Mehmet Akcakaya
Detecting and Mitigating Adversarial Attacks on Deep Learning-Based MRI Reconstruction Without Any Retraining
Jan 03, 2025



Abstract:Deep learning (DL) methods, especially those based on physics-driven DL, have become the state-of-the-art for reconstructing sub-sampled magnetic resonance imaging (MRI) data. However, studies have shown that these methods are susceptible to small adversarial input perturbations, or attacks, resulting in major distortions in the output images. Various strategies have been proposed to reduce the effects of these attacks, but they require retraining and may lower reconstruction quality for non-perturbed/clean inputs. In this work, we propose a novel approach for detecting and mitigating adversarial attacks on MRI reconstruction models without any retraining. Our detection strategy is based on the idea of cyclic measurement consistency. The output of the model is mapped to another set of MRI measurements for a different sub-sampling pattern, and this synthesized data is reconstructed with the same model. Intuitively, without an attack, the second reconstruction is expected to be consistent with the first, while with an attack, disruptions are present. Subsequently, this idea is extended to devise a novel objective function, which is minimized within a small ball around the attack input for mitigation. Experimental results show that our method substantially reduces the impact of adversarial perturbations across different datasets, attack types/strengths and PD-DL networks, and qualitatively and quantitatively outperforms conventional mitigation methods that involve retraining.
The International Workshop on Osteoarthritis Imaging Knee MRI Segmentation Challenge: A Multi-Institute Evaluation and Analysis Framework on a Standardized Dataset
May 26, 2020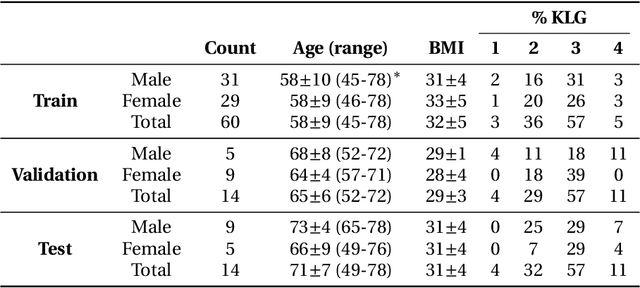
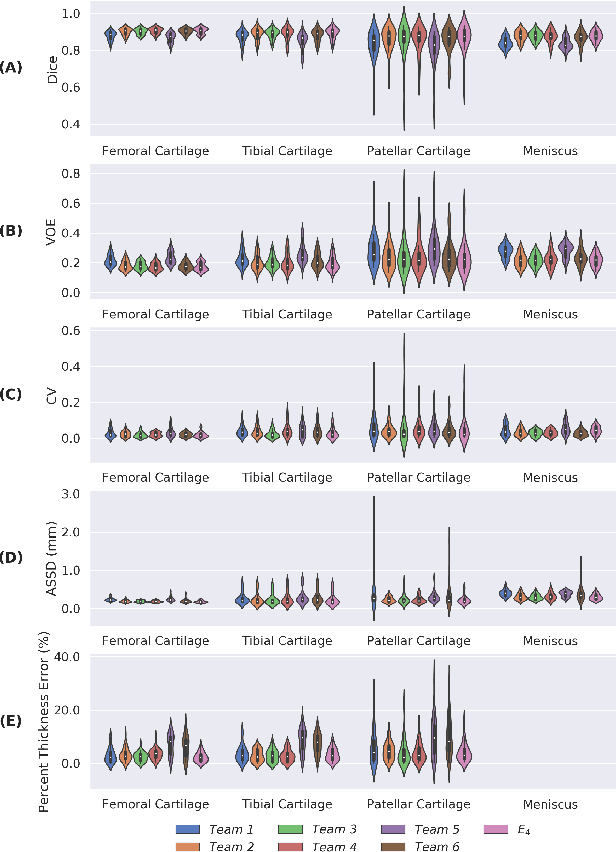
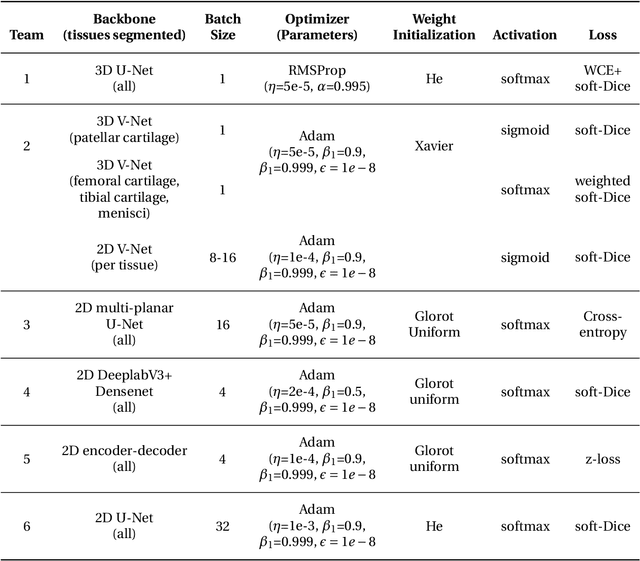
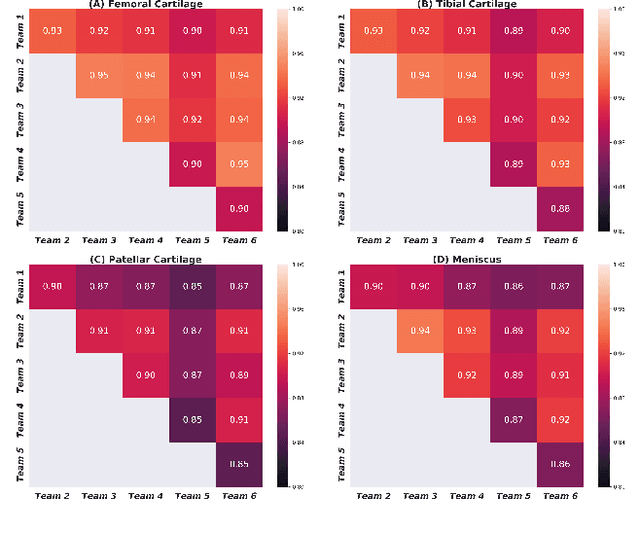
Abstract:Purpose: To organize a knee MRI segmentation challenge for characterizing the semantic and clinical efficacy of automatic segmentation methods relevant for monitoring osteoarthritis progression. Methods: A dataset partition consisting of 3D knee MRI from 88 subjects at two timepoints with ground-truth articular (femoral, tibial, patellar) cartilage and meniscus segmentations was standardized. Challenge submissions and a majority-vote ensemble were evaluated using Dice score, average symmetric surface distance, volumetric overlap error, and coefficient of variation on a hold-out test set. Similarities in network segmentations were evaluated using pairwise Dice correlations. Articular cartilage thickness was computed per-scan and longitudinally. Correlation between thickness error and segmentation metrics was measured using Pearson's coefficient. Two empirical upper bounds for ensemble performance were computed using combinations of model outputs that consolidated true positives and true negatives. Results: Six teams (T1-T6) submitted entries for the challenge. No significant differences were observed across all segmentation metrics for all tissues (p=1.0) among the four top-performing networks (T2, T3, T4, T6). Dice correlations between network pairs were high (>0.85). Per-scan thickness errors were negligible among T1-T4 (p=0.99) and longitudinal changes showed minimal bias (<0.03mm). Low correlations (<0.41) were observed between segmentation metrics and thickness error. The majority-vote ensemble was comparable to top performing networks (p=1.0). Empirical upper bound performances were similar for both combinations (p=1.0). Conclusion: Diverse networks learned to segment the knee similarly where high segmentation accuracy did not correlate to cartilage thickness accuracy. Voting ensembles did not outperform individual networks but may help regularize individual models.
Deep Learning Methods for Parallel Magnetic Resonance Image Reconstruction
Apr 01, 2019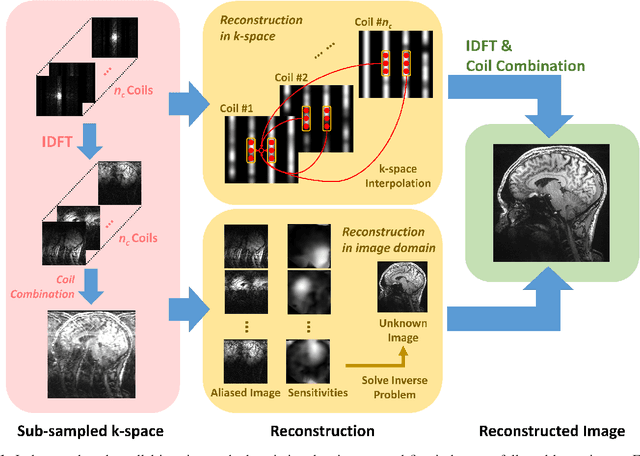
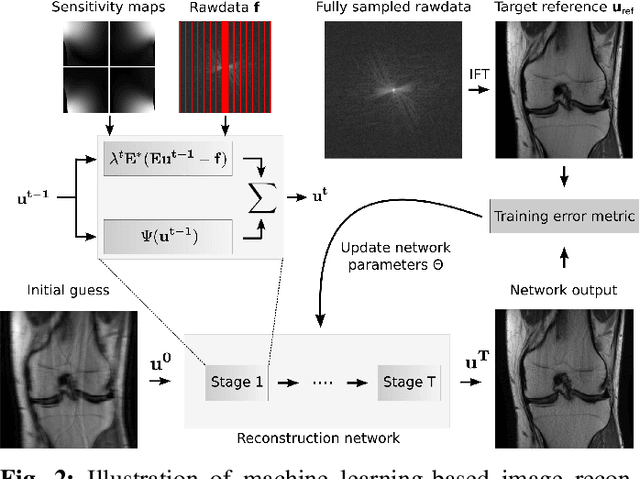
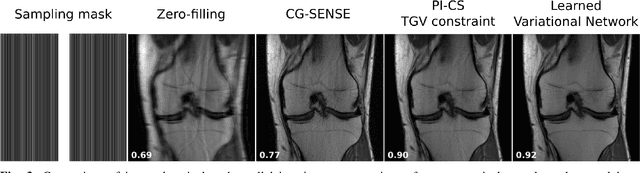

Abstract:Following the success of deep learning in a wide range of applications, neural network-based machine learning techniques have received interest as a means of accelerating magnetic resonance imaging (MRI). A number of ideas inspired by deep learning techniques from computer vision and image processing have been successfully applied to non-linear image reconstruction in the spirit of compressed sensing for both low dose computed tomography and accelerated MRI. The additional integration of multi-coil information to recover missing k-space lines in the MRI reconstruction process, is still studied less frequently, even though it is the de-facto standard for currently used accelerated MR acquisitions. This manuscript provides an overview of the recent machine learning approaches that have been proposed specifically for improving parallel imaging. A general background introduction to parallel MRI is given that is structured around the classical view of image space and k-space based methods. Both linear and non-linear methods are covered, followed by a discussion of recent efforts to further improve parallel imaging using machine learning, and specifically using artificial neural networks. Image-domain based techniques that introduce improved regularizers are covered as well as k-space based methods, where the focus is on better interpolation strategies using neural networks. Issues and open problems are discussed as well as recent efforts for producing open datasets and benchmarks for the community.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge