Umair Khan
Staining normalization in histopathology: Method benchmarking using multicenter dataset
Jun 23, 2025Abstract:Hematoxylin and Eosin (H&E) has been the gold standard in tissue analysis for decades, however, tissue specimens stained in different laboratories vary, often significantly, in appearance. This variation poses a challenge for both pathologists' and AI-based downstream analysis. Minimizing stain variation computationally is an active area of research. To further investigate this problem, we collected a unique multi-center tissue image dataset, wherein tissue samples from colon, kidney, and skin tissue blocks were distributed to 66 different labs for routine H&E staining. To isolate staining variation, other factors affecting the tissue appearance were kept constant. Further, we used this tissue image dataset to compare the performance of eight different stain normalization methods, including four traditional methods, namely, histogram matching, Macenko, Vahadane, and Reinhard normalization, and two deep learning-based methods namely CycleGAN and Pixp2pix, both with two variants each. We used both quantitative and qualitative evaluation to assess the performance of these methods. The dataset's inter-laboratory staining variation could also guide strategies to improve model generalizability through varied training data
Physical Color Calibration of Digital Pathology Scanners for Robust Artificial Intelligence Assisted Cancer Diagnosis
Jul 07, 2023



Abstract:The potential of artificial intelligence (AI) in digital pathology is limited by technical inconsistencies in the production of whole slide images (WSIs), leading to degraded AI performance and posing a challenge for widespread clinical application as fine-tuning algorithms for each new site is impractical. Changes in the imaging workflow can also lead to compromised diagnoses and patient safety risks. We evaluated whether physical color calibration of scanners can standardize WSI appearance and enable robust AI performance. We employed a color calibration slide in four different laboratories and evaluated its impact on the performance of an AI system for prostate cancer diagnosis on 1,161 WSIs. Color standardization resulted in consistently improved AI model calibration and significant improvements in Gleason grading performance. The study demonstrates that physical color calibration provides a potential solution to the variation introduced by different scanners, making AI-based cancer diagnostics more reliable and applicable in clinical settings.
A single-cell gene expression language model
Oct 25, 2022Abstract:Gene regulation is a dynamic process that connects genotype and phenotype. Given the difficulty of physically mapping mammalian gene circuitry, we require new computational methods to learn regulatory rules. Natural language is a valuable analogy to the communication of regulatory control. Machine learning systems model natural language by explicitly learning context dependencies between words. We propose a similar system applied to single-cell RNA expression profiles to learn context dependencies between genes. Our model, Exceiver, is trained across a diversity of cell types using a self-supervised task formulated for discrete count data, accounting for feature sparsity. We found agreement between the similarity profiles of latent sample representations and learned gene embeddings with respect to biological annotations. We evaluated Exceiver on a new dataset and a downstream prediction task and found that pretraining supports transfer learning. Our work provides a framework to model gene regulation on a single-cell level and transfer knowledge to downstream tasks.
Deformation equivariant cross-modality image synthesis with paired non-aligned training data
Aug 26, 2022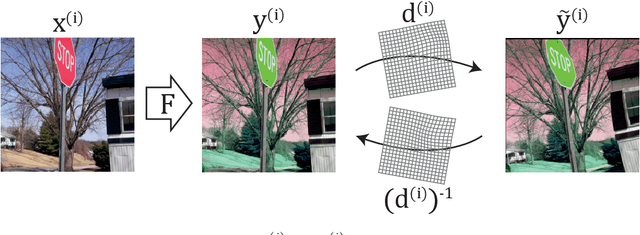



Abstract:Cross-modality image synthesis is an active research topic with multiple medical clinically relevant applications. Recently, methods allowing training with paired but misaligned data have started to emerge. However, no robust and well-performing methods applicable to a wide range of real world data sets exist. In this work, we propose a generic solution to the problem of cross-modality image synthesis with paired but non-aligned data by introducing new deformation equivariance encouraging loss functions. The method consists of joint training of an image synthesis network together with separate registration networks and allows adversarial training conditioned on the input even with misaligned data. The work lowers the bar for new clinical applications by allowing effortless training of cross-modality image synthesis networks for more difficult data sets and opens up opportunities for the development of new generic learning based cross-modality registration algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge