Trang Pham
Improving Minimax Estimation Rates for Contaminated Mixture of Multinomial Logistic Experts via Expert Heterogeneity
Jan 31, 2026Abstract:Contaminated mixture of experts (MoE) is motivated by transfer learning methods where a pre-trained model, acting as a frozen expert, is integrated with an adapter model, functioning as a trainable expert, in order to learn a new task. Despite recent efforts to analyze the convergence behavior of parameter estimation in this model, there are still two unresolved problems in the literature. First, the contaminated MoE model has been studied solely in regression settings, while its theoretical foundation in classification settings remains absent. Second, previous works on MoE models for classification capture pointwise convergence rates for parameter estimation without any guaranty of minimax optimality. In this work, we close these gaps by performing, for the first time, the convergence analysis of a contaminated mixture of multinomial logistic experts with homogeneous and heterogeneous structures, respectively. In each regime, we characterize uniform convergence rates for estimating parameters under challenging settings where ground-truth parameters vary with the sample size. Furthermore, we also establish corresponding minimax lower bounds to ensure that these rates are minimax optimal. Notably, our theories offer an important insight into the design of contaminated MoE, that is, expert heterogeneity yields faster parameter estimation rates and, therefore, is more sample-efficient than expert homogeneity.
Tree-Sliced Wasserstein Distance with Nonlinear Projection
May 02, 2025



Abstract:Tree-Sliced methods have recently emerged as an alternative to the traditional Sliced Wasserstein (SW) distance, replacing one-dimensional lines with tree-based metric spaces and incorporating a splitting mechanism for projecting measures. This approach enhances the ability to capture the topological structures of integration domains in Sliced Optimal Transport while maintaining low computational costs. Building on this foundation, we propose a novel nonlinear projectional framework for the Tree-Sliced Wasserstein (TSW) distance, substituting the linear projections in earlier versions with general projections, while ensuring the injectivity of the associated Radon Transform and preserving the well-definedness of the resulting metric. By designing appropriate projections, we construct efficient metrics for measures on both Euclidean spaces and spheres. Finally, we validate our proposed metric through extensive numerical experiments for Euclidean and spherical datasets. Applications include gradient flows, self-supervised learning, and generative models, where our methods demonstrate significant improvements over recent SW and TSW variants.
Spherical Tree-Sliced Wasserstein Distance
Mar 14, 2025



Abstract:Sliced Optimal Transport (OT) simplifies the OT problem in high-dimensional spaces by projecting supports of input measures onto one-dimensional lines and then exploiting the closed-form expression of the univariate OT to reduce the computational burden of OT. Recently, the Tree-Sliced method has been introduced to replace these lines with more intricate structures, known as tree systems. This approach enhances the ability to capture topological information of integration domains in Sliced OT while maintaining low computational cost. Inspired by this approach, in this paper, we present an adaptation of tree systems on OT problems for measures supported on a sphere. As a counterpart to the Radon transform variant on tree systems, we propose a novel spherical Radon transform with a new integration domain called spherical trees. By leveraging this transform and exploiting the spherical tree structures, we derive closed-form expressions for OT problems on the sphere. Consequently, we obtain an efficient metric for measures on the sphere, named Spherical Tree-Sliced Wasserstein (STSW) distance. We provide an extensive theoretical analysis to demonstrate the topology of spherical trees and the well-definedness and injectivity of our Radon transform variant, which leads to an orthogonally invariant distance between spherical measures. Finally, we conduct a wide range of numerical experiments, including gradient flows and self-supervised learning, to assess the performance of our proposed metric, comparing it to recent benchmarks.
Distance-Based Tree-Sliced Wasserstein Distance
Mar 14, 2025



Abstract:To overcome computational challenges of Optimal Transport (OT), several variants of Sliced Wasserstein (SW) has been developed in the literature. These approaches exploit the closed-form expression of the univariate OT by projecting measures onto (one-dimensional) lines. However, projecting measures onto low-dimensional spaces can lead to a loss of topological information. Tree-Sliced Wasserstein distance on Systems of Lines (TSW-SL) has emerged as a promising alternative that replaces these lines with a more advanced structure called tree systems. The tree structures enhance the ability to capture topological information of the metric while preserving computational efficiency. However, at the core of TSW-SL, the splitting maps, which serve as the mechanism for pushing forward measures onto tree systems, focus solely on the position of the measure supports while disregarding the projecting domains. Moreover, the specific splitting map used in TSW-SL leads to a metric that is not invariant under Euclidean transformations, a typically expected property for OT on Euclidean space. In this work, we propose a novel class of splitting maps that generalizes the existing one studied in TSW-SL enabling the use of all positional information from input measures, resulting in a novel Distance-based Tree-Sliced Wasserstein (Db-TSW) distance. In addition, we introduce a simple tree sampling process better suited for Db-TSW, leading to an efficient GPU-friendly implementation for tree systems, similar to the original SW. We also provide a comprehensive theoretical analysis of proposed class of splitting maps to verify the injectivity of the corresponding Radon Transform, and demonstrate that Db-TSW is an Euclidean invariant metric. We empirically show that Db-TSW significantly improves accuracy compared to recent SW variants while maintaining low computational cost via a wide range of experiments.
Tree-Sliced Wasserstein Distance on a System of Lines
Jun 19, 2024



Abstract:Sliced Wasserstein (SW) distance in Optimal Transport (OT) is widely used in various applications thanks to its statistical effectiveness and computational efficiency. On the other hand, Tree Wassenstein (TW) and Tree-sliced Wassenstein (TSW) are instances of OT for probability measures where its ground cost is a tree metric. TSW also has a low computational complexity, i.e. linear to the number of edges in the tree. Especially, TSW is identical to SW when the tree is a chain. While SW is prone to loss of topological information of input measures due to relying on one-dimensional projection, TSW is more flexible and has a higher degree of freedom by choosing a tree rather than a line to alleviate the curse of dimensionality in SW. However, for practical applications, popular tree metric sampling methods are heavily built upon given supports, which limits their capacity to adapt to new supports. In this paper, we propose the Tree-Sliced Wasserstein distance on a System of Lines (TSW-SL), which brings a connection between SW and TSW. Compared to SW and TSW, our TSW-SL benefits from the higher degree of freedom of TSW while being suitable to dynamic settings as SW. In TSW-SL, we use a variant of the Radon Transform to project measures onto a system of lines, resulting in measures on a space with a tree metric, then leverage TW to efficiently compute distances between them. We empirically verify the advantages of TSW-SL over the traditional SW by conducting a variety of experiments on gradient flows, image style transfer, and generative models.
Statistical Advantages of Perturbing Cosine Router in Sparse Mixture of Experts
May 23, 2024



Abstract:The cosine router in sparse Mixture of Experts (MoE) has recently emerged as an attractive alternative to the conventional linear router. Indeed, the cosine router demonstrates favorable performance in image and language tasks and exhibits better ability to mitigate the representation collapse issue, which often leads to parameter redundancy and limited representation potentials. Despite its empirical success, a comprehensive analysis of the cosine router in sparse MoE has been lacking. Considering the least square estimation of the cosine routing sparse MoE, we demonstrate that due to the intrinsic interaction of the model parameters in the cosine router via some partial differential equations, regardless of the structures of the experts, the estimation rates of experts and model parameters can be as slow as $\mathcal{O}(1/\log^{\tau}(n))$ where $\tau > 0$ is some constant and $n$ is the sample size. Surprisingly, these pessimistic non-polynomial convergence rates can be circumvented by the widely used technique in practice to stabilize the cosine router -- simply adding noises to the $\mathbb{L}_{2}$ norms in the cosine router, which we refer to as \textit{perturbed cosine router}. Under the strongly identifiable settings of the expert functions, we prove that the estimation rates for both the experts and model parameters under the perturbed cosine routing sparse MoE are significantly improved to polynomial rates. Finally, we conduct extensive simulation studies in both synthetic and real data settings to empirically validate our theoretical results.
Mixture of Experts Meets Prompt-Based Continual Learning
May 23, 2024Abstract:Exploiting the power of pre-trained models, prompt-based approaches stand out compared to other continual learning solutions in effectively preventing catastrophic forgetting, even with very few learnable parameters and without the need for a memory buffer. While existing prompt-based continual learning methods excel in leveraging prompts for state-of-the-art performance, they often lack a theoretical explanation for the effectiveness of prompting. This paper conducts a theoretical analysis to unravel how prompts bestow such advantages in continual learning, thus offering a new perspective on prompt design. We first show that the attention block of pre-trained models like Vision Transformers inherently encodes a special mixture of experts architecture, characterized by linear experts and quadratic gating score functions. This realization drives us to provide a novel view on prefix tuning, reframing it as the addition of new task-specific experts, thereby inspiring the design of a novel gating mechanism termed Non-linear Residual Gates (NoRGa). Through the incorporation of non-linear activation and residual connection, NoRGa enhances continual learning performance while preserving parameter efficiency. The effectiveness of NoRGa is substantiated both theoretically and empirically across diverse benchmarks and pretraining paradigms.
Relational dynamic memory networks
Oct 30, 2018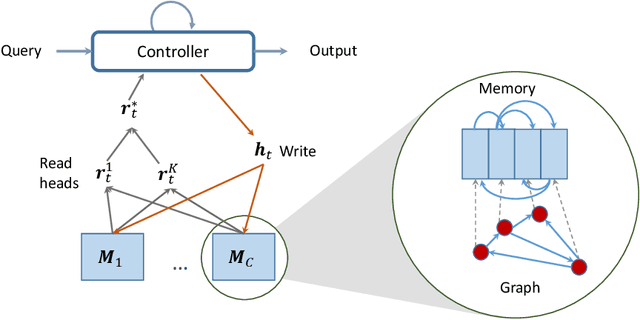
Abstract:Neural networks excel in detecting regular patterns but are less successful in representing and manipulating complex data structures, possibly due to the lack of an external memory. This has led to the recent development of a new line of architectures known as Memory-Augmented Neural Networks (MANNs), each of which consists of a neural network that interacts with an external memory matrix. However, this RAM-like memory matrix is unstructured and thus does not naturally encode structured objects. Here we design a new MANN dubbed Relational Dynamic Memory Network (RMDN) to bridge the gap. Like existing MANNs, RMDN has a neural controller but its memory is structured as multi-relational graphs. RMDN uses the memory to represent and manipulate graph-structured data in response to query; and as a neural network, RMDN is trainable from labeled data. Thus RMDN learns to answer queries about a set of graph-structured objects without explicit programming. We evaluate the capability of RMDN on several important prediction problems, including software vulnerability, molecular bioactivity and chemical-chemical interaction. Results demonstrate the efficacy of the proposed model.
Graph Memory Networks for Molecular Activity Prediction
Jan 27, 2018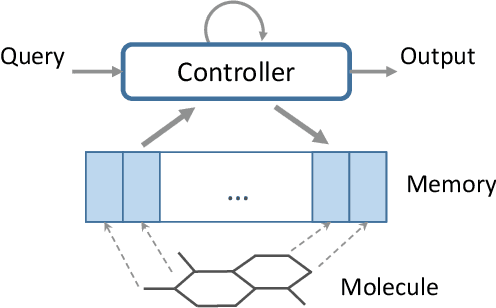
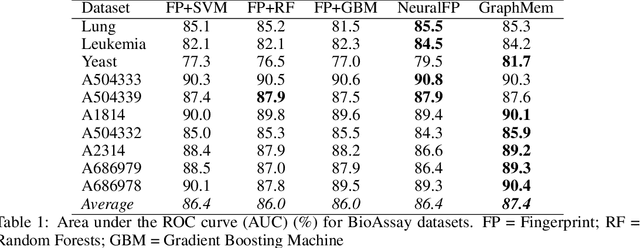

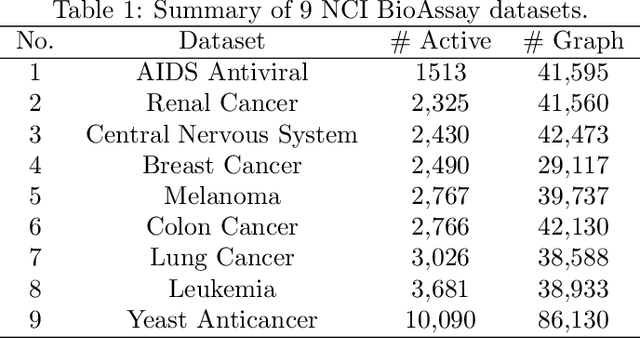
Abstract:Molecular activity prediction is critical in drug design. Machine learning techniques such as kernel methods and random forests have been successful for this task. These models require fixed-size feature vectors as input while the molecules are variable in size and structure. As a result, fixed-size fingerprint representation is poor in handling substructures for large molecules. In addition, molecular activity tests, or a so-called BioAssays, are relatively small in the number of tested molecules due to its complexity. Here we approach the problem through deep neural networks as they are flexible in modeling structured data such as grids, sequences and graphs. We train multiple BioAssays using a multi-task learning framework, which combines information from multiple sources to improve the performance of prediction, especially on small datasets. We propose Graph Memory Network (GraphMem), a memory-augmented neural network to model the graph structure in molecules. GraphMem consists of a recurrent controller coupled with an external memory whose cells dynamically interact and change through a multi-hop reasoning process. Applied to the molecules, the dynamic interactions enable an iterative refinement of the representation of molecular graphs with multiple bond types. GraphMem is capable of jointly training on multiple datasets by using a specific-task query fed to the controller as an input. We demonstrate the effectiveness of the proposed model for separately and jointly training on more than 100K measurements, spanning across 9 BioAssay activity tests.
Graph Classification via Deep Learning with Virtual Nodes
Aug 14, 2017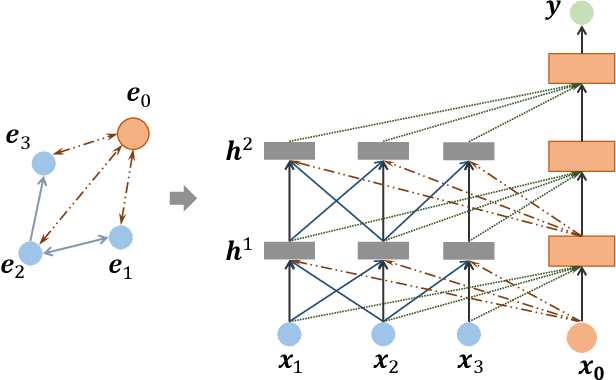

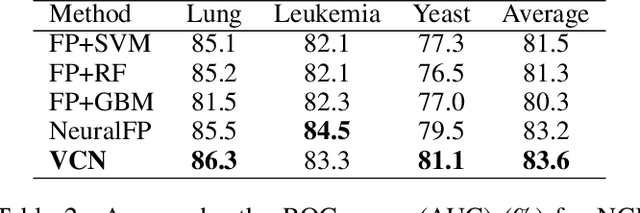
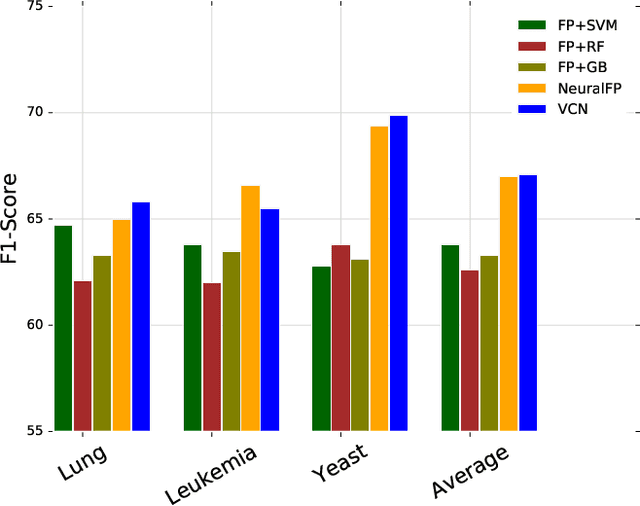
Abstract:Learning representation for graph classification turns a variable-size graph into a fixed-size vector (or matrix). Such a representation works nicely with algebraic manipulations. Here we introduce a simple method to augment an attributed graph with a virtual node that is bidirectionally connected to all existing nodes. The virtual node represents the latent aspects of the graph, which are not immediately available from the attributes and local connectivity structures. The expanded graph is then put through any node representation method. The representation of the virtual node is then the representation of the entire graph. In this paper, we use the recently introduced Column Network for the expanded graph, resulting in a new end-to-end graph classification model dubbed Virtual Column Network (VCN). The model is validated on two tasks: (i) predicting bio-activity of chemical compounds, and (ii) finding software vulnerability from source code. Results demonstrate that VCN is competitive against well-established rivals.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge