Martin Eklund
Department of Medical Epidemiology and Biostatistics, Karolinska Institutet, Stockholm, Sweden
AI-based Prediction of Biochemical Recurrence from Biopsy and Prostatectomy Samples
Jan 28, 2026Abstract:Biochemical recurrence (BCR) after radical prostatectomy (RP) is a surrogate marker for aggressive prostate cancer with adverse outcomes, yet current prognostic tools remain imprecise. We trained an AI-based model on diagnostic prostate biopsy slides from the STHLM3 cohort (n = 676) to predict patient-specific risk of BCR, using foundation models and attention-based multiple instance learning. Generalizability was assessed across three external RP cohorts: LEOPARD (n = 508), CHIMERA (n = 95), and TCGA-PRAD (n = 379). The image-based approach achieved 5-year time-dependent AUCs of 0.64, 0.70, and 0.70, respectively. Integrating clinical variables added complementary prognostic value and enabled statistically significant risk stratification. Compared with guideline-based CAPRA-S, AI incrementally improved postoperative prognostication. These findings suggest biopsy-trained histopathology AI can generalize across specimen types to support preoperative and postoperative decision making, but the added value of AI-based multimodal approaches over simpler predictive models should be critically scrutinized in further studies.
Artificial Intelligence-Assisted Prostate Cancer Diagnosis for Reduced Use of Immunohistochemistry
Mar 31, 2025Abstract:Prostate cancer diagnosis heavily relies on histopathological evaluation, which is subject to variability. While immunohistochemical staining (IHC) assists in distinguishing benign from malignant tissue, it involves increased work, higher costs, and diagnostic delays. Artificial intelligence (AI) presents a promising solution to reduce reliance on IHC by accurately classifying atypical glands and borderline morphologies in hematoxylin & eosin (H&E) stained tissue sections. In this study, we evaluated an AI model's ability to minimize IHC use without compromising diagnostic accuracy by retrospectively analyzing prostate core needle biopsies from routine diagnostics at three different pathology sites. These cohorts were composed exclusively of difficult cases where the diagnosing pathologists required IHC to finalize the diagnosis. The AI model demonstrated area under the curve values of 0.951-0.993 for detecting cancer in routine H&E-stained slides. Applying sensitivity-prioritized diagnostic thresholds reduced the need for IHC staining by 44.4%, 42.0%, and 20.7% in the three cohorts investigated, without a single false negative prediction. This AI model shows potential for optimizing IHC use, streamlining decision-making in prostate pathology, and alleviating resource burdens.
Foundation Models -- A Panacea for Artificial Intelligence in Pathology?
Feb 28, 2025
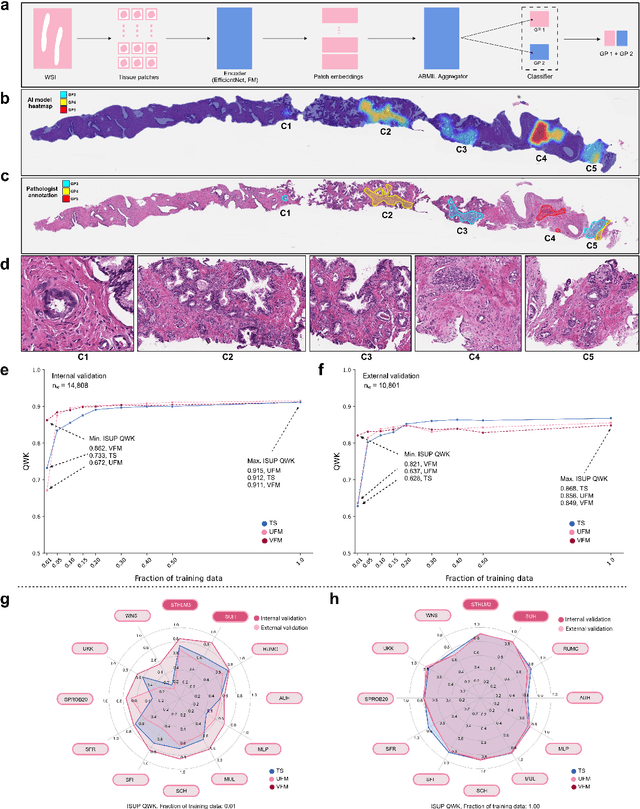

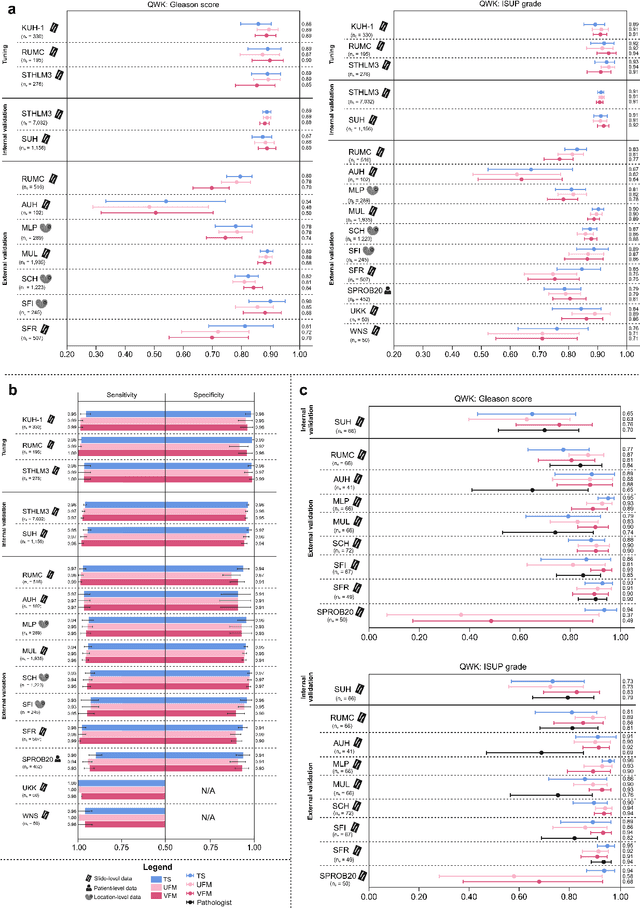
Abstract:The role of artificial intelligence (AI) in pathology has evolved from aiding diagnostics to uncovering predictive morphological patterns in whole slide images (WSIs). Recently, foundation models (FMs) leveraging self-supervised pre-training have been widely advocated as a universal solution for diverse downstream tasks. However, open questions remain about their clinical applicability and generalization advantages over end-to-end learning using task-specific (TS) models. Here, we focused on AI with clinical-grade performance for prostate cancer diagnosis and Gleason grading. We present the largest validation of AI for this task, using over 100,000 core needle biopsies from 7,342 patients across 15 sites in 11 countries. We compared two FMs with a fully end-to-end TS model in a multiple instance learning framework. Our findings challenge assumptions that FMs universally outperform TS models. While FMs demonstrated utility in data-scarce scenarios, their performance converged with - and was in some cases surpassed by - TS models when sufficient labeled training data were available. Notably, extensive task-specific training markedly reduced clinically significant misgrading, misdiagnosis of challenging morphologies, and variability across different WSI scanners. Additionally, FMs used up to 35 times more energy than the TS model, raising concerns about their sustainability. Our results underscore that while FMs offer clear advantages for rapid prototyping and research, their role as a universal solution for clinically applicable medical AI remains uncertain. For high-stakes clinical applications, rigorous validation and consideration of task-specific training remain critically important. We advocate for integrating the strengths of FMs and end-to-end learning to achieve robust and resource-efficient AI pathology solutions fit for clinical use.
Reconsidering evaluation practices in modular systems: On the propagation of errors in MRI prostate cancer detection
Sep 15, 2023Abstract:Magnetic resonance imaging has evolved as a key component for prostate cancer (PCa) detection, substantially increasing the radiologist workload. Artificial intelligence (AI) systems can support radiological assessment by segmenting and classifying lesions in clinically significant (csPCa) and non-clinically significant (ncsPCa). Commonly, AI systems for PCa detection involve an automatic prostate segmentation followed by the lesion detection using the extracted prostate. However, evaluation reports are typically presented in terms of detection under the assumption of the availability of a highly accurate segmentation and an idealistic scenario, omitting the propagation of errors between modules. For that purpose, we evaluate the effect of two different segmentation networks (s1 and s2) with heterogeneous performances in the detection stage and compare it with an idealistic setting (s1:89.90+-2.23 vs 88.97+-3.06 ncsPCa, P<.001, 89.30+-4.07 and 88.12+-2.71 csPCa, P<.001). Our results depict the relevance of a holistic evaluation, accounting for all the sub-modules involved in the system.
Leveraging multi-view data without annotations for prostate MRI segmentation: A contrastive approach
Aug 12, 2023

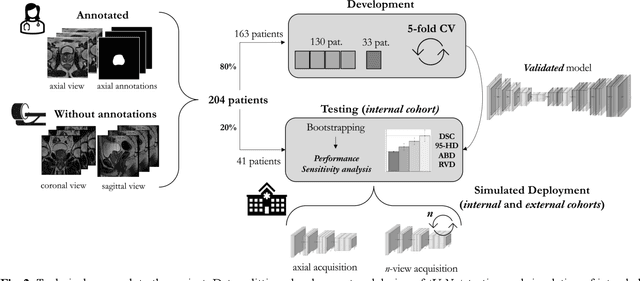

Abstract:An accurate prostate delineation and volume characterization can support the clinical assessment of prostate cancer. A large amount of automatic prostate segmentation tools consider exclusively the axial MRI direction in spite of the availability as per acquisition protocols of multi-view data. Further, when multi-view data is exploited, manual annotations and availability at test time for all the views is commonly assumed. In this work, we explore a contrastive approach at training time to leverage multi-view data without annotations and provide flexibility at deployment time in the event of missing views. We propose a triplet encoder and single decoder network based on U-Net, tU-Net (triplet U-Net). Our proposed architecture is able to exploit non-annotated sagittal and coronal views via contrastive learning to improve the segmentation from a volumetric perspective. For that purpose, we introduce the concept of inter-view similarity in the latent space. To guide the training, we combine a dice score loss calculated with respect to the axial view and its manual annotations together with a multi-view contrastive loss. tU-Net shows statistical improvement in dice score coefficient (DSC) with respect to only axial view (91.25+-0.52% compared to 86.40+-1.50%,P<.001). Sensitivity analysis reveals the volumetric positive impact of the contrastive loss when paired with tU-Net (2.85+-1.34% compared to 3.81+-1.88%,P<.001). Further, our approach shows good external volumetric generalization in an in-house dataset when tested with multi-view data (2.76+-1.89% compared to 3.92+-3.31%,P=.002), showing the feasibility of exploiting non-annotated multi-view data through contrastive learning whilst providing flexibility at deployment in the event of missing views.
Prostate Age Gap : An MRI surrogate marker of aging for prostate cancer detection
Aug 10, 2023Abstract:Background: Prostate cancer (PC) MRI-based risk calculators are commonly based on biological (e.g. PSA), MRI markers (e.g. volume), and patient age. Whilst patient age measures the amount of years an individual has existed, biological age (BA) might better reflect the physiology of an individual. However, surrogates from prostate MRI and linkage with clinically significant PC (csPC) remain to be explored. Purpose: To obtain and evaluate Prostate Age Gap (PAG) as an MRI marker tool for csPC risk. Study type: Retrospective. Population: A total of 7243 prostate MRI slices from 468 participants who had undergone prostate biopsies. A deep learning model was trained on 3223 MRI slices cropped around the gland from 81 low-grade PC (ncsPC, Gleason score <=6) and 131 negative cases and tested on the remaining 256 participants. Assessment: Chronological age was defined as the age of the participant at the time of the visit and used to train the deep learning model to predict the age of the patient. Following, we obtained PAG, defined as the model predicted age minus the patient's chronological age. Multivariate logistic regression models were used to estimate the association through odds ratio (OR) and predictive value of PAG and compared against PSA levels and PI-RADS>=3. Statistical tests: T-test, Mann-Whitney U test, Permutation test and ROC curve analysis. Results: The multivariate adjusted model showed a significant difference in the odds of clinically significant PC (csPC, Gleason score >=7) (OR =3.78, 95% confidence interval (CI):2.32-6.16, P <.001). PAG showed a better predictive ability when compared to PI-RADS>=3 and adjusted by other risk factors, including PSA levels: AUC =0.981 vs AUC =0.704, p<.001. Conclusion: PAG was significantly associated with the risk of clinically significant PC and outperformed other well-established PC risk factors.
Physical Color Calibration of Digital Pathology Scanners for Robust Artificial Intelligence Assisted Cancer Diagnosis
Jul 07, 2023



Abstract:The potential of artificial intelligence (AI) in digital pathology is limited by technical inconsistencies in the production of whole slide images (WSIs), leading to degraded AI performance and posing a challenge for widespread clinical application as fine-tuning algorithms for each new site is impractical. Changes in the imaging workflow can also lead to compromised diagnoses and patient safety risks. We evaluated whether physical color calibration of scanners can standardize WSI appearance and enable robust AI performance. We employed a color calibration slide in four different laboratories and evaluated its impact on the performance of an AI system for prostate cancer diagnosis on 1,161 WSIs. Color standardization resulted in consistently improved AI model calibration and significant improvements in Gleason grading performance. The study demonstrates that physical color calibration provides a potential solution to the variation introduced by different scanners, making AI-based cancer diagnostics more reliable and applicable in clinical settings.
Using deep learning to detect patients at risk for prostate cancer despite benign biopsies
Jul 31, 2021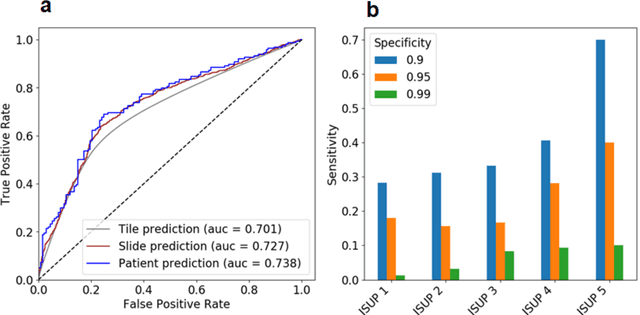

Abstract:Background: Transrectal ultrasound guided systematic biopsies of the prostate is a routine procedure to establish a prostate cancer diagnosis. However, the 10-12 prostate core biopsies only sample a relatively small volume of the prostate, and tumour lesions in regions between biopsy cores can be missed, leading to a well-known low sensitivity to detect clinically relevant cancer. As a proof-of-principle, we developed and validated a deep convolutional neural network model to distinguish between morphological patterns in benign prostate biopsy whole slide images from men with and without established cancer. Methods: This study included 14,354 hematoxylin and eosin stained whole slide images from benign prostate biopsies from 1,508 men in two groups: men without an established prostate cancer (PCa) diagnosis and men with at least one core biopsy diagnosed with PCa. 80% of the participants were assigned as training data and used for model optimization (1,211 men), and the remaining 20% (297 men) as a held-out test set used to evaluate model performance. An ensemble of 10 deep convolutional neural network models was optimized for classification of biopsies from men with and without established cancer. Hyperparameter optimization and model selection was performed by cross-validation in the training data . Results: Area under the receiver operating characteristic curve (ROC-AUC) was estimated as 0.727 (bootstrap 95% CI: 0.708-0.745) on biopsy level and 0.738 (bootstrap 95% CI: 0.682 - 0.796) on man level. At a specificity of 0.9 the model had an estimated sensitivity of 0.348. Conclusion: The developed model has the ability to detect men with risk of missed PCa due to under-sampling of the prostate. The proposed model has the potential to reduce the number of false negative cases in routine systematic prostate biopsies and to indicate men who could benefit from MRI-guided re-biopsy.
Transcriptome-wide prediction of prostate cancer gene expression from histopathology images using co-expression based convolutional neural networks
Apr 19, 2021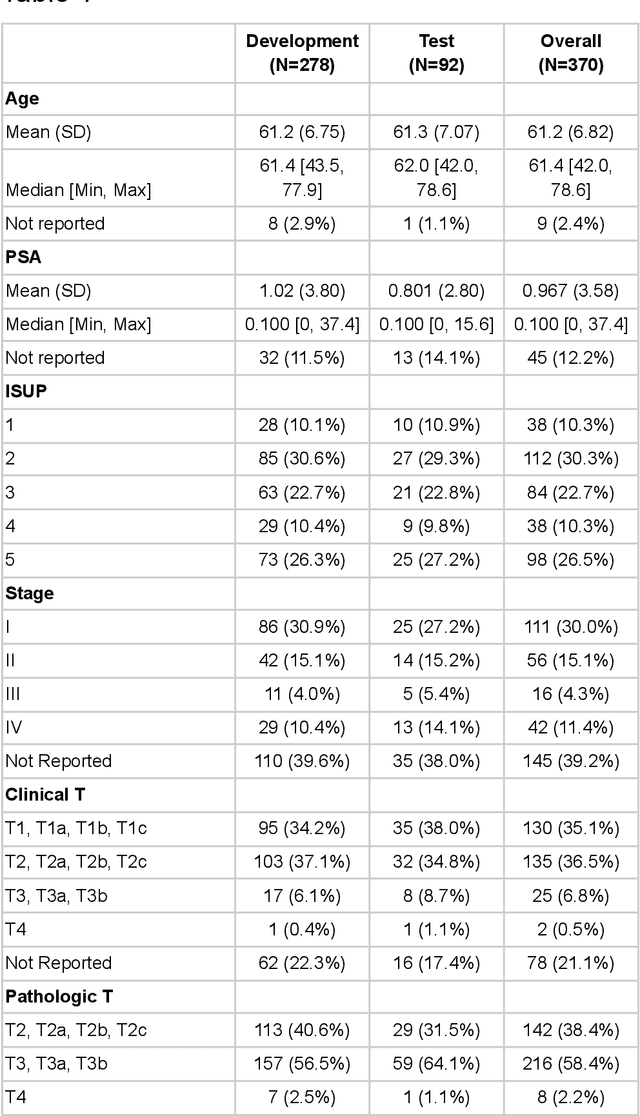
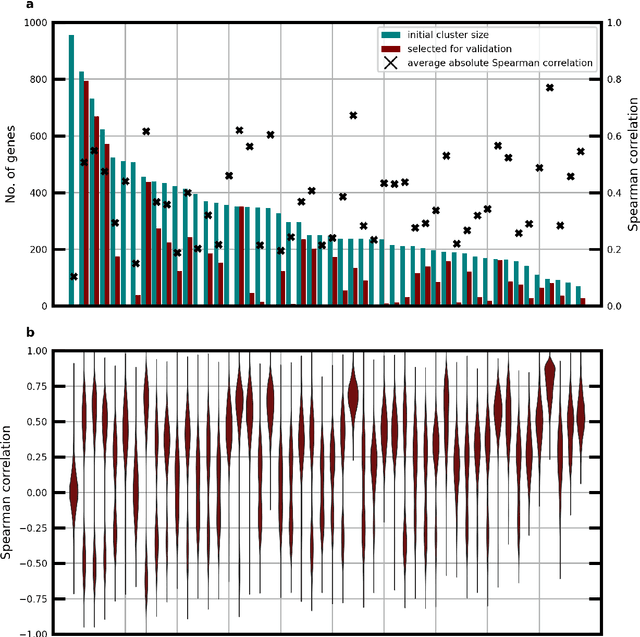
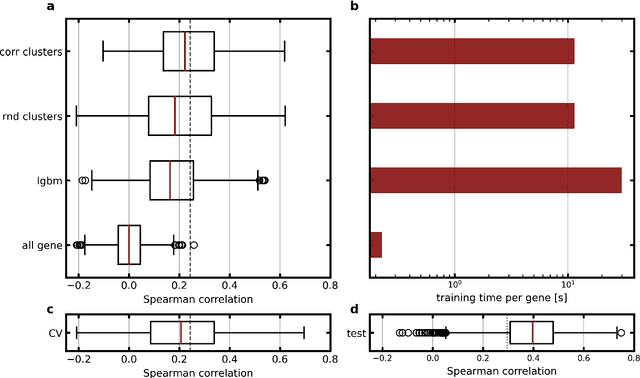
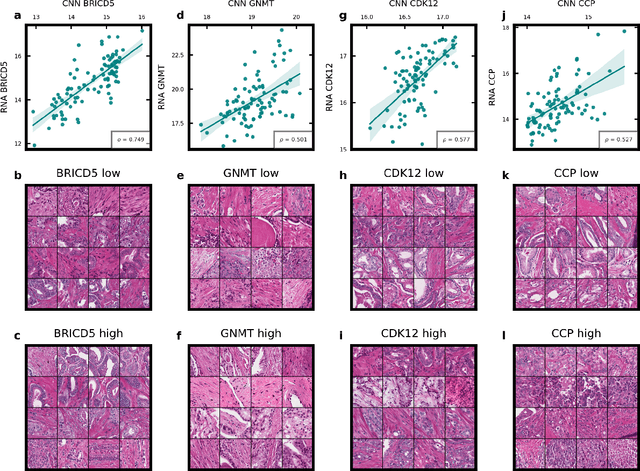
Abstract:Molecular phenotyping by gene expression profiling is common in contemporary cancer research and in molecular diagnostics. However, molecular profiling remains costly and resource intense to implement, and is just starting to be introduced into clinical diagnostics. Molecular changes, including genetic alterations and gene expression changes, occuring in tumors cause morphological changes in tissue, which can be observed on the microscopic level. The relationship between morphological patterns and some of the molecular phenotypes can be exploited to predict molecular phenotypes directly from routine haematoxylin and eosin (H&E) stained whole slide images (WSIs) using deep convolutional neural networks (CNNs). In this study, we propose a new, computationally efficient approach for disease specific modelling of relationships between morphology and gene expression, and we conducted the first transcriptome-wide analysis in prostate cancer, using CNNs to predict bulk RNA-sequencing estimates from WSIs of H&E stained tissue. The work is based on the TCGA PRAD study and includes both WSIs and RNA-seq data for 370 patients. Out of 15586 protein coding and sufficiently frequently expressed transcripts, 6618 had predicted expression significantly associated with RNA-seq estimates (FDR-adjusted p-value < 1*10-4) in a cross-validation. 5419 (81.9%) of these were subsequently validated in a held-out test set. We also demonstrate the ability to predict a prostate cancer specific cell cycle progression score directly from WSIs. These findings suggest that contemporary computer vision models offer an inexpensive and scalable solution for prediction of gene expression phenotypes directly from WSIs, providing opportunity for cost-effective large-scale research studies and molecular diagnostics.
Detection of Perineural Invasion in Prostate Needle Biopsies with Deep Neural Networks
Apr 03, 2020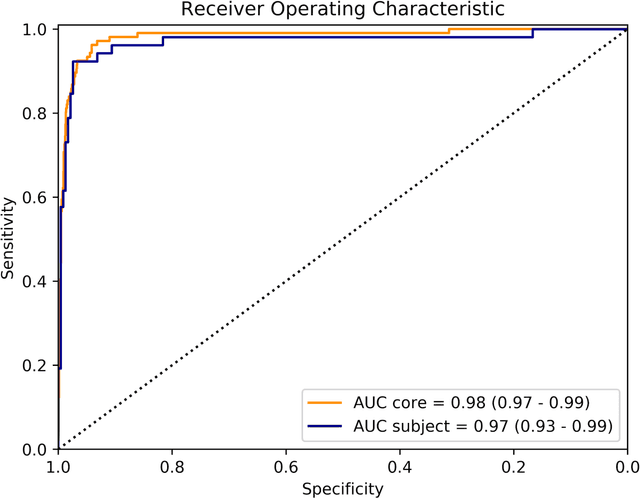
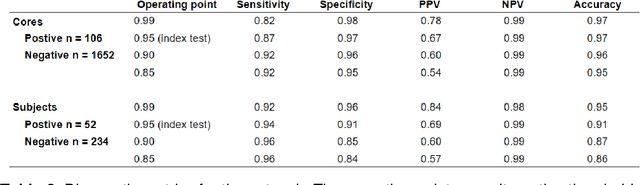
Abstract:Background: The detection of perineural invasion (PNI) by carcinoma in prostate biopsies has been shown to be associated with poor prognosis. The assessment and quantification of PNI is; however, labor intensive. In the study we aimed to develop an algorithm based on deep neural networks to aid pathologists in this task. Methods: We collected, digitized and pixel-wise annotated the PNI findings in each of the approximately 80,000 biopsy cores from the 7,406 men who underwent biopsy in the prospective and diagnostic STHLM3 trial between 2012 and 2014. In total, 485 biopsy cores showed PNI. We also digitized more than 10% (n=8,318) of the PNI negative biopsy cores. Digitized biopsies from a random selection of 80% of the men were used to build deep neural networks, and the remaining 20% were used to evaluate the performance of the algorithm. Results: For the detection of PNI in prostate biopsy cores the network had an estimated area under the receiver operating characteristics curve of 0.98 (95% CI 0.97-0.99) based on 106 PNI positive cores and 1,652 PNI negative cores in the independent test set. For the pre-specified operating point this translates to sensitivity of 0.87 and specificity of 0.97. The corresponding positive and negative predictive values were 0.67 and 0.99, respectively. For localizing the regions of PNI within a slide we estimated an average intersection over union of 0.50 (CI: 0.46-0.55). Conclusion: We have developed an algorithm based on deep neural networks for detecting PNI in prostate biopsies with apparently acceptable diagnostic properties. These algorithms have the potential to aid pathologists in the day-to-day work by drastically reducing the number of biopsy cores that need to be assessed for PNI and by highlighting regions of diagnostic interest.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge