Pak-Hei Yeung
Beyond Benchmarks of IUGC: Rethinking Requirements of Deep Learning Methods for Intrapartum Ultrasound Biometry from Fetal Ultrasound Videos
Feb 13, 2026Abstract:A substantial proportion (45\%) of maternal deaths, neonatal deaths, and stillbirths occur during the intrapartum phase, with a particularly high burden in low- and middle-income countries. Intrapartum biometry plays a critical role in monitoring labor progression; however, the routine use of ultrasound in resource-limited settings is hindered by a shortage of trained sonographers. To address this challenge, the Intrapartum Ultrasound Grand Challenge (IUGC), co-hosted with MICCAI 2024, was launched. The IUGC introduces a clinically oriented multi-task automatic measurement framework that integrates standard plane classification, fetal head-pubic symphysis segmentation, and biometry, enabling algorithms to exploit complementary task information for more accurate estimation. Furthermore, the challenge releases the largest multi-center intrapartum ultrasound video dataset to date, comprising 774 videos (68,106 frames) collected from three hospitals, providing a robust foundation for model training and evaluation. In this study, we present a comprehensive overview of the challenge design, review the submissions from eight participating teams, and analyze their methods from five perspectives: preprocessing, data augmentation, learning strategy, model architecture, and post-processing. In addition, we perform a systematic analysis of the benchmark results to identify key bottlenecks, explore potential solutions, and highlight open challenges for future research. Although encouraging performance has been achieved, our findings indicate that the field remains at an early stage, and further in-depth investigation is required before large-scale clinical deployment. All benchmark solutions and the complete dataset have been publicly released to facilitate reproducible research and promote continued advances in automatic intrapartum ultrasound biometry.
FedDEAP: Adaptive Dual-Prompt Tuning for Multi-Domain Federated Learning
Oct 21, 2025Abstract:Federated learning (FL) enables multiple clients to collaboratively train machine learning models without exposing local data, balancing performance and privacy. However, domain shift and label heterogeneity across clients often hinder the generalization of the aggregated global model. Recently, large-scale vision-language models like CLIP have shown strong zero-shot classification capabilities, raising the question of how to effectively fine-tune CLIP across domains in a federated setting. In this work, we propose an adaptive federated prompt tuning framework, FedDEAP, to enhance CLIP's generalization in multi-domain scenarios. Our method includes the following three key components: (1) To mitigate the loss of domain-specific information caused by label-supervised tuning, we disentangle semantic and domain-specific features in images by using semantic and domain transformation networks with unbiased mappings; (2) To preserve domain-specific knowledge during global prompt aggregation, we introduce a dual-prompt design with a global semantic prompt and a local domain prompt to balance shared and personalized information; (3) To maximize the inclusion of semantic and domain information from images in the generated text features, we align textual and visual representations under the two learned transformations to preserve semantic and domain consistency. Theoretical analysis and extensive experiments on four datasets demonstrate the effectiveness of our method in enhancing the generalization of CLIP for federated image recognition across multiple domains.
Semi-Supervised 3D Medical Segmentation from 2D Natural Images Pretrained Model
Sep 18, 2025

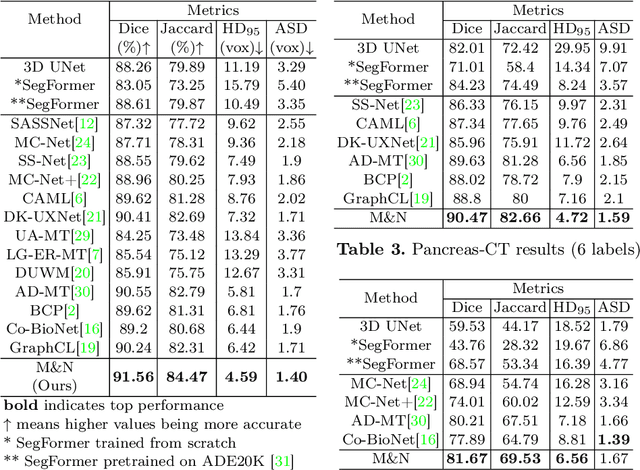

Abstract:This paper explores the transfer of knowledge from general vision models pretrained on 2D natural images to improve 3D medical image segmentation. We focus on the semi-supervised setting, where only a few labeled 3D medical images are available, along with a large set of unlabeled images. To tackle this, we propose a model-agnostic framework that progressively distills knowledge from a 2D pretrained model to a 3D segmentation model trained from scratch. Our approach, M&N, involves iterative co-training of the two models using pseudo-masks generated by each other, along with our proposed learning rate guided sampling that adaptively adjusts the proportion of labeled and unlabeled data in each training batch to align with the models' prediction accuracy and stability, minimizing the adverse effect caused by inaccurate pseudo-masks. Extensive experiments on multiple publicly available datasets demonstrate that M&N achieves state-of-the-art performance, outperforming thirteen existing semi-supervised segmentation approaches under all different settings. Importantly, ablation studies show that M&N remains model-agnostic, allowing seamless integration with different architectures. This ensures its adaptability as more advanced models emerge. The code is available at https://github.com/pakheiyeung/M-N.
Automated Fetal Biometry Assessment with Deep Ensembles using Sparse-Sampling of 2D Intrapartum Ultrasound Images
May 20, 2025



Abstract:The International Society of Ultrasound advocates Intrapartum Ultrasound (US) Imaging in Obstetrics and Gynecology (ISUOG) to monitor labour progression through changes in fetal head position. Two reliable ultrasound-derived parameters that are used to predict outcomes of instrumental vaginal delivery are the angle of progression (AoP) and head-symphysis distance (HSD). In this work, as part of the Intrapartum Ultrasounds Grand Challenge (IUGC) 2024, we propose an automated fetal biometry measurement pipeline to reduce intra- and inter-observer variability and improve measurement reliability. Our pipeline consists of three key tasks: (i) classification of standard planes (SP) from US videos, (ii) segmentation of fetal head and pubic symphysis from the detected SPs, and (iii) computation of the AoP and HSD from the segmented regions. We perform sparse sampling to mitigate class imbalances and reduce spurious correlations in task (i), and utilize ensemble-based deep learning methods for task (i) and (ii) to enhance generalizability under different US acquisition settings. Finally, to promote robustness in task iii) with respect to the structural fidelity of measurements, we retain the largest connected components and apply ellipse fitting to the segmentations. Our solution achieved ACC: 0.9452, F1: 0.9225, AUC: 0.983, MCC: 0.8361, DSC: 0.918, HD: 19.73, ASD: 5.71, $\Delta_{AoP}$: 8.90 and $\Delta_{HSD}$: 14.35 across an unseen hold-out set of 4 patients and 224 US frames. The results from the proposed automated pipeline can improve the understanding of labour arrest causes and guide the development of clinical risk stratification tools for efficient and effective prenatal care.
Bridging the Inter-Domain Gap through Low-Level Features for Cross-Modal Medical Image Segmentation
May 17, 2025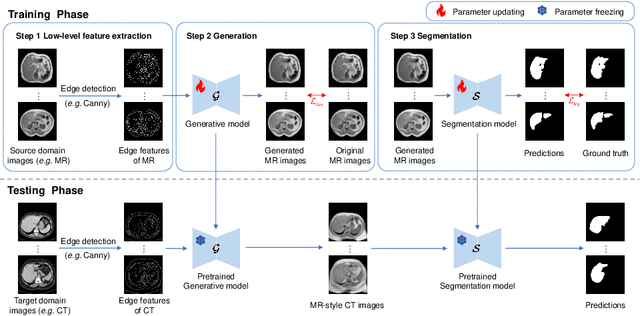
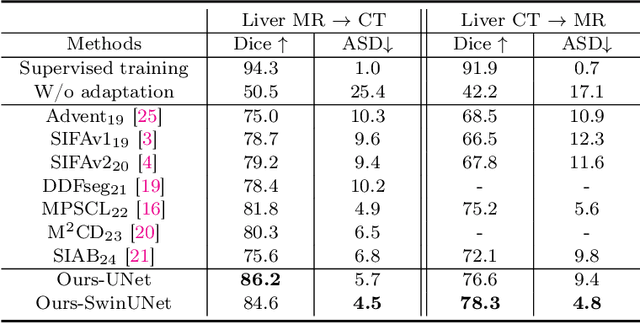
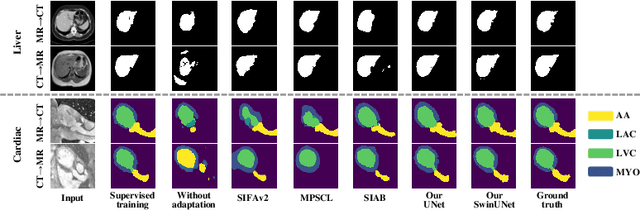
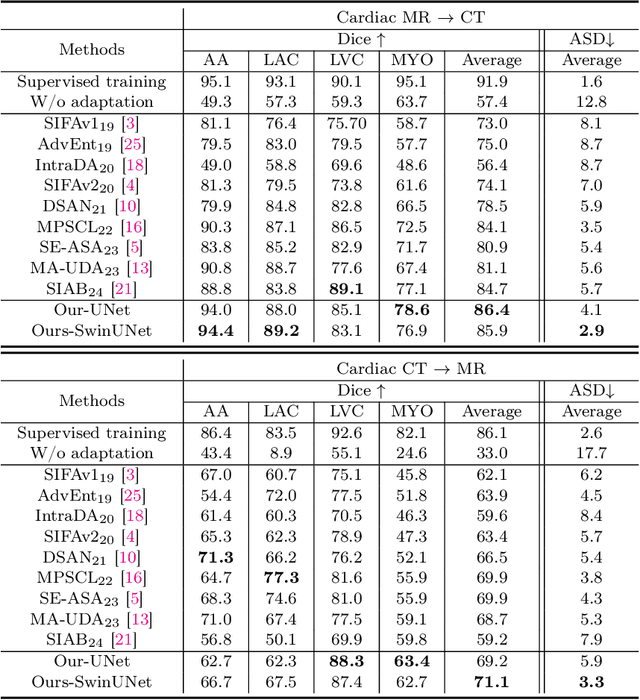
Abstract:This paper addresses the task of cross-modal medical image segmentation by exploring unsupervised domain adaptation (UDA) approaches. We propose a model-agnostic UDA framework, LowBridge, which builds on a simple observation that cross-modal images share some similar low-level features (e.g., edges) as they are depicting the same structures. Specifically, we first train a generative model to recover the source images from their edge features, followed by training a segmentation model on the generated source images, separately. At test time, edge features from the target images are input to the pretrained generative model to generate source-style target domain images, which are then segmented using the pretrained segmentation network. Despite its simplicity, extensive experiments on various publicly available datasets demonstrate that \proposed achieves state-of-the-art performance, outperforming eleven existing UDA approaches under different settings. Notably, further ablation studies show that \proposed is agnostic to different types of generative and segmentation models, suggesting its potential to be seamlessly plugged with the most advanced models to achieve even more outstanding results in the future. The code is available at https://github.com/JoshuaLPF/LowBridge.
Efficient Fourier Filtering Network with Contrastive Learning for UAV-based Unaligned Bi-modal Salient Object Detection
Nov 06, 2024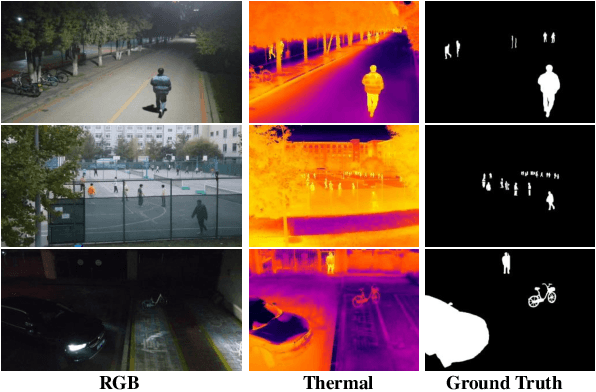
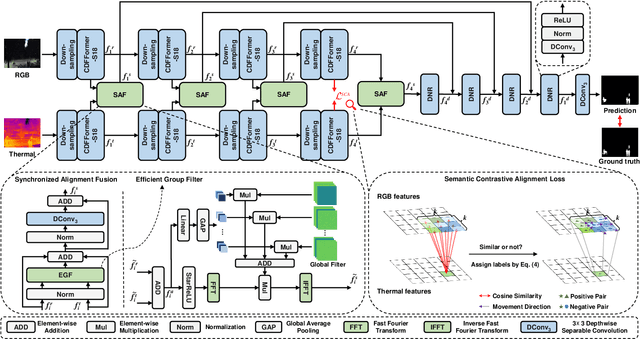
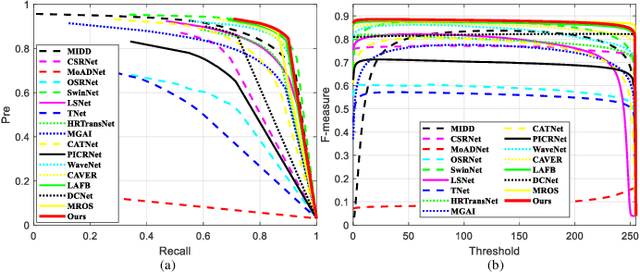

Abstract:Unmanned aerial vehicle (UAV)-based bi-modal salient object detection (BSOD) aims to segment salient objects in a scene utilizing complementary cues in unaligned RGB and thermal image pairs. However, the high computational expense of existing UAV-based BSOD models limits their applicability to real-world UAV devices. To address this problem, we propose an efficient Fourier filter network with contrastive learning that achieves both real-time and accurate performance. Specifically, we first design a semantic contrastive alignment loss to align the two modalities at the semantic level, which facilitates mutual refinement in a parameter-free way. Second, inspired by the fast Fourier transform that obtains global relevance in linear complexity, we propose synchronized alignment fusion, which aligns and fuses bi-modal features in the channel and spatial dimensions by a hierarchical filtering mechanism. Our proposed model, AlignSal, reduces the number of parameters by 70.0%, decreases the floating point operations by 49.4%, and increases the inference speed by 152.5% compared to the cutting-edge BSOD model (i.e., MROS). Extensive experiments on the UAV RGB-T 2400 and three weakly aligned datasets demonstrate that AlignSal achieves both real-time inference speed and better performance and generalizability compared to sixteen state-of-the-art BSOD models across most evaluation metrics. In addition, our ablation studies further verify AlignSal's potential in boosting the performance of existing aligned BSOD models on UAV-based unaligned data. The code is available at: https://github.com/JoshuaLPF/AlignSal.
Geometric Transformation Uncertainty for Improving 3D Fetal Brain Pose Prediction from Freehand 2D Ultrasound Videos
May 21, 2024Abstract:Accurately localizing two-dimensional (2D) ultrasound (US) fetal brain images in the 3D brain, using minimal computational resources, is an important task for automated US analysis of fetal growth and development. We propose an uncertainty-aware deep learning model for automated 3D plane localization in 2D fetal brain images. Specifically, a multi-head network is trained to jointly regress 3D plane pose from 2D images in terms of different geometric transformations. The model explicitly learns to predict uncertainty to allocate higher weight to inputs with low variances across different transformations to improve performance. Our proposed method, QAERTS, demonstrates superior pose estimation accuracy than the state-of-the-art and most of the uncertainty-based approaches, leading to 9% improvement on plane angle (PA) for localization accuracy, and 8% on normalized cross-correlation (NCC) for sampled image quality. QAERTS also demonstrates efficiency, containing 5$\times$ fewer parameters than ensemble-based approach, making it advantageous in resource-constrained settings. In addition, QAERTS proves to be more robust to noise effects observed in freehand US scanning by leveraging rotational discontinuities and explicit output uncertainties.
RapidVol: Rapid Reconstruction of 3D Ultrasound Volumes from Sensorless 2D Scans
Apr 16, 2024



Abstract:Two-dimensional (2D) freehand ultrasonography is one of the most commonly used medical imaging modalities, particularly in obstetrics and gynaecology. However, it only captures 2D cross-sectional views of inherently 3D anatomies, losing valuable contextual information. As an alternative to requiring costly and complex 3D ultrasound scanners, 3D volumes can be constructed from 2D scans using machine learning. However this usually requires long computational time. Here, we propose RapidVol: a neural representation framework to speed up slice-to-volume ultrasound reconstruction. We use tensor-rank decomposition, to decompose the typical 3D volume into sets of tri-planes, and store those instead, as well as a small neural network. A set of 2D ultrasound scans, with their ground truth (or estimated) 3D position and orientation (pose) is all that is required to form a complete 3D reconstruction. Reconstructions are formed from real fetal brain scans, and then evaluated by requesting novel cross-sectional views. When compared to prior approaches based on fully implicit representation (e.g. neural radiance fields), our method is over 3x quicker, 46% more accurate, and if given inaccurate poses is more robust. Further speed-up is also possible by reconstructing from a structural prior rather than from scratch.
Adaptive 3D Localization of 2D Freehand Ultrasound Brain Images
Sep 12, 2022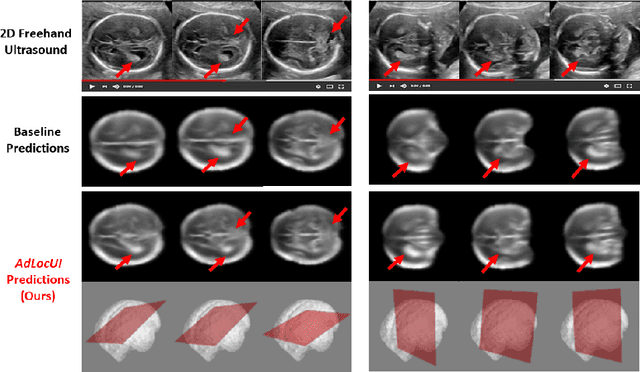
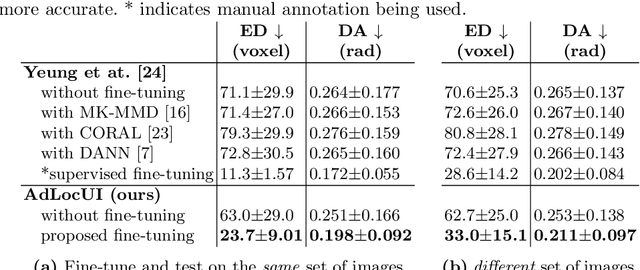
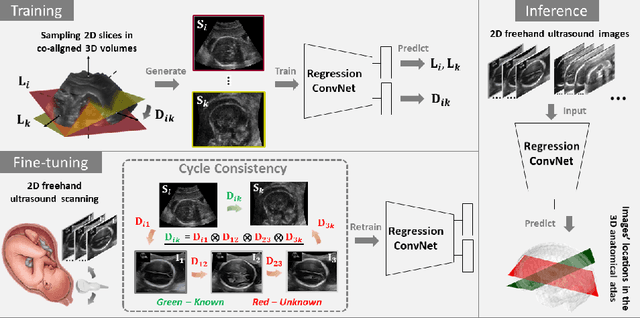
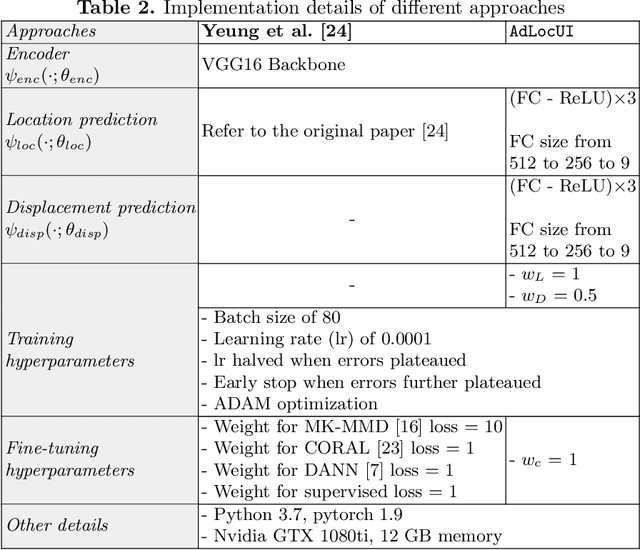
Abstract:Two-dimensional (2D) freehand ultrasound is the mainstay in prenatal care and fetal growth monitoring. The task of matching corresponding cross-sectional planes in the 3D anatomy for a given 2D ultrasound brain scan is essential in freehand scanning, but challenging. We propose AdLocUI, a framework that Adaptively Localizes 2D Ultrasound Images in the 3D anatomical atlas without using any external tracking sensor.. We first train a convolutional neural network with 2D slices sampled from co-aligned 3D ultrasound volumes to predict their locations in the 3D anatomical atlas. Next, we fine-tune it with 2D freehand ultrasound images using a novel unsupervised cycle consistency, which utilizes the fact that the overall displacement of a sequence of images in the 3D anatomical atlas is equal to the displacement from the first image to the last in that sequence. We demonstrate that AdLocUI can adapt to three different ultrasound datasets, acquired with different machines and protocols, and achieves significantly better localization accuracy than the baselines. AdLocUI can be used for sensorless 2D freehand ultrasound guidance by the bedside. The source code is available at https://github.com/pakheiyeung/AdLocUI.
ImplicitVol: Sensorless 3D Ultrasound Reconstruction with Deep Implicit Representation
Sep 24, 2021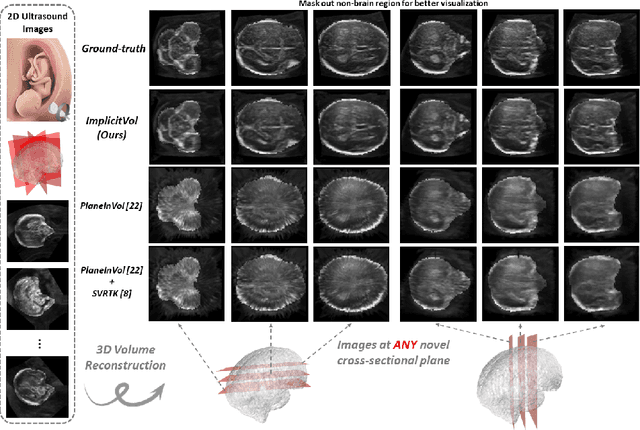
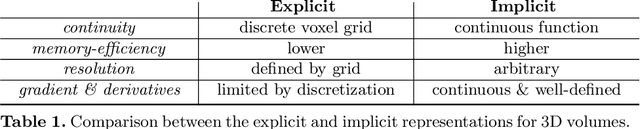

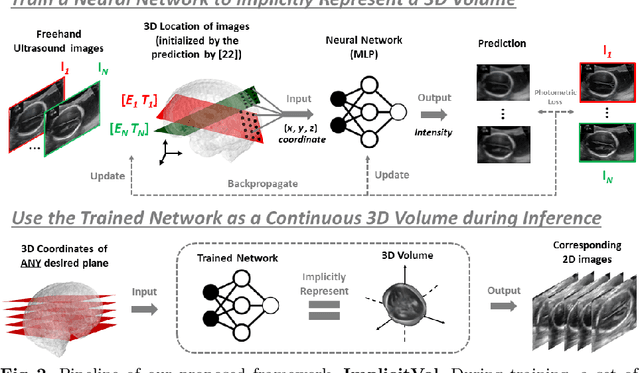
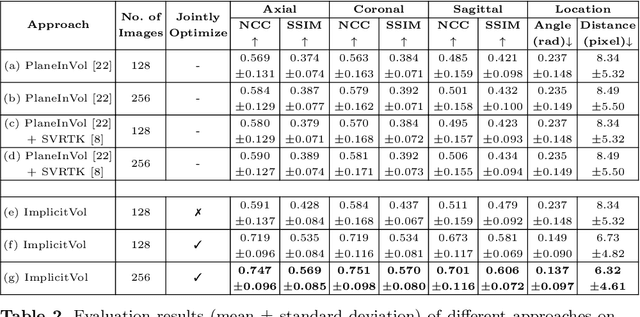
Abstract:The objective of this work is to achieve sensorless reconstruction of a 3D volume from a set of 2D freehand ultrasound images with deep implicit representation. In contrast to the conventional way that represents a 3D volume as a discrete voxel grid, we do so by parameterizing it as the zero level-set of a continuous function, i.e. implicitly representing the 3D volume as a mapping from the spatial coordinates to the corresponding intensity values. Our proposed model, termed as ImplicitVol, takes a set of 2D scans and their estimated locations in 3D as input, jointly re?fing the estimated 3D locations and learning a full reconstruction of the 3D volume. When testing on real 2D ultrasound images, novel cross-sectional views that are sampled from ImplicitVol show significantly better visual quality than those sampled from existing reconstruction approaches, outperforming them by over 30% (NCC and SSIM), between the output and ground-truth on the 3D volume testing data. The code will be made publicly available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge