Sobhan Shafiei
Colored Kimia Path24 Dataset: Configurations and Benchmarks with Deep Embeddings
Feb 15, 2021
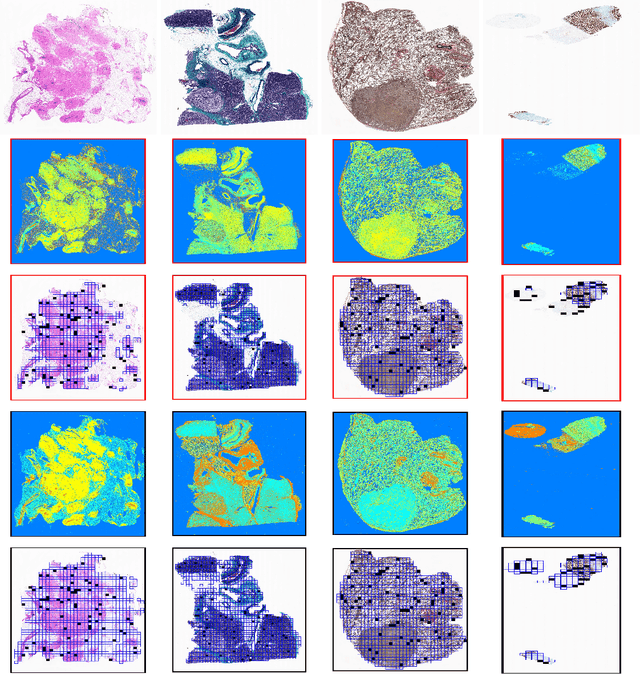
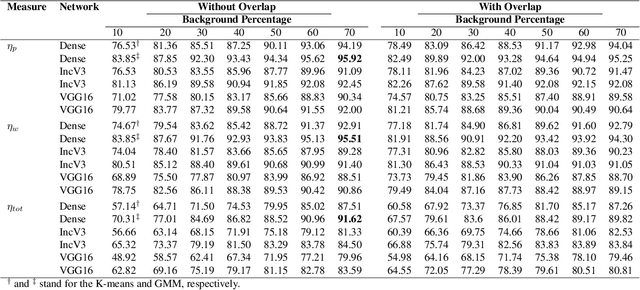
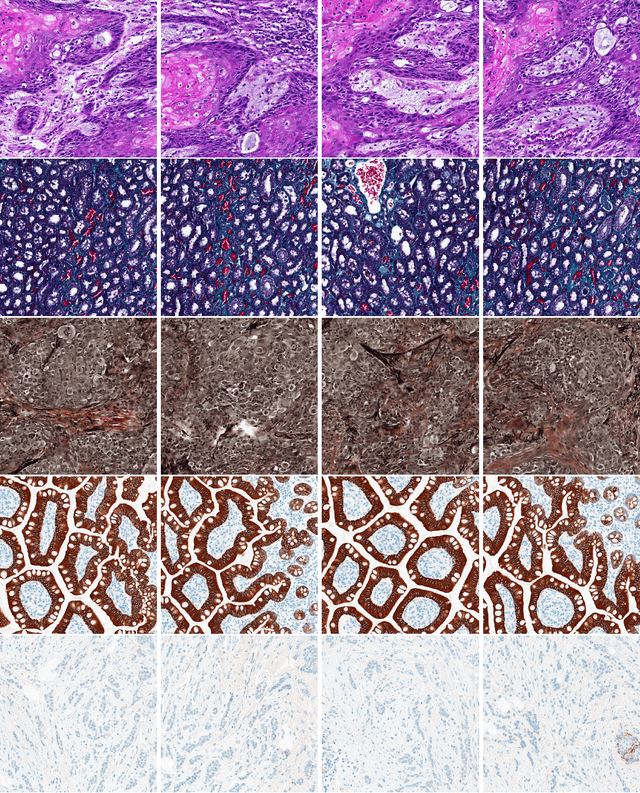
Abstract:The Kimia Path24 dataset has been introduced as a classification and retrieval dataset for digital pathology. Although it provides multi-class data, the color information has been neglected in the process of extracting patches. The staining information plays a major role in the recognition of tissue patterns. To address this drawback, we introduce the color version of Kimia Path24 by recreating sample patches from all 24 scans to propose Kimia Path24C. We run extensive experiments to determine the best configuration for selected patches. To provide preliminary results for setting a benchmark for the new dataset, we utilize VGG16, InceptionV3 and DenseNet-121 model as feature extractors. Then, we use these feature vectors to retrieve test patches. The accuracy of image retrieval using DenseNet was 95.92% while the highest accuracy using InceptionV3 and VGG16 reached 92.45% and 92%, respectively. We also experimented with "deep barcodes" and established that with a small loss in accuracy (e.g., 93.43% for binarized features for DenseNet instead of 95.92% when the features themselves are used), the search operations can be significantly accelerated.
Fine-Tuning and Training of DenseNet for Histopathology Image Representation Using TCGA Diagnostic Slides
Jan 20, 2021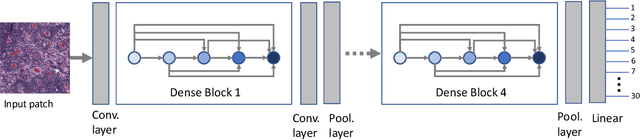
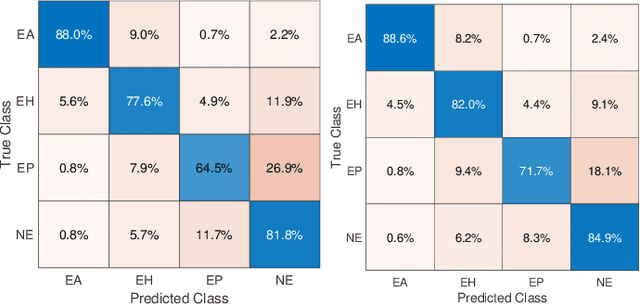
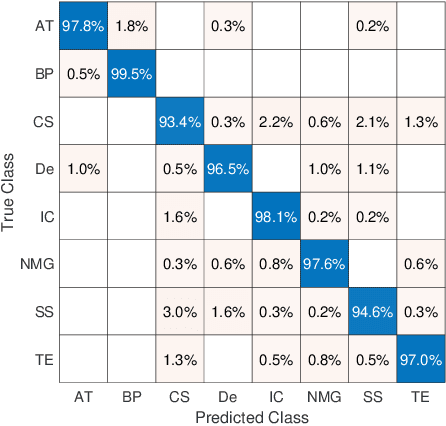
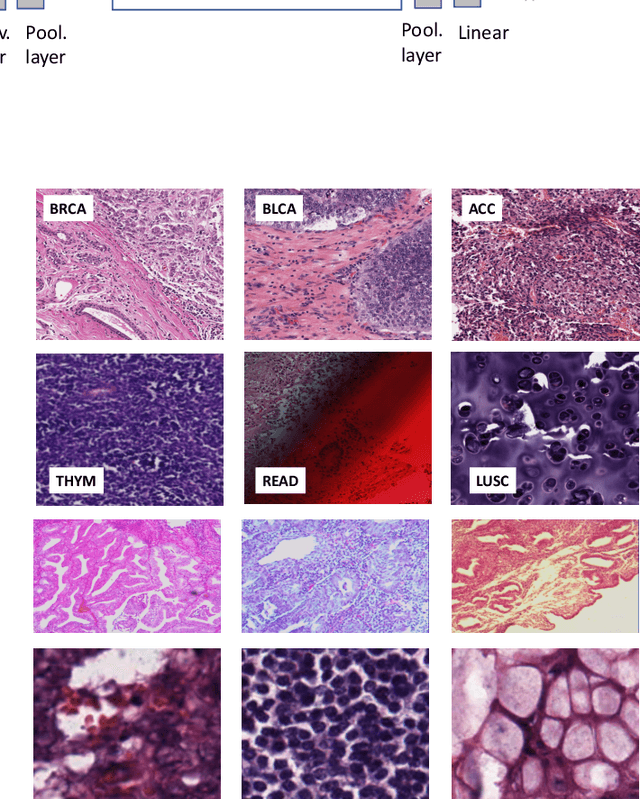
Abstract:Feature vectors provided by pre-trained deep artificial neural networks have become a dominant source for image representation in recent literature. Their contribution to the performance of image analysis can be improved through finetuning. As an ultimate solution, one might even train a deep network from scratch with the domain-relevant images, a highly desirable option which is generally impeded in pathology by lack of labeled images and the computational expense. In this study, we propose a new network, namely KimiaNet, that employs the topology of the DenseNet with four dense blocks, fine-tuned and trained with histopathology images in different configurations. We used more than 240,000 image patches with 1000x1000 pixels acquired at 20x magnification through our proposed "highcellularity mosaic" approach to enable the usage of weak labels of 7,126 whole slide images of formalin-fixed paraffin-embedded human pathology samples publicly available through the The Cancer Genome Atlas (TCGA) repository. We tested KimiaNet using three public datasets, namely TCGA, endometrial cancer images, and colorectal cancer images by evaluating the performance of search and classification when corresponding features of different networks are used for image representation. As well, we designed and trained multiple convolutional batch-normalized ReLU (CBR) networks. The results show that KimiaNet provides superior results compared to the original DenseNet and smaller CBR networks when used as feature extractor to represent histopathology images.
Supervision and Source Domain Impact on Representation Learning: A Histopathology Case Study
May 10, 2020

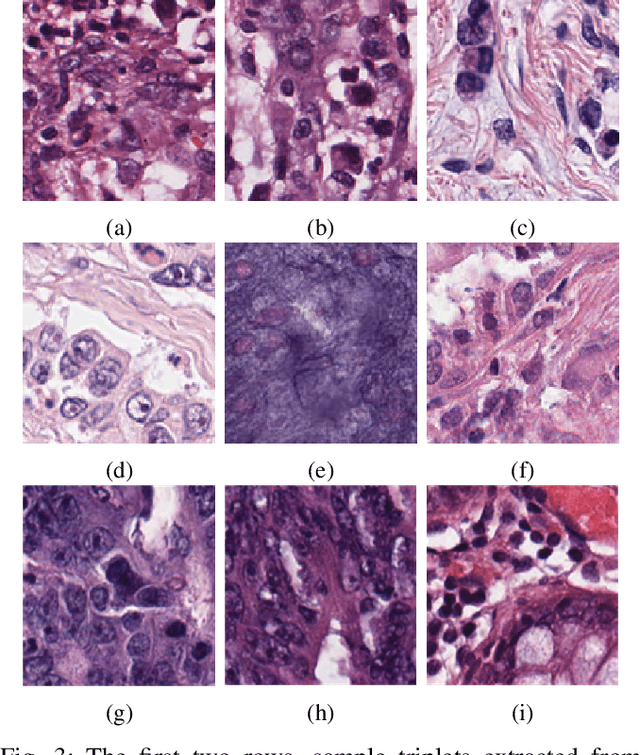

Abstract:As many algorithms depend on a suitable representation of data, learning unique features is considered a crucial task. Although supervised techniques using deep neural networks have boosted the performance of representation learning, the need for a large set of labeled data limits the application of such methods. As an example, high-quality delineations of regions of interest in the field of pathology is a tedious and time-consuming task due to the large image dimensions. In this work, we explored the performance of a deep neural network and triplet loss in the area of representation learning. We investigated the notion of similarity and dissimilarity in pathology whole-slide images and compared different setups from unsupervised and semi-supervised to supervised learning in our experiments. Additionally, different approaches were tested, applying few-shot learning on two publicly available pathology image datasets. We achieved high accuracy and generalization when the learned representations were applied to two different pathology datasets.
Fisher Discriminant Triplet and Contrastive Losses for Training Siamese Networks
Apr 05, 2020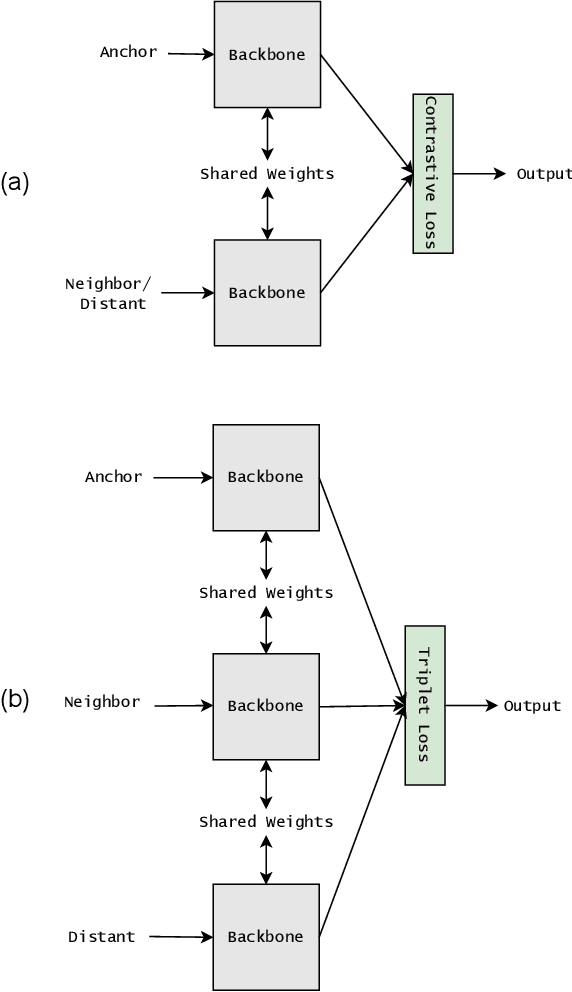
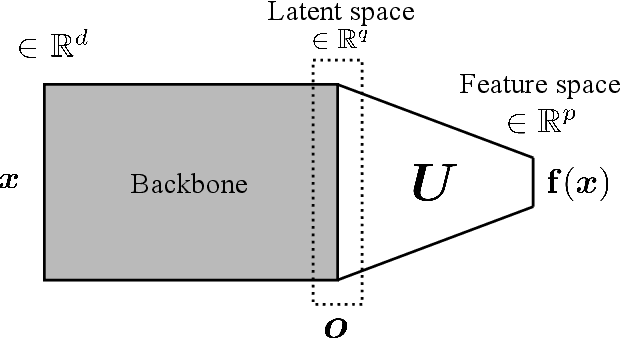
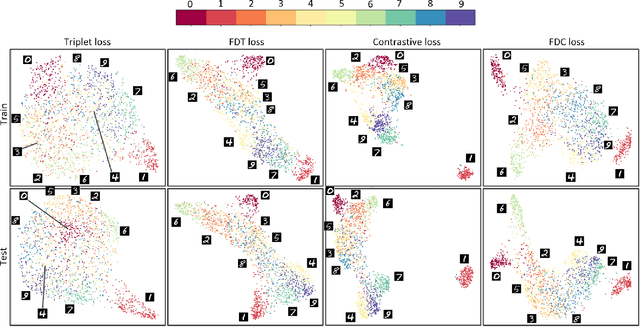
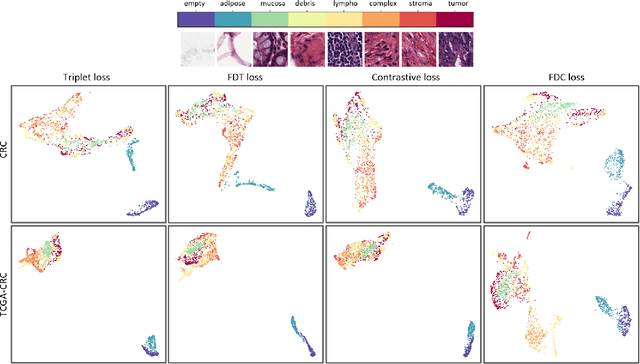
Abstract:Siamese neural network is a very powerful architecture for both feature extraction and metric learning. It usually consists of several networks that share weights. The Siamese concept is topology-agnostic and can use any neural network as its backbone. The two most popular loss functions for training these networks are the triplet and contrastive loss functions. In this paper, we propose two novel loss functions, named Fisher Discriminant Triplet (FDT) and Fisher Discriminant Contrastive (FDC). The former uses anchor-neighbor-distant triplets while the latter utilizes pairs of anchor-neighbor and anchor-distant samples. The FDT and FDC loss functions are designed based on the statistical formulation of the Fisher Discriminant Analysis (FDA), which is a linear subspace learning method. Our experiments on the MNIST and two challenging and publicly available histopathology datasets show the effectiveness of the proposed loss functions.
Pan-Cancer Diagnostic Consensus Through Searching Archival Histopathology Images Using Artificial Intelligence
Nov 20, 2019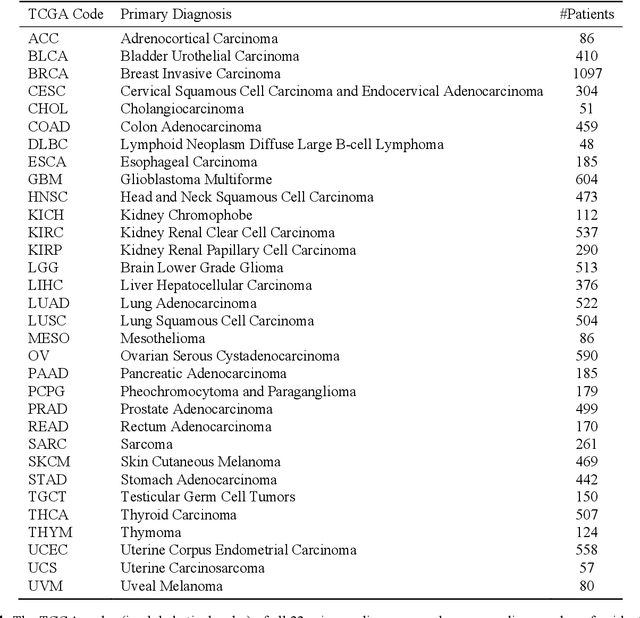
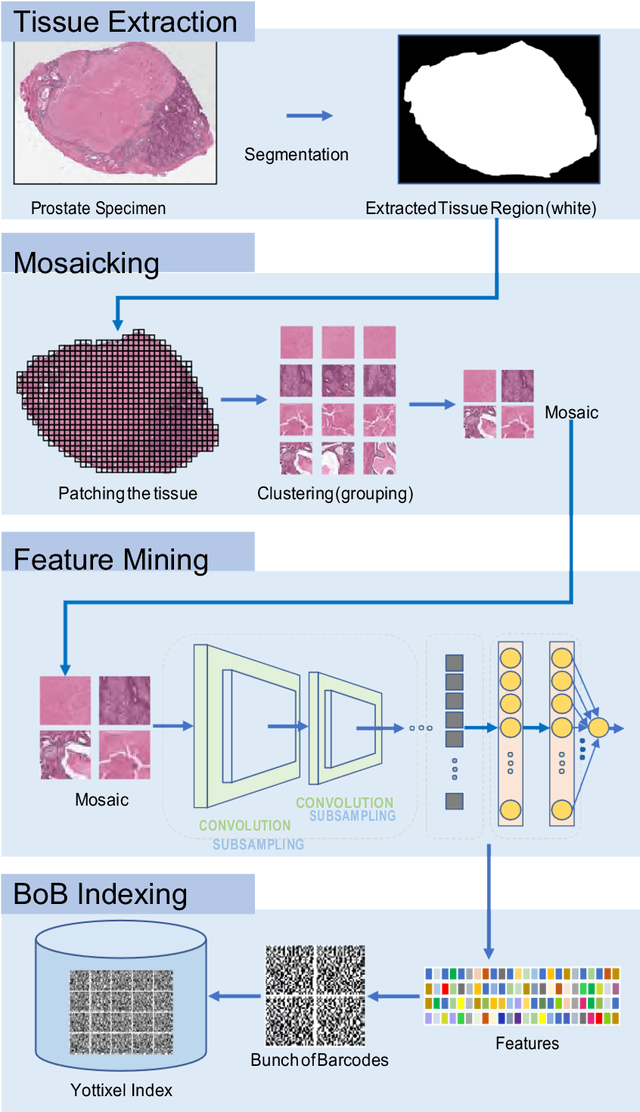

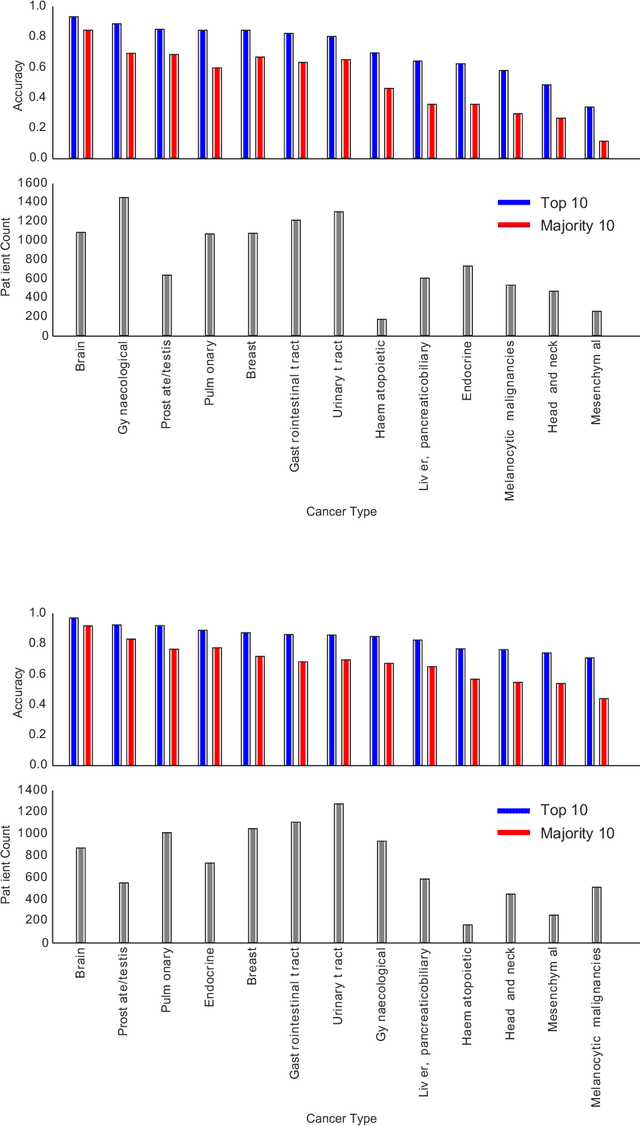
Abstract:The emergence of digital pathology has opened new horizons for histopathology and cytology. Artificial-intelligence algorithms are able to operate on digitized slides to assist pathologists with diagnostic tasks. Whereas machine learning involving classification and segmentation methods have obvious benefits for image analysis in pathology, image search represents a fundamental shift in computational pathology. Matching the pathology of new patients with already diagnosed and curated cases offers pathologist a novel approach to improve diagnostic accuracy through visual inspection of similar cases and computational majority vote for consensus building. In this study, we report the results from searching the largest public repository (The Cancer Genome Atlas [TCGA] program by National Cancer Institute, USA) of whole slide images from almost 11,000 patients depicting different types of malignancies. For the first time, we successfully indexed and searched almost 30,000 high-resolution digitized slides constituting 16 terabytes of data comprised of 20 million 1000x1000 pixels image patches. The TCGA image database covers 25 anatomic sites and contains 32 cancer subtypes. High-performance storage and GPU power were employed for experimentation. The results were assessed with conservative "majority voting" to build consensus for subtype diagnosis through vertical search and demonstrated high accuracy values for both frozen sections slides (e.g., bladder urothelial carcinoma 93%, kidney renal clear cell carcinoma 97%, and ovarian serous cystadenocarcinoma 99%) and permanent histopathology slides (e.g., prostate adenocarcinoma 98%, skin cutaneous melanoma 99%, and thymoma 100%). The key finding of this validation study was that computational consensus appears to be possible for rendering diagnoses if a sufficiently large number of searchable cases are available for each cancer subtype.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge