Mingxin Liu
RISE-Video: Can Video Generators Decode Implicit World Rules?
Feb 05, 2026Abstract:While generative video models have achieved remarkable visual fidelity, their capacity to internalize and reason over implicit world rules remains a critical yet under-explored frontier. To bridge this gap, we present RISE-Video, a pioneering reasoning-oriented benchmark for Text-Image-to-Video (TI2V) synthesis that shifts the evaluative focus from surface-level aesthetics to deep cognitive reasoning. RISE-Video comprises 467 meticulously human-annotated samples spanning eight rigorous categories, providing a structured testbed for probing model intelligence across diverse dimensions, ranging from commonsense and spatial dynamics to specialized subject domains. Our framework introduces a multi-dimensional evaluation protocol consisting of four metrics: \textit{Reasoning Alignment}, \textit{Temporal Consistency}, \textit{Physical Rationality}, and \textit{Visual Quality}. To further support scalable evaluation, we propose an automated pipeline leveraging Large Multimodal Models (LMMs) to emulate human-centric assessment. Extensive experiments on 11 state-of-the-art TI2V models reveal pervasive deficiencies in simulating complex scenarios under implicit constraints, offering critical insights for the advancement of future world-simulating generative models.
SPWOOD: Sparse Partial Weakly-Supervised Oriented Object Detection
Feb 03, 2026Abstract:A consistent trend throughout the research of oriented object detection has been the pursuit of maintaining comparable performance with fewer and weaker annotations. This is particularly crucial in the remote sensing domain, where the dense object distribution and a wide variety of categories contribute to prohibitively high costs. Based on the supervision level, existing oriented object detection algorithms can be broadly grouped into fully supervised, semi-supervised, and weakly supervised methods. Within the scope of this work, we further categorize them to include sparsely supervised and partially weakly-supervised methods. To address the challenges of large-scale labeling, we introduce the first Sparse Partial Weakly-Supervised Oriented Object Detection framework, designed to efficiently leverage only a few sparse weakly-labeled data and plenty of unlabeled data. Our framework incorporates three key innovations: (1) We design a Sparse-annotation-Orientation-and-Scale-aware Student (SOS-Student) model to separate unlabeled objects from the background in a sparsely-labeled setting, and learn orientation and scale information from orientation-agnostic or scale-agnostic weak annotations. (2) We construct a novel Multi-level Pseudo-label Filtering strategy that leverages the distribution of model predictions, which is informed by the model's multi-layer predictions. (3) We propose a unique sparse partitioning approach, ensuring equal treatment for each category. Extensive experiments on the DOTA and DIOR datasets show that our framework achieves a significant performance gain over traditional oriented object detection methods mentioned above, offering a highly cost-effective solution. Our code is publicly available at https://github.com/VisionXLab/SPWOOD.
Partial Weakly-Supervised Oriented Object Detection
Jul 03, 2025
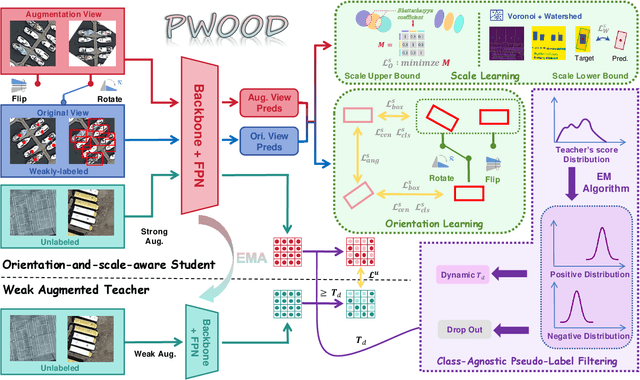


Abstract:The growing demand for oriented object detection (OOD) across various domains has driven significant research in this area. However, the high cost of dataset annotation remains a major concern. Current mainstream OOD algorithms can be mainly categorized into three types: (1) fully supervised methods using complete oriented bounding box (OBB) annotations, (2) semi-supervised methods using partial OBB annotations, and (3) weakly supervised methods using weak annotations such as horizontal boxes or points. However, these algorithms inevitably increase the cost of models in terms of annotation speed or annotation cost. To address this issue, we propose:(1) the first Partial Weakly-Supervised Oriented Object Detection (PWOOD) framework based on partially weak annotations (horizontal boxes or single points), which can efficiently leverage large amounts of unlabeled data, significantly outperforming weakly supervised algorithms trained with partially weak annotations, also offers a lower cost solution; (2) Orientation-and-Scale-aware Student (OS-Student) model capable of learning orientation and scale information with only a small amount of orientation-agnostic or scale-agnostic weak annotations; and (3) Class-Agnostic Pseudo-Label Filtering strategy (CPF) to reduce the model's sensitivity to static filtering thresholds. Comprehensive experiments on DOTA-v1.0/v1.5/v2.0 and DIOR datasets demonstrate that our PWOOD framework performs comparably to, or even surpasses, traditional semi-supervised algorithms.
Point2RBox-v2: Rethinking Point-supervised Oriented Object Detection with Spatial Layout Among Instances
Feb 07, 2025



Abstract:With the rapidly increasing demand for oriented object detection (OOD), recent research involving weakly-supervised detectors for learning OOD from point annotations has gained great attention. In this paper, we rethink this challenging task setting with the layout among instances and present Point2RBox-v2. At the core are three principles: 1) Gaussian overlap loss. It learns an upper bound for each instance by treating objects as 2D Gaussian distributions and minimizing their overlap. 2) Voronoi watershed loss. It learns a lower bound for each instance through watershed on Voronoi tessellation. 3) Consistency loss. It learns the size/rotation variation between two output sets with respect to an input image and its augmented view. Supplemented by a few devised techniques, e.g. edge loss and copy-paste, the detector is further enhanced. To our best knowledge, Point2RBox-v2 is the first approach to explore the spatial layout among instances for learning point-supervised OOD. Our solution is elegant and lightweight, yet it is expected to give a competitive performance especially in densely packed scenes: 62.61%/86.15%/34.71% on DOTA/HRSC/FAIR1M. Code is available at https://github.com/VisionXLab/point2rbox-v2.
Unleashing the Infinity Power of Geometry: A Novel Geometry-Aware Transformer for Whole Slide Histopathology Image Analysis
Feb 08, 2024


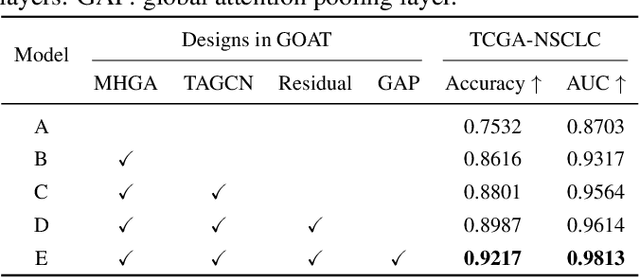
Abstract:The histopathology analysis is of great significance for the diagnosis and prognosis of cancers, however, it has great challenges due to the enormous heterogeneity of gigapixel whole slide images (WSIs) and the intricate representation of pathological features. However, recent methods have not adequately exploited geometrical representation in WSIs which is significant in disease diagnosis. Therefore, we proposed a novel weakly-supervised framework, Geometry-Aware Transformer (GOAT), in which we urge the model to pay attention to the geometric characteristics within the tumor microenvironment which often serve as potent indicators. In addition, a context-aware attention mechanism is designed to extract and enhance the morphological features within WSIs.
MGCT: Mutual-Guided Cross-Modality Transformer for Survival Outcome Prediction using Integrative Histopathology-Genomic Features
Nov 20, 2023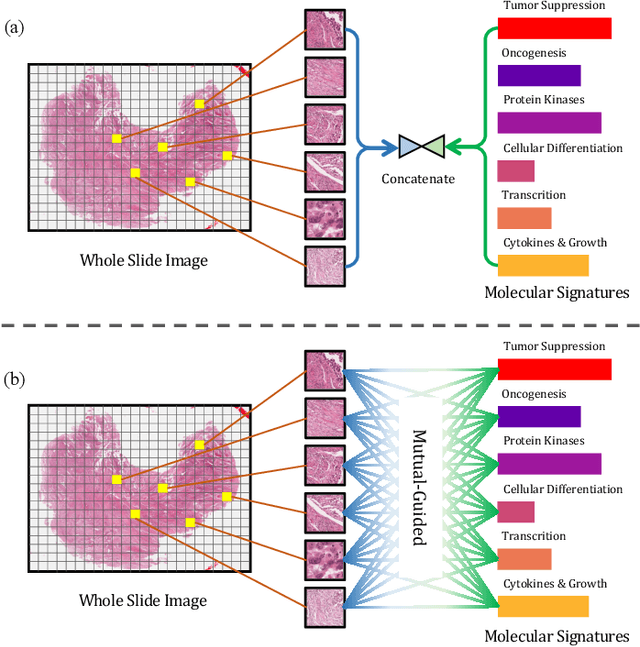
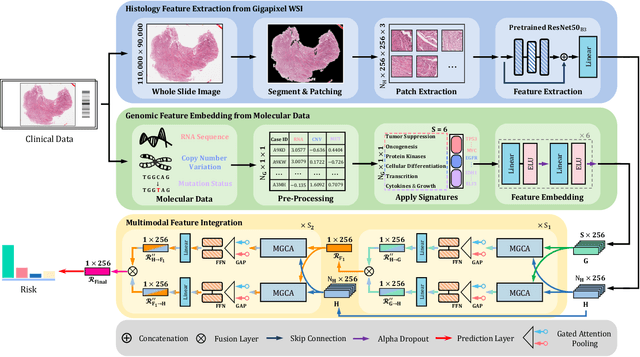
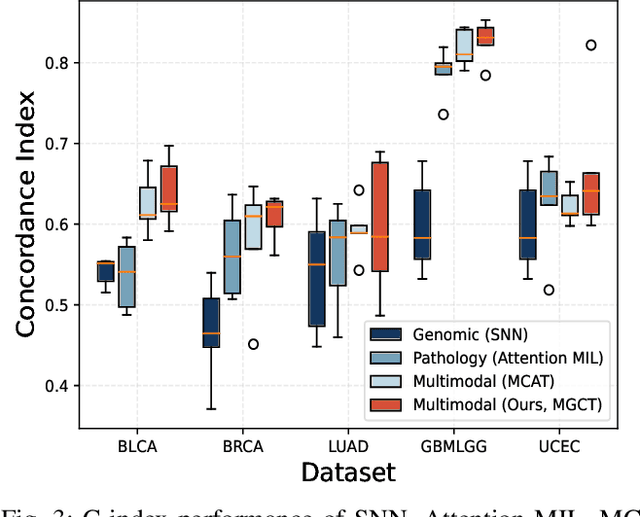
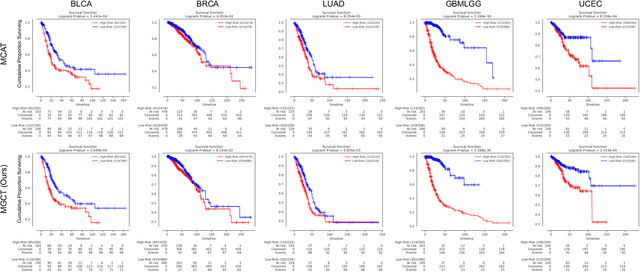
Abstract:The rapidly emerging field of deep learning-based computational pathology has shown promising results in utilizing whole slide images (WSIs) to objectively prognosticate cancer patients. However, most prognostic methods are currently limited to either histopathology or genomics alone, which inevitably reduces their potential to accurately predict patient prognosis. Whereas integrating WSIs and genomic features presents three main challenges: (1) the enormous heterogeneity of gigapixel WSIs which can reach sizes as large as 150,000x150,000 pixels; (2) the absence of a spatially corresponding relationship between histopathology images and genomic molecular data; and (3) the existing early, late, and intermediate multimodal feature fusion strategies struggle to capture the explicit interactions between WSIs and genomics. To ameliorate these issues, we propose the Mutual-Guided Cross-Modality Transformer (MGCT), a weakly-supervised, attention-based multimodal learning framework that can combine histology features and genomic features to model the genotype-phenotype interactions within the tumor microenvironment. To validate the effectiveness of MGCT, we conduct experiments using nearly 3,600 gigapixel WSIs across five different cancer types sourced from The Cancer Genome Atlas (TCGA). Extensive experimental results consistently emphasize that MGCT outperforms the state-of-the-art (SOTA) methods.
IKEA Object State Dataset: A 6DoF object pose estimation dataset and benchmark for multi-state assembly objects
Nov 16, 2021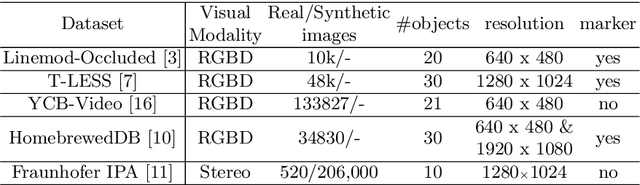
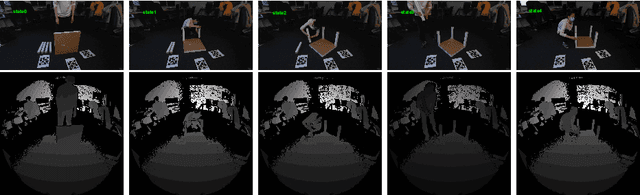


Abstract:Utilizing 6DoF(Degrees of Freedom) pose information of an object and its components is critical for object state detection tasks. We present IKEA Object State Dataset, a new dataset that contains IKEA furniture 3D models, RGBD video of the assembly process, the 6DoF pose of furniture parts and their bounding box. The proposed dataset will be available at https://github.com/mxllmx/IKEAObjectStateDataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge