Mikhail Bessmeltsev
SketchingReality: From Freehand Scene Sketches To Photorealistic Images
Feb 16, 2026Abstract:Recent years have witnessed remarkable progress in generative AI, with natural language emerging as the most common conditioning input. As underlying models grow more powerful, researchers are exploring increasingly diverse conditioning signals, such as depth maps, edge maps, camera parameters, and reference images, to give users finer control over generation. Among different modalities, sketches are a natural and long-standing form of human communication, enabling rapid expression of visual concepts. Previous literature has largely focused on edge maps, often misnamed 'sketches', yet algorithms that effectively handle true freehand sketches, with their inherent abstraction and distortions, remain underexplored. We pursue the challenging goal of balancing photorealism with sketch adherence when generating images from freehand input. A key obstacle is the absence of ground-truth, pixel-aligned images: by their nature, freehand sketches do not have a single correct alignment. To address this, we propose a modulation-based approach that prioritizes semantic interpretation of the sketch over strict adherence to individual edge positions. We further introduce a novel loss that enables training on freehand sketches without requiring ground-truth pixel-aligned images. We show that our method outperforms existing approaches in both semantic alignment with freehand sketch inputs and in the realism and overall quality of the generated images.
FreSeg: Frenet-Frame-based Part Segmentation for 3D Curvilinear Structures
Apr 19, 2024



Abstract:Part segmentation is a crucial task for 3D curvilinear structures like neuron dendrites and blood vessels, enabling the analysis of dendritic spines and aneurysms with scientific and clinical significance. However, their diversely winded morphology poses a generalization challenge to existing deep learning methods, which leads to labor-intensive manual correction. In this work, we propose FreSeg, a framework of part segmentation tasks for 3D curvilinear structures. With Frenet-Frame-based point cloud transformation, it enables the models to learn more generalizable features and have significant performance improvements on tasks involving elongated and curvy geometries. We evaluate FreSeg on 2 datasets: 1) DenSpineEM, an in-house dataset for dendritic spine segmentation, and 2) IntrA, a public 3D dataset for intracranial aneurysm segmentation. Further, we will release the DenSpineEM dataset, which includes roughly 6,000 spines from 69 dendrites from 3 public electron microscopy (EM) datasets, to foster the development of effective dendritic spine instance extraction methods and, consequently, large-scale connectivity analysis to better understand mammalian brains.
Volumetric Parameterization of the Placenta to a Flattened Template
Nov 15, 2021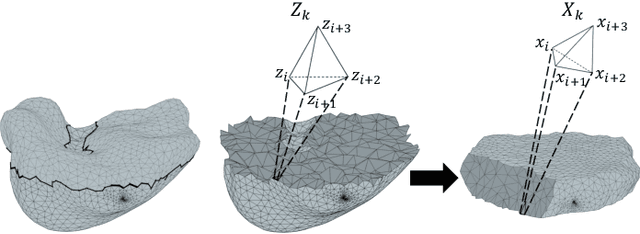
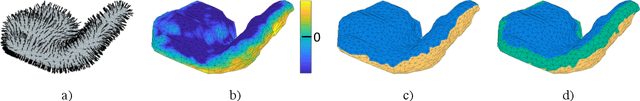
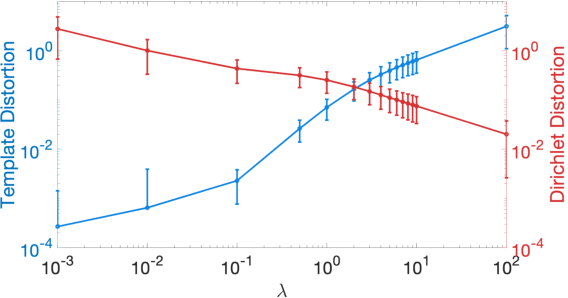
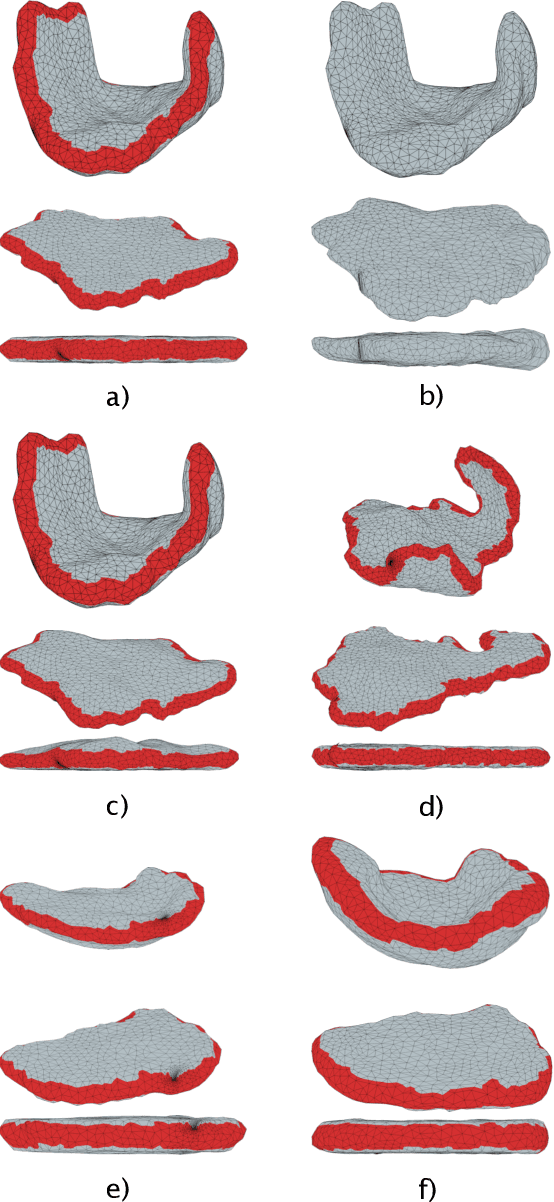
Abstract:We present a volumetric mesh-based algorithm for parameterizing the placenta to a flattened template to enable effective visualization of local anatomy and function. MRI shows potential as a research tool as it provides signals directly related to placental function. However, due to the curved and highly variable in vivo shape of the placenta, interpreting and visualizing these images is difficult. We address interpretation challenges by mapping the placenta so that it resembles the familiar ex vivo shape. We formulate the parameterization as an optimization problem for mapping the placental shape represented by a volumetric mesh to a flattened template. We employ the symmetric Dirichlet energy to control local distortion throughout the volume. Local injectivity in the mapping is enforced by a constrained line search during the gradient descent optimization. We validate our method using a research study of 111 placental shapes extracted from BOLD MRI images. Our mapping achieves sub-voxel accuracy in matching the template while maintaining low distortion throughout the volume. We demonstrate how the resulting flattening of the placenta improves visualization of anatomy and function. Our code is freely available at https://github.com/mabulnaga/placenta-flattening .
Placental Flattening via Volumetric Parameterization
Mar 14, 2019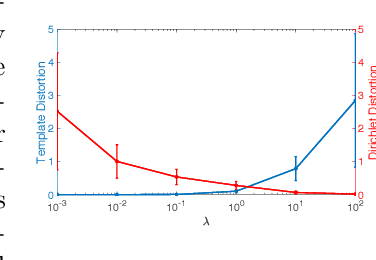
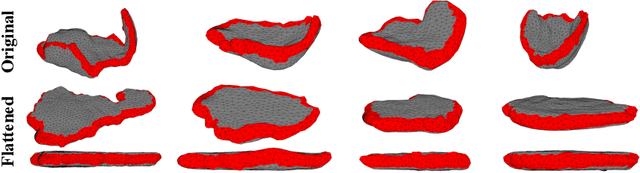
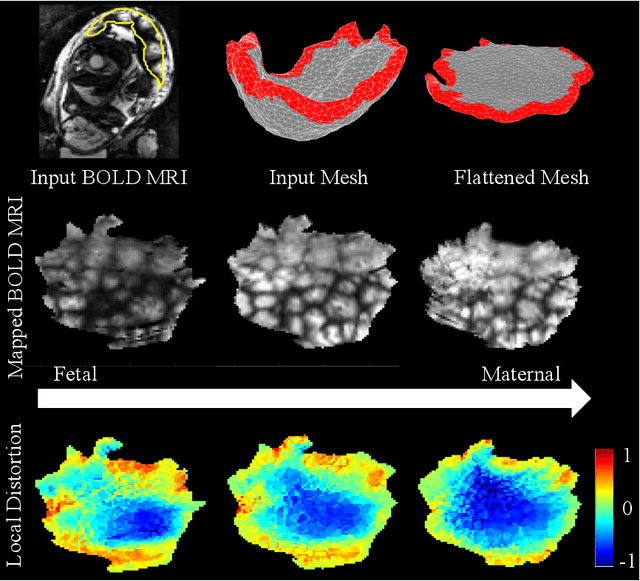

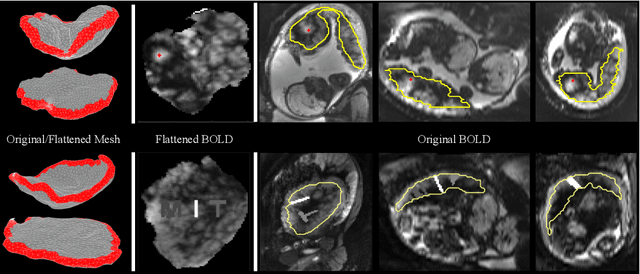
Abstract:We present a volumetric mesh-based algorithm for flattening the placenta to a canonical template to enable effective visualization of local anatomy and function. Monitoring placental function in vivo promises to support pregnancy assessment and to improve care outcomes. We aim to alleviate visualization and interpretation challenges presented by the shape of the placenta when it is attached to the curved uterine wall. We flatten the volumetric mesh that captures placental shape to resemble the well-studied ex vivo shape. We formulate our method as a map from the in vivo shape to a flattened template that minimizes the symmetric Dirichlet energy density to control distortion throughout the volume. Local injectivity is enforced via constrained line search during gradient descent. We evaluate the proposed method on 28 placenta shapes extracted from MRI images in a study of placental function. We achieve sub-voxel accuracy in mapping the boundary of the placenta to the template while successfully controlling distortion throughout the volume. We illustrate how the resulting mapping of the placenta enhances visualization of the placental anatomy and function. Our code is freely available at https://github.com/mabulnaga/placenta-flattening .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge