William Parker
Segmentation of Pulmonary Opacification in Chest CT Scans of COVID-19 Patients
Jul 08, 2020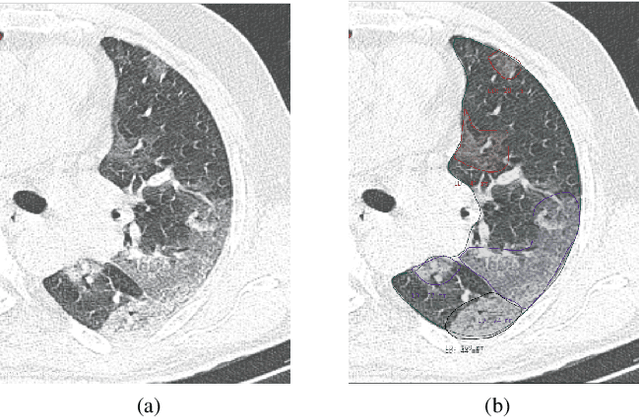
Abstract:The Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) has rapidly spread into a global pandemic. A form of pneumonia, presenting as opacities with in a patient's lungs, is the most common presentation associated with this virus, and great attention has gone into how these changes relate to patient morbidity and mortality. In this work we provide open source models for the segmentation of patterns of pulmonary opacification on chest Computed Tomography (CT) scans which have been correlated with various stages and severities of infection. We have collected 663 chest CT scans of COVID-19 patients from healthcare centers around the world, and created pixel wise segmentation labels for nearly 25,000 slices that segment 6 different patterns of pulmonary opacification. We provide open source implementations and pre-trained weights for multiple segmentation models trained on our dataset. Our best model achieves an opacity Intersection-Over-Union score of 0.76 on our test set, demonstrates successful domain adaptation, and predicts the volume of opacification within 1.7\% of expert radiologists. Additionally, we present an analysis of the inter-observer variability inherent to this task, and propose methods for appropriate probabilistic approaches.
A Weakly Supervised Region-Based Active Learning Method for COVID-19 Segmentation in CT Images
Jul 07, 2020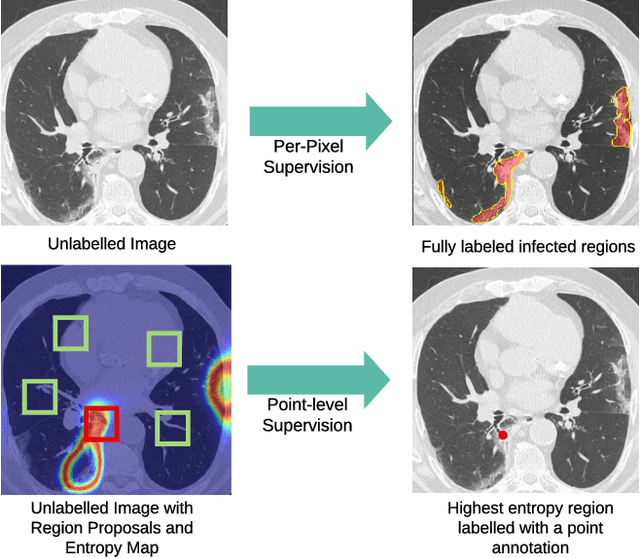
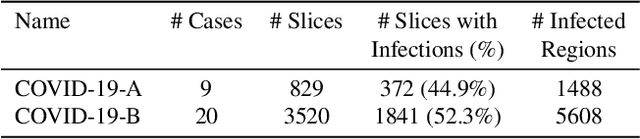
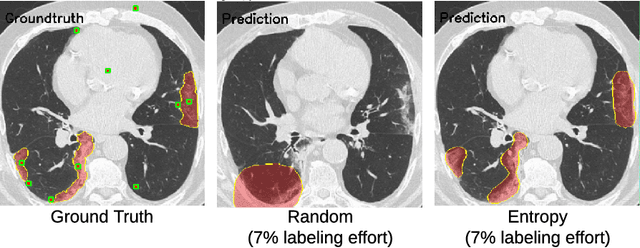
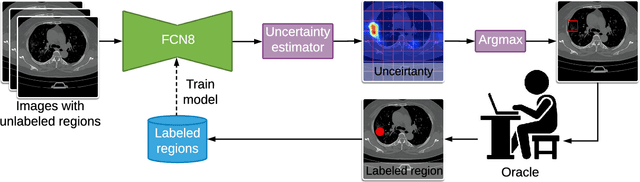
Abstract:One of the key challenges in the battle against the Coronavirus (COVID-19) pandemic is to detect and quantify the severity of the disease in a timely manner. Computed tomographies (CT) of the lungs are effective for assessing the state of the infection. Unfortunately, labeling CT scans can take a lot of time and effort, with up to 150 minutes per scan. We address this challenge introducing a scalable, fast, and accurate active learning system that accelerates the labeling of CT scan images. Conventionally, active learning methods require the labelers to annotate whole images with full supervision, but that can lead to wasted efforts as many of the annotations could be redundant. Thus, our system presents the annotator with unlabeled regions that promise high information content and low annotation cost. Further, the system allows annotators to label regions using point-level supervision, which is much cheaper to acquire than per-pixel annotations. Our experiments on open-source COVID-19 datasets show that using an entropy-based method to rank unlabeled regions yields to significantly better results than random labeling of these regions. Also, we show that labeling small regions of images is more efficient than labeling whole images. Finally, we show that with only 7\% of the labeling effort required to label the whole training set gives us around 90\% of the performance obtained by training the model on the fully annotated training set. Code is available at: \url{https://github.com/IssamLaradji/covid19_active_learning}.
A Weakly Supervised Consistency-based Learning Method for COVID-19 Segmentation in CT Images
Jul 07, 2020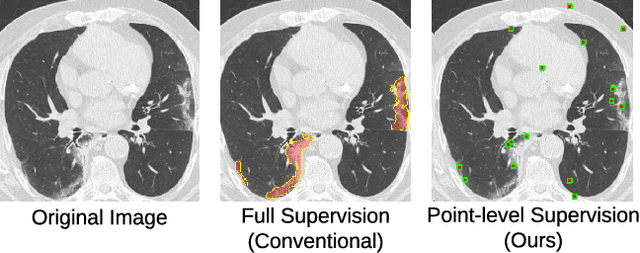
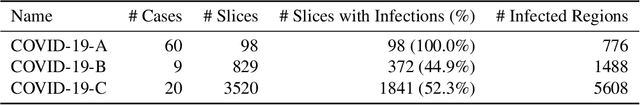

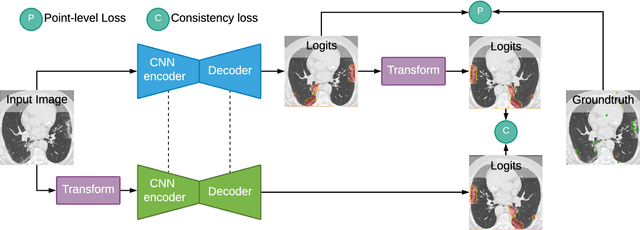
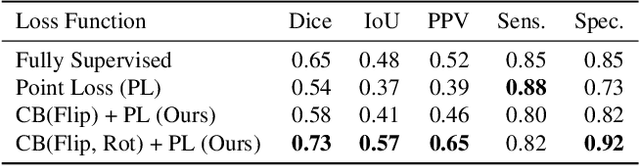
Abstract:Coronavirus Disease 2019 (COVID-19) has spread aggressively across the world causing an existential health crisis. Thus, having a system that automatically detects COVID-19 in tomography (CT) images can assist in quantifying the severity of the illness. Unfortunately, labelling chest CT scans requires significant domain expertise, time, and effort. We address these labelling challenges by only requiring point annotations, a single pixel for each infected region on a CT image. This labeling scheme allows annotators to label a pixel in a likely infected region, only taking 1-3 seconds, as opposed to 10-15 seconds to segment a region. Conventionally, segmentation models train on point-level annotations using the cross-entropy loss function on these labels. However, these models often suffer from low precision. Thus, we propose a consistency-based (CB) loss function that encourages the output predictions to be consistent with spatial transformations of the input images. The experiments on 3 open-source COVID-19 datasets show that this loss function yields significant improvement over conventional point-level loss functions and almost matches the performance of models trained with full supervision with much less human effort. Code is available at: \url{https://github.com/IssamLaradji/covid19_weak_supervision}.
Quantification of Tomographic Patterns associated with COVID-19 from Chest CT
Apr 28, 2020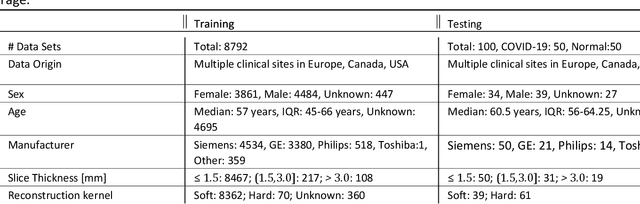
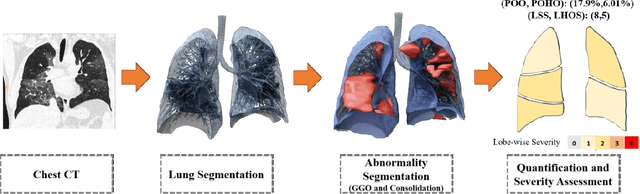
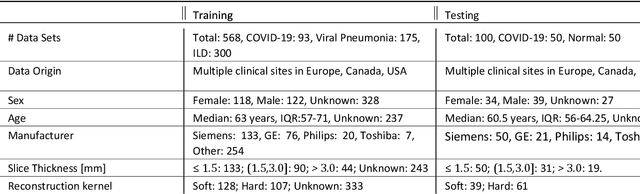
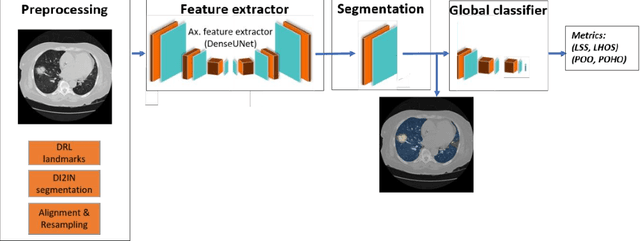
Abstract:Purpose: To present a method that automatically detects and quantifies abnormal tomographic patterns commonly present in COVID-19, namely Ground Glass Opacities (GGO) and consolidations. Given that high opacity abnormalities (i.e., consolidations) were shown to correlate with severe disease, the paper introduces two combined severity measures (Percentage of Opacity, Percentage of High Opacity) and (Lung Severity Score, Lung High Opacity Score). They quantify the extent of overall COVID-19 abnormalities and the presence of high opacity abnormalities, global and lobe-wise, respectively, being computed based on 3D segmentations of lesions, lungs, and lobes. Materials and Methods: The proposed method takes as input a non-contrasted Chest CT and segments the lesions, lungs, and lobes in 3D. It outputs two combined measures of the severity of lung/lobe involvement, quantifying both the extent of COVID-19 abnormalities and presence of high opacities, based on deep learning and deep reinforcement learning. The first measure (POO, POHO) is global, while the second (LSS, LHOS) is lobe-wise. Evaluation is reported on CTs of 100 subjects (50 COVID-19 confirmed and 50 controls) from institutions from Canada, Europe and US. Ground truth is established by manual annotations of lesions, lungs, and lobes. Results: Pearson Correlation Coefficient between method prediction and ground truth is 0.97 (POO), 0.98 (POHO), 0.96 (LSS), 0.96 (LHOS). Automated processing time to compute the severity scores is 10 seconds/case vs 30 mins needed for manual annotations. Conclusion: A new method identifies regions of abnormalities seen in COVID-19 non-contrasted Chest CT and computes (POO, POHO) and (LSS, LHOS) severity scores.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge