Shireen Elhabian
MorphoFlow: Sparse-Supervised Generative Shape Modeling with Adaptive Latent Relevance
Apr 13, 2026Abstract:Statistical shape modeling (SSM) is central to population level analysis of anatomical variability, yet most existing approaches rely on densely annotated segmentations and fixed latent representations. These requirements limit scalability and reduce flexibility when modeling complex anatomical variation. We introduce MorphoFlow, a sparse supervised generative shape modeling framework that learns compact probabilistic shape representations directly from sparse surface annotations. MorphoFlow integrates neural implicit shape representations with an autodecoder formulation and autoregressive normalizing flows to learn an expressive probabilistic density over the latent shape space. The neural implicit representation enables resolution-agnostic modeling of 3D anatomy, while the autodecoder formulation supports direct optimization of per-instance latent codes under sparse supervision. The autoregressive flow captures the distribution of latent anatomical variability providing a tractable, likelihood-based generative model of shapes. To promote compact and structured latent representations, we incorporate adaptive latent relevance weighting through sparsity-inducing priors, enabling the model to regulate the contribution of individual latent dimensions according to their relevance to the underlying anatomical variation while preserving generative expressivity. The resulting latent space supports uncertainty quantification and anatomically plausible shape synthesis without manual latent dimensionality tuning. Evaluation on publicly available lumbar vertebrae and femur datasets demonstrates accurate high-resolution reconstruction from sparse inputs and recovery of structured modes of anatomical variation consistent with population level trends.
MedConcept: Unsupervised Concept Discovery for Interpretability in Medical VLMs
Apr 13, 2026Abstract:While medical Vision-Language models (VLMs) achieve strong performance on tasks such as tumor or organ segmentation and diagnosis prediction, their opaque latent representations limit clinical trust and the ability to explain predictions. Interpretability of these multimodal representations are therefore essential for the trustworthy clinical deployment of pretrained medical VLMs. However, current interpretability methods, such as gradient- or attention-based visualizations, are often limited to specific tasks such as classification. Moreover, they do not provide concept-level explanations derived from shared pretrained representations that can be reused across downstream tasks. We introduce MedConcept, a framework that uncovers latent medical concepts in a fully unsupervised manner and grounds them in clinically verifiable textual semantics. MedConcept identifies sparse neuron-level concept activations from pretrained VLM representations and translates them into pseudo-report-style summaries, enabling physician-level inspection of internal model reasoning. To address the lack of quantitative evaluation in concept-based interpretability, we introduce a quantitative semantic verification protocol that leverages an independent pretrained medical LLM as a frozen external evaluator to assess concept alignment with radiology reports. We define three concept scores, Aligned, Unaligned, and Uncertain, to quantify semantic support, contradiction, or ambiguity relative to radiology reports and use them exclusively for post hoc evaluation. These scores provide a quantitative baseline for assessing interpretability in medical VLMs. All codes, prompt and data to be released on acceptance. Ke
SIMPLER: H&E-Informed Representation Learning for Structured Illumination Microscopy
Apr 11, 2026Abstract:Structured Illumination Microscopy (SIM) enables rapid, high-contrast optical sectioning of fresh tissue without staining or physical sectioning, making it promising for intraoperative and point-of-care diagnostics. Recent foundation and large-scale self-supervised models in digital pathology have demonstrated strong performance on section-based modalities such as Hematoxylin and Eosin (H&E) and immunohistochemistry (IHC). However, these approaches are predominantly trained on thin tissue sections and do not explicitly address thick-tissue fluorescence modalities such as SIM. When transferred directly to SIM, performance is constrained by substantial modality shift, and naive fine-tuning often overfits to modality-specific appearance rather than underlying histological structure. We introduce SIMPLER (Structured Illumination Microscopy-Powered Learning for Embedding Representations), a cross-modality self-supervised pretraining framework that leverages H&E as a semantic anchor to learn reusable SIM representations. H&E encodes rich cellular and glandular structure aligned with established clinical annotations, while SIM provides rapid, nondestructive imaging of fresh tissue. During pretraining, SIM and H&E are progressively aligned through adversarial, contrastive, and reconstruction-based objectives, encouraging SIM embeddings to internalize histological structure from H&E without collapsing modality-specific characteristics. A single pretrained SIMPLER encoder transfers across multiple downstream tasks, including multiple instance learning and morphological clustering, consistently outperforming SIM models trained from scratch or H&E-only pretraining. Importantly, joint alignment enhances SIM performance without degrading H&E representations, demonstrating asymmetric enrichment rather
AC-MIL: Weakly Supervised Atrial LGE-MRI Quality Assessment via Adversarial Concept Disentanglement
Apr 11, 2026Abstract:High-quality Late Gadolinium Enhancement (LGE) MRI can be helpful for atrial fibrillation management, yet scan quality is frequently compromised by patient motion, irregular breathing, and suboptimal image acquisition timing. While Multiple Instance Learning (MIL) has emerged as a powerful tool for automated quality assessment under weak supervision, current state-of-the-art methods map localized visual evidence to a single, opaque global feature vector. This black box approach fails to provide actionable feedback on specific failure modes, obscuring whether a scan degrades due to motion blur, inadequate contrast, or a lack of anatomical context. In this paper, we propose Adversarial Concept-MIL (AC-MIL), a weakly supervised framework that decomposes global image quality into clinically defined radiological concepts using only volume-level supervision. To capture latent quality variations without entangling predefined concepts, our framework incorporates an unsupervised residual branch guided by an adversarial erasure mechanism to strictly prevent information leakage. Furthermore, we introduce a spatial diversity constraint that penalizes overlap between distinct concept attention maps, ensuring localized and interpretable feature extraction. Extensive experiments on a clinical dataset of atrial LGE-MRI volumes demonstrate that AC-MIL successfully opens the MIL black box, providing highly localized spatial concept maps that allow clinicians to pinpoint the specific causes of non-diagnostic scans. Crucially, our framework achieves this deep clinical transparency while maintaining highly competitive ordinal grading performance against existing baselines. Code to be released on acceptance.
Unrolled Reconstruction with Integrated Super-Resolution for Accelerated 3D LGE MRI
Mar 18, 2026Abstract:Accelerated 3D late gadolinium enhancement (LGE) MRI requires robust reconstruction methods to recover thin atrial structures from undersampled k-space data. While unrolled model-based networks effectively integrate physics-driven data consistency with learned priors, they operate at the acquired resolution and may fail to fully recover high-frequency detail. We propose a hybrid unrolled reconstruction framework in which an Enhanced Deep Super-Resolution (EDSR) network replaces the proximal operator within each iteration of the optimization loop, enabling joint super-resolution enhancement and data consistency enforcement. The model is trained end-to-end on retrospectively undersampled preclinical 3D LGE datasets and compared against compressed sensing, Model-Based Deep Learning (MoDL), and self-guided Deep Image Prior (DIP) baselines. Across acceleration factors, the proposed method consistently improves PSNR and SSIM over standard unrolled reconstruction and better preserves fine cardiac structures, leading to improved LA (left atrium) segmentation performance. These results demonstrate that integrating super-resolution priors directly within model-based reconstruction provides measurable gains in accelerated 3D LGE MRI.
TimeSynth: A Framework for Uncovering Systematic Biases in Time Series Forecasting
Feb 11, 2026Abstract:Time series forecasting is a fundamental tool with wide ranging applications, yet recent debates question whether complex nonlinear architectures truly outperform simple linear models. Prior claims of dominance of the linear model often stem from benchmarks that lack diverse temporal dynamics and employ biased evaluation protocols. We revisit this debate through TimeSynth, a structured framework that emulates key properties of real world time series,including non-stationarity, periodicity, trends, and phase modulation by creating synthesized signals whose parameters are derived from real-world time series. Evaluating four model families Linear, Multi Layer Perceptrons (MLP), Convolutional Neural Networks (CNNs), and Transformers, we find a systematic bias in linear models: they collapse to simple oscillation regardless of signal complexity. Nonlinear models avoid this collapse and gain clear advantages as signal complexity increases. Notably, Transformers and CNN based models exhibit slightly greater adaptability to complex modulated signals compared to MLPs. Beyond clean forecasting, the framework highlights robustness differences under distribution and noise shifts and removes biases of prior benchmarks by using independent instances for train, test, and validation for each signal family. Collectively, TimeSynth provides a principled foundation for understanding when different forecasting approaches succeed or fail, moving beyond oversimplified claims of model equivalence.
Mesh2SSM++: A Probabilistic Framework for Unsupervised Learning of Statistical Shape Model of Anatomies from Surface Meshes
Feb 11, 2025
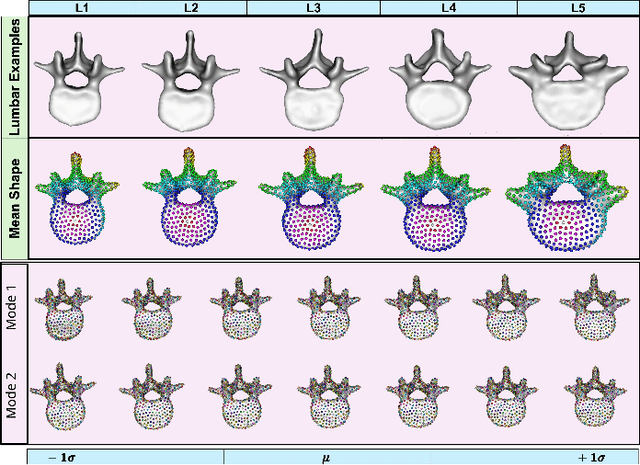
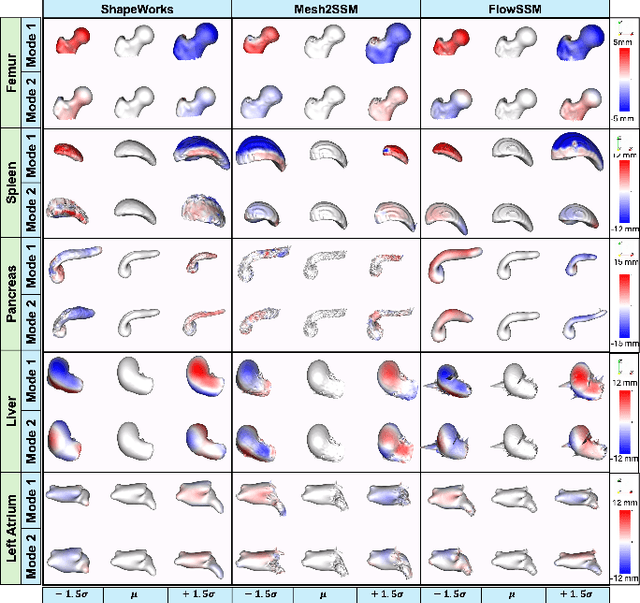
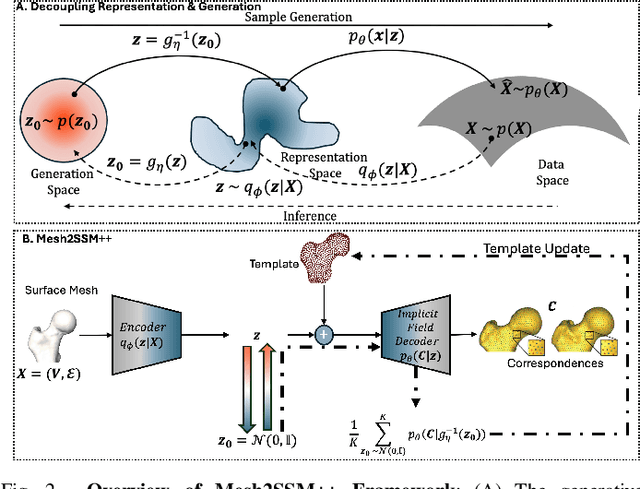
Abstract:Anatomy evaluation is crucial for understanding the physiological state, diagnosing abnormalities, and guiding medical interventions. Statistical shape modeling (SSM) is vital in this process. By enabling the extraction of quantitative morphological shape descriptors from MRI and CT scans, SSM provides comprehensive descriptions of anatomical variations within a population. However, the effectiveness of SSM in anatomy evaluation hinges on the quality and robustness of the shape models. While deep learning techniques show promise in addressing these challenges by learning complex nonlinear representations of shapes, existing models still have limitations and often require pre-established shape models for training. To overcome these issues, we propose Mesh2SSM++, a novel approach that learns to estimate correspondences from meshes in an unsupervised manner. This method leverages unsupervised, permutation-invariant representation learning to estimate how to deform a template point cloud into subject-specific meshes, forming a correspondence-based shape model. Additionally, our probabilistic formulation allows learning a population-specific template, reducing potential biases associated with template selection. A key feature of Mesh2SSM++ is its ability to quantify aleatoric uncertainty, which captures inherent data variability and is essential for ensuring reliable model predictions and robust decision-making in clinical tasks, especially under challenging imaging conditions. Through extensive validation across diverse anatomies, evaluation metrics, and downstream tasks, we demonstrate that Mesh2SSM++ outperforms existing methods. Its ability to operate directly on meshes, combined with computational efficiency and interpretability through its probabilistic framework, makes it an attractive alternative to traditional and deep learning-based SSM approaches.
MORPH-LER: Log-Euclidean Regularization for Population-Aware Image Registration
Feb 04, 2025



Abstract:Spatial transformations that capture population-level morphological statistics are critical for medical image analysis. Commonly used smoothness regularizers for image registration fail to integrate population statistics, leading to anatomically inconsistent transformations. Inverse consistency regularizers promote geometric consistency but lack population morphometrics integration. Regularizers that constrain deformation to low-dimensional manifold methods address this. However, they prioritize reconstruction over interpretability and neglect diffeomorphic properties, such as group composition and inverse consistency. We introduce MORPH-LER, a Log-Euclidean regularization framework for population-aware unsupervised image registration. MORPH-LER learns population morphometrics from spatial transformations to guide and regularize registration networks, ensuring anatomically plausible deformations. It features a bottleneck autoencoder that computes the principal logarithm of deformation fields via iterative square-root predictions. It creates a linearized latent space that respects diffeomorphic properties and enforces inverse consistency. By integrating a registration network with a diffeomorphic autoencoder, MORPH-LER produces smooth, meaningful deformation fields. The framework offers two main contributions: (1) a data-driven regularization strategy that incorporates population-level anatomical statistics to enhance transformation validity and (2) a linearized latent space that enables compact and interpretable deformation fields for efficient population morphometrics analysis. We validate MORPH-LER across two families of deep learning-based registration networks, demonstrating its ability to produce anatomically accurate, computationally efficient, and statistically meaningful transformations on the OASIS-1 brain imaging dataset.
LEDA: Log-Euclidean Diffeomorphic Autoencoder for Efficient Statistical Analysis of Diffeomorphism
Dec 20, 2024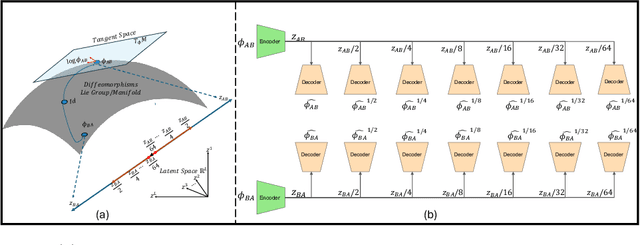
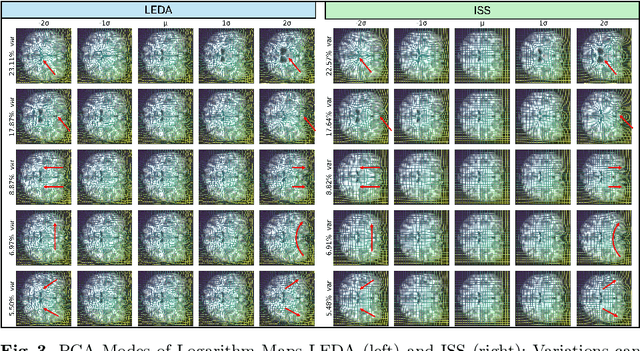
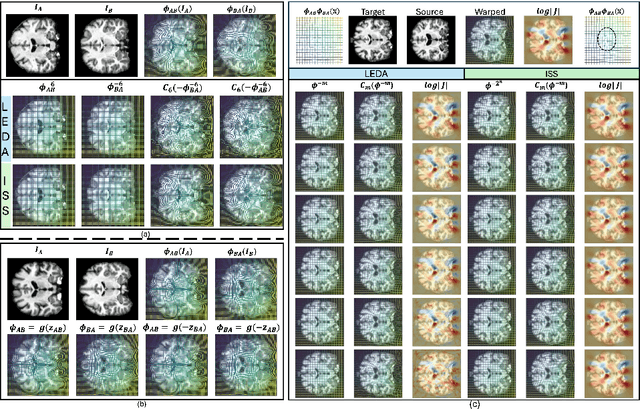

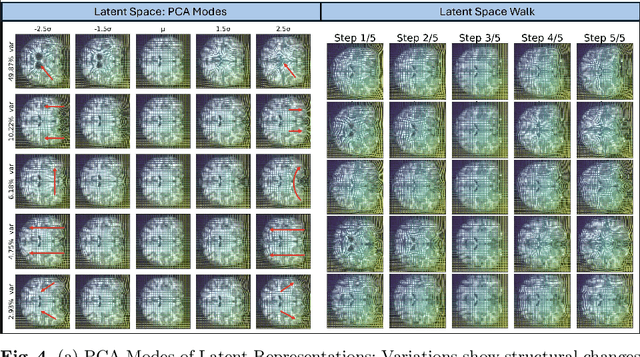
Abstract:Image registration is a core task in computational anatomy that establishes correspondences between images. Invertible deformable registration, which computes a deformation field and handles complex, non-linear transformation, is essential for tracking anatomical variations, especially in neuroimaging applications where inter-subject differences and longitudinal changes are key. Analyzing the deformation fields is challenging due to their non-linearity, limiting statistical analysis. However, traditional approaches for analyzing deformation fields are computationally expensive, sensitive to initialization, and prone to numerical errors, especially when the deformation is far from the identity. To address these limitations, we propose the Log-Euclidean Diffeomorphic Autoencoder (LEDA), an innovative framework designed to compute the principal logarithm of deformation fields by efficiently predicting consecutive square roots. LEDA operates within a linearized latent space that adheres to the diffeomorphisms group action laws, enhancing our model's robustness and applicability. We also introduce a loss function to enforce inverse consistency, ensuring accurate latent representations of deformation fields. Extensive experiments with the OASIS-1 dataset demonstrate the effectiveness of LEDA in accurately modeling and analyzing complex non-linear deformations while maintaining inverse consistency. Additionally, we evaluate its ability to capture and incorporate clinical variables, enhancing its relevance for clinical applications.
HAMIL-QA: Hierarchical Approach to Multiple Instance Learning for Atrial LGE MRI Quality Assessment
Jul 09, 2024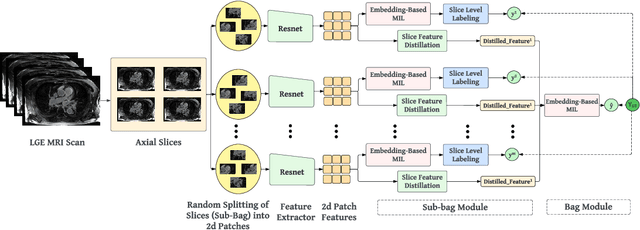
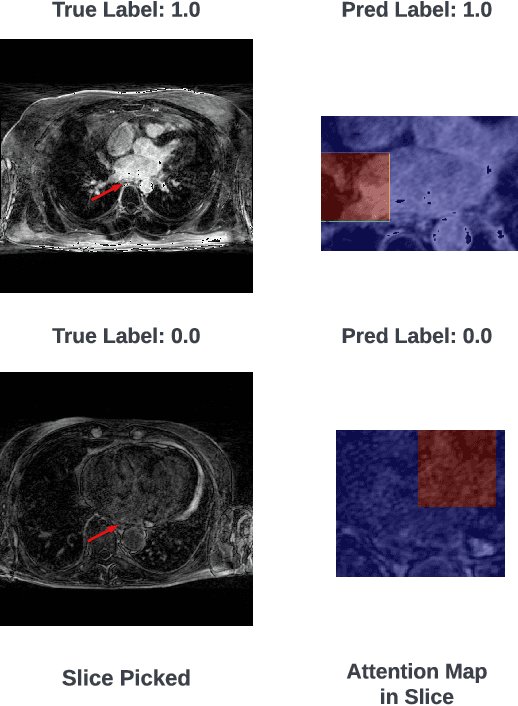
Abstract:The accurate evaluation of left atrial fibrosis via high-quality 3D Late Gadolinium Enhancement (LGE) MRI is crucial for atrial fibrillation management but is hindered by factors like patient movement and imaging variability. The pursuit of automated LGE MRI quality assessment is critical for enhancing diagnostic accuracy, standardizing evaluations, and improving patient outcomes. The deep learning models aimed at automating this process face significant challenges due to the scarcity of expert annotations, high computational costs, and the need to capture subtle diagnostic details in highly variable images. This study introduces HAMIL-QA, a multiple instance learning (MIL) framework, designed to overcome these obstacles. HAMIL-QA employs a hierarchical bag and sub-bag structure that allows for targeted analysis within sub-bags and aggregates insights at the volume level. This hierarchical MIL approach reduces reliance on extensive annotations, lessens computational load, and ensures clinically relevant quality predictions by focusing on diagnostically critical image features. Our experiments show that HAMIL-QA surpasses existing MIL methods and traditional supervised approaches in accuracy, AUROC, and F1-Score on an LGE MRI scan dataset, demonstrating its potential as a scalable solution for LGE MRI quality assessment automation. The code is available at: $\href{https://github.com/arf111/HAMIL-QA}{\text{this https URL}}$
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge