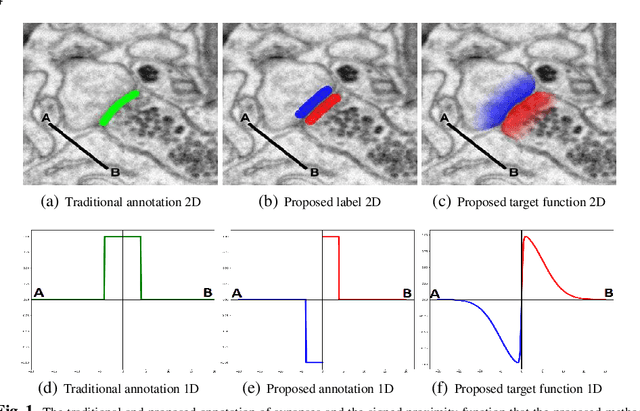



Abstract:Synaptic connectivity detection is a critical task for neural reconstruction from Electron Microscopy (EM) data. Most of the existing algorithms for synapse detection do not identify the cleft location and direction of connectivity simultaneously. The few methods that computes direction along with contact location have only been demonstrated to work on either dyadic (most common in vertebrate brain) or polyadic (found in fruit fly brain) synapses, but not on both types. In this paper, we present an algorithm to automatically predict the location as well as the direction of both dyadic and polyadic synapses. The proposed algorithm first generates candidate synaptic connections from voxelwise predictions of signed proximity generated by a 3D U-net. A second 3D CNN then prunes the set of candidates to produce the final detection of cleft and connectivity orientation. Experimental results demonstrate that the proposed method outperforms the existing methods for determining synapses in both rodent and fruit fly brain.
RhoanaNet Pipeline: Dense Automatic Neural Annotation
Nov 21, 2016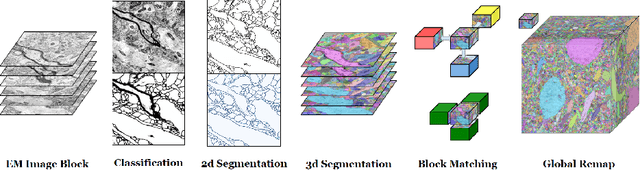
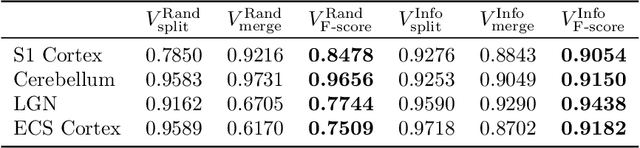
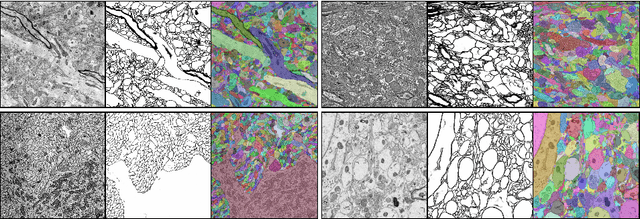
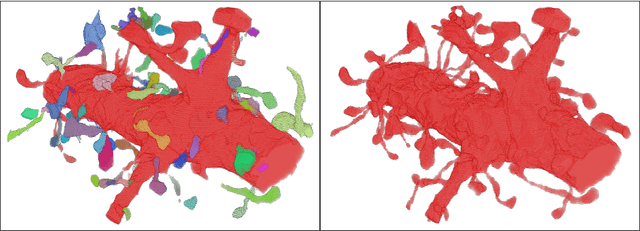
Abstract:Reconstructing a synaptic wiring diagram, or connectome, from electron microscopy (EM) images of brain tissue currently requires many hours of manual annotation or proofreading (Kasthuri and Lichtman, 2010; Lichtman and Sanes, 2008; Seung, 2009). The desire to reconstruct ever larger and more complex networks has pushed the collection of ever larger EM datasets. A cubic millimeter of raw imaging data would take up 1 PB of storage and present an annotation project that would be impractical without relying heavily on automatic segmentation methods. The RhoanaNet image processing pipeline was developed to automatically segment large volumes of EM data and ease the burden of manual proofreading and annotation. Based on (Kaynig et al., 2015), we updated every stage of the software pipeline to provide better throughput performance and higher quality segmentation results. We used state of the art deep learning techniques to generate improved membrane probability maps, and Gala (Nunez-Iglesias et al., 2014) was used to agglomerate 2D segments into 3D objects. We applied the RhoanaNet pipeline to four densely annotated EM datasets, two from mouse cortex, one from cerebellum and one from mouse lateral geniculate nucleus (LGN). All training and test data is made available for benchmark comparisons. The best segmentation results obtained gave $V^\text{Info}_\text{F-score}$ scores of 0.9054 and 09182 for the cortex datasets, 0.9438 for LGN, and 0.9150 for Cerebellum. The RhoanaNet pipeline is open source software. All source code, training data, test data, and annotations for all four benchmark datasets are available at www.rhoana.org.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge