Suzanne Tamang
Domain Shift Analysis in Chest Radiographs Classification in a Veterans Healthcare Administration Population
Jul 30, 2024



Abstract:Objectives: This study aims to assess the impact of domain shift on chest X-ray classification accuracy and to analyze the influence of ground truth label quality and demographic factors such as age group, sex, and study year. Materials and Methods: We used a DenseNet121 model pretrained MIMIC-CXR dataset for deep learning-based multilabel classification using ground truth labels from radiology reports extracted using the CheXpert and CheXbert Labeler. We compared the performance of the 14 chest X-ray labels on the MIMIC-CXR and Veterans Healthcare Administration chest X-ray dataset (VA-CXR). The VA-CXR dataset comprises over 259k chest X-ray images spanning between the years 2010 and 2022. Results: The validation of ground truth and the assessment of multi-label classification performance across various NLP extraction tools revealed that the VA-CXR dataset exhibited lower disagreement rates than the MIMIC-CXR datasets. Additionally, there were notable differences in AUC scores between models utilizing CheXpert and CheXbert. When evaluating multi-label classification performance across different datasets, minimal domain shift was observed in unseen datasets, except for the label "Enlarged Cardiomediastinum." The study year's subgroup analyses exhibited the most significant variations in multi-label classification model performance. These findings underscore the importance of considering domain shifts in chest X-ray classification tasks, particularly concerning study years. Conclusion: Our study reveals the significant impact of domain shift and demographic factors on chest X-ray classification, emphasizing the need for improved transfer learning and equitable model development. Addressing these challenges is crucial for advancing medical imaging and enhancing patient care.
VISION: Toward a Standardized Process for Radiology Image Management at the National Level
Apr 29, 2024Abstract:The compilation and analysis of radiological images poses numerous challenges for researchers. The sheer volume of data as well as the computational needs of algorithms capable of operating on images are extensive. Additionally, the assembly of these images alone is difficult, as these exams may differ widely in terms of clinical context, structured annotation available for model training, modality, and patient identifiers. In this paper, we describe our experiences and challenges in establishing a trusted collection of radiology images linked to the United States Department of Veterans Affairs (VA) electronic health record database. We also discuss implications in making this repository research-ready for medical investigators. Key insights include uncovering the specific procedures required for transferring images from a clinical to a research-ready environment, as well as roadblocks and bottlenecks in this process that may hinder future efforts at automation.
Question-Answering System Extracts Information on Injection Drug Use from Clinical Progress Notes
May 15, 2023Abstract:Injection drug use (IDU) is a dangerous health behavior that increases mortality and morbidity. Identifying IDU early and initiating harm reduction interventions can benefit individuals at risk. However, extracting IDU behaviors from patients' electronic health records (EHR) is difficult because there is no International Classification of Disease (ICD) code and the only place IDU information can be indicated are unstructured free-text clinical progress notes. Although natural language processing (NLP) can efficiently extract this information from unstructured data, there are no validated tools. To address this gap in clinical information, we design and demonstrate a question-answering (QA) framework to extract information on IDU from clinical progress notes. Unlike other methods discussed in the literature, the QA model is able to extract various types of information without being constrained by predefined entities, relations, or concepts. Our framework involves two main steps: (1) generating a gold-standard QA dataset and (2) developing and testing the QA model. This paper also demonstrates the QA model's ability to extract IDU-related information on temporally out-of-distribution data. The results indicate that the majority (51%) of the extracted information by the QA model exactly matches the gold-standard answer and 73% of them contain the gold-standard answer with some additional surrounding words.
Snomed2Vec: Random Walk and Poincaré Embeddings of a Clinical Knowledge Base for Healthcare Analytics
Jul 19, 2019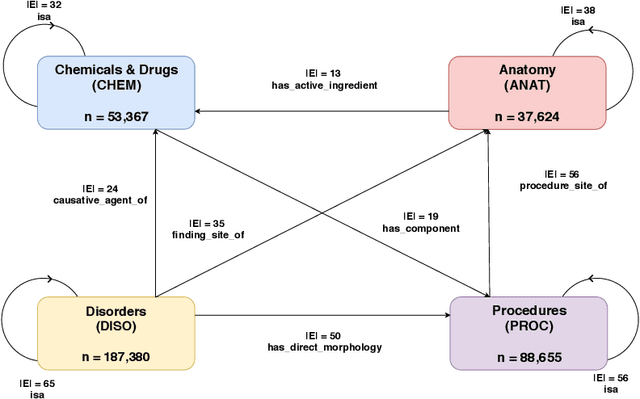
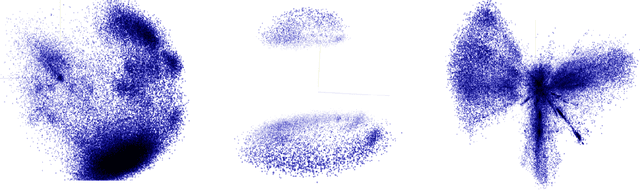
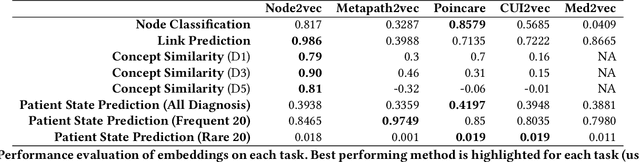
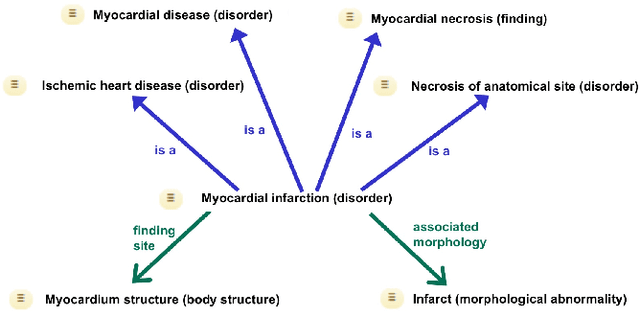

Abstract:Representation learning methods that transform encoded data (e.g., diagnosis and drug codes) into continuous vector spaces (i.e., vector embeddings) are critical for the application of deep learning in healthcare. Initial work in this area explored the use of variants of the word2vec algorithm to learn embeddings for medical concepts from electronic health records or medical claims datasets. We propose learning embeddings for medical concepts by using graph-based representation learning methods on SNOMED-CT, a widely popular knowledge graph in the healthcare domain with numerous operational and research applications. Current work presents an empirical analysis of various embedding methods, including the evaluation of their performance on multiple tasks of biomedical relevance (node classification, link prediction, and patient state prediction). Our results show that concept embeddings derived from the SNOMED-CT knowledge graph significantly outperform state-of-the-art embeddings, showing 5-6x improvement in ``concept similarity" and 6-20\% improvement in patient diagnosis.
A Knowledge Graph-based Approach for Exploring the U.S. Opioid Epidemic
May 27, 2019
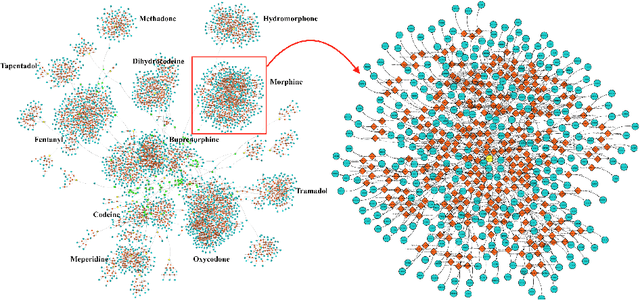
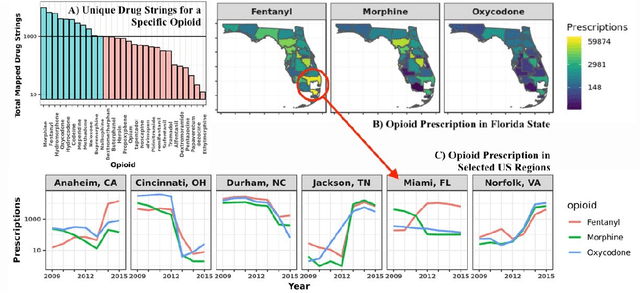
Abstract:The United States is in the midst of an opioid epidemic with recent estimates indicating that more than 130 people die every day due to drug overdose. The over-prescription and addiction to opioid painkillers, heroin, and synthetic opioids, has led to a public health crisis and created a huge social and economic burden. Statistical learning methods that use data from multiple clinical centers across the US to detect opioid over-prescribing trends and predict possible opioid misuse are required. However, the semantic heterogeneity in the representation of clinical data across different centers makes the development and evaluation of such methods difficult and non-trivial. We create the Opioid Drug Knowledge Graph (ODKG) -- a network of opioid-related drugs, active ingredients, formulations, combinations, and brand names. We use the ODKG to normalize drug strings in a clinical data warehouse consisting of patient data from over 400 healthcare facilities in 42 different states. We showcase the use of ODKG to generate summary statistics of opioid prescription trends across US regions. These methods and resources can aid the development of advanced and scalable models to monitor the opioid epidemic and to detect illicit opioid misuse behavior. Our work is relevant to policymakers and pain researchers who wish to systematically assess factors that contribute to opioid over-prescribing and iatrogenic opioid addiction in the US.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge