Toyonori Tsuzuki
Department of Surgical Pathology, School of Medicine, Aichi Medical University, Nagoya, Japan
Foundation Models -- A Panacea for Artificial Intelligence in Pathology?
Feb 28, 2025
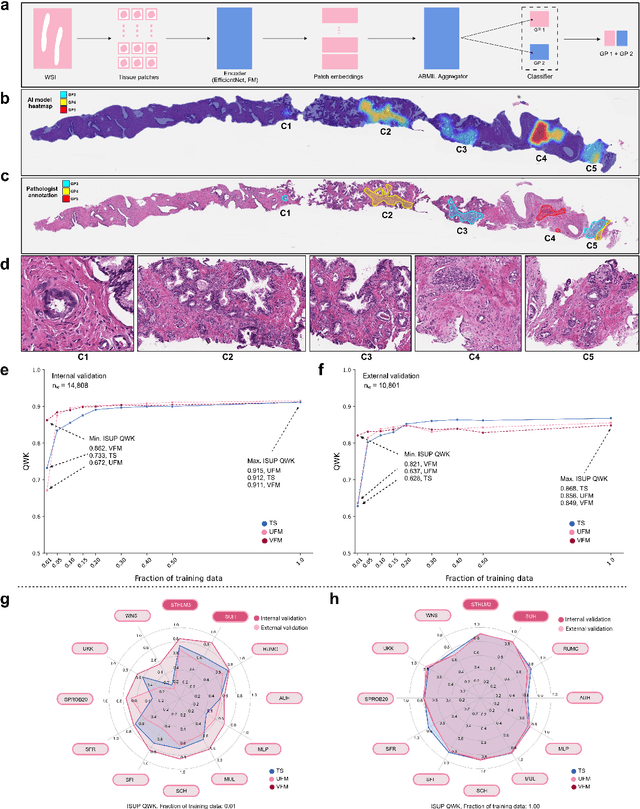

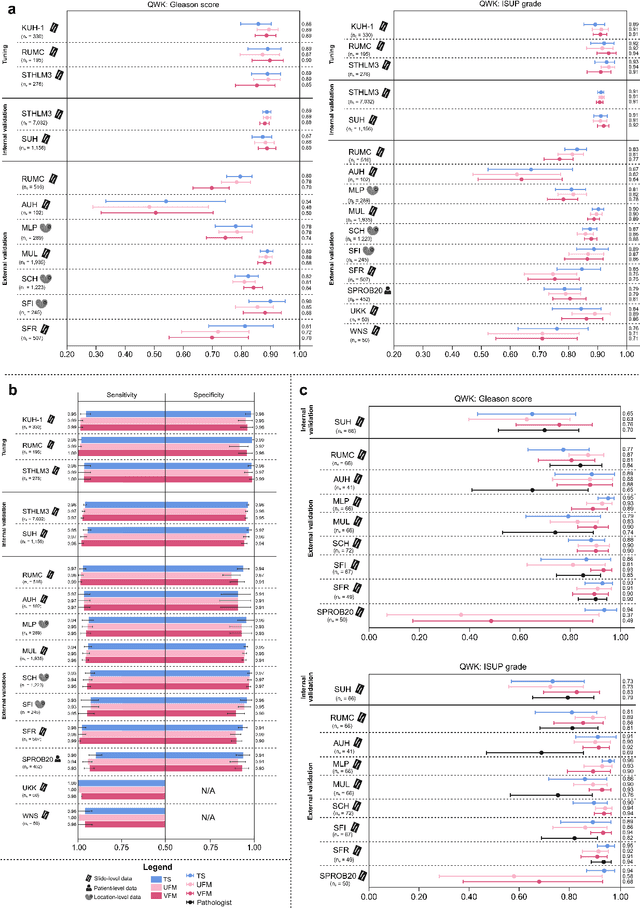
Abstract:The role of artificial intelligence (AI) in pathology has evolved from aiding diagnostics to uncovering predictive morphological patterns in whole slide images (WSIs). Recently, foundation models (FMs) leveraging self-supervised pre-training have been widely advocated as a universal solution for diverse downstream tasks. However, open questions remain about their clinical applicability and generalization advantages over end-to-end learning using task-specific (TS) models. Here, we focused on AI with clinical-grade performance for prostate cancer diagnosis and Gleason grading. We present the largest validation of AI for this task, using over 100,000 core needle biopsies from 7,342 patients across 15 sites in 11 countries. We compared two FMs with a fully end-to-end TS model in a multiple instance learning framework. Our findings challenge assumptions that FMs universally outperform TS models. While FMs demonstrated utility in data-scarce scenarios, their performance converged with - and was in some cases surpassed by - TS models when sufficient labeled training data were available. Notably, extensive task-specific training markedly reduced clinically significant misgrading, misdiagnosis of challenging morphologies, and variability across different WSI scanners. Additionally, FMs used up to 35 times more energy than the TS model, raising concerns about their sustainability. Our results underscore that while FMs offer clear advantages for rapid prototyping and research, their role as a universal solution for clinically applicable medical AI remains uncertain. For high-stakes clinical applications, rigorous validation and consideration of task-specific training remain critically important. We advocate for integrating the strengths of FMs and end-to-end learning to achieve robust and resource-efficient AI pathology solutions fit for clinical use.
Detection of Perineural Invasion in Prostate Needle Biopsies with Deep Neural Networks
Apr 03, 2020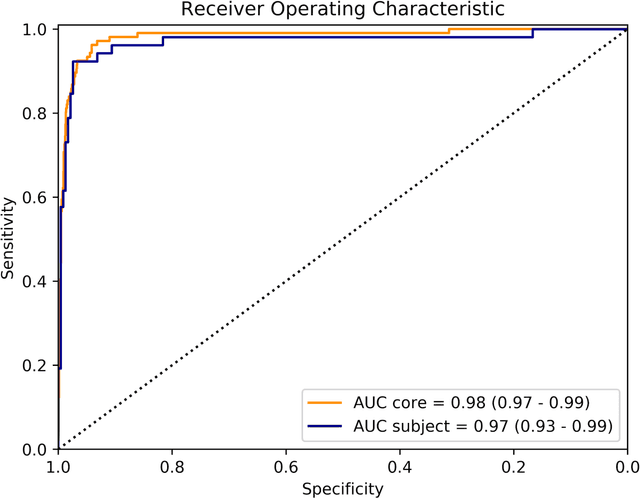
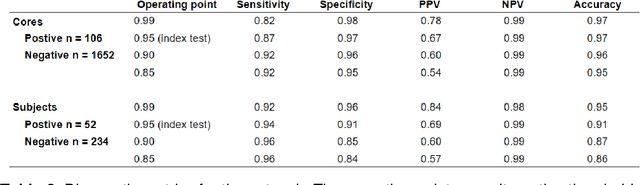
Abstract:Background: The detection of perineural invasion (PNI) by carcinoma in prostate biopsies has been shown to be associated with poor prognosis. The assessment and quantification of PNI is; however, labor intensive. In the study we aimed to develop an algorithm based on deep neural networks to aid pathologists in this task. Methods: We collected, digitized and pixel-wise annotated the PNI findings in each of the approximately 80,000 biopsy cores from the 7,406 men who underwent biopsy in the prospective and diagnostic STHLM3 trial between 2012 and 2014. In total, 485 biopsy cores showed PNI. We also digitized more than 10% (n=8,318) of the PNI negative biopsy cores. Digitized biopsies from a random selection of 80% of the men were used to build deep neural networks, and the remaining 20% were used to evaluate the performance of the algorithm. Results: For the detection of PNI in prostate biopsy cores the network had an estimated area under the receiver operating characteristics curve of 0.98 (95% CI 0.97-0.99) based on 106 PNI positive cores and 1,652 PNI negative cores in the independent test set. For the pre-specified operating point this translates to sensitivity of 0.87 and specificity of 0.97. The corresponding positive and negative predictive values were 0.67 and 0.99, respectively. For localizing the regions of PNI within a slide we estimated an average intersection over union of 0.50 (CI: 0.46-0.55). Conclusion: We have developed an algorithm based on deep neural networks for detecting PNI in prostate biopsies with apparently acceptable diagnostic properties. These algorithms have the potential to aid pathologists in the day-to-day work by drastically reducing the number of biopsy cores that need to be assessed for PNI and by highlighting regions of diagnostic interest.
Pathologist-Level Grading of Prostate Biopsies with Artificial Intelligence
Jul 02, 2019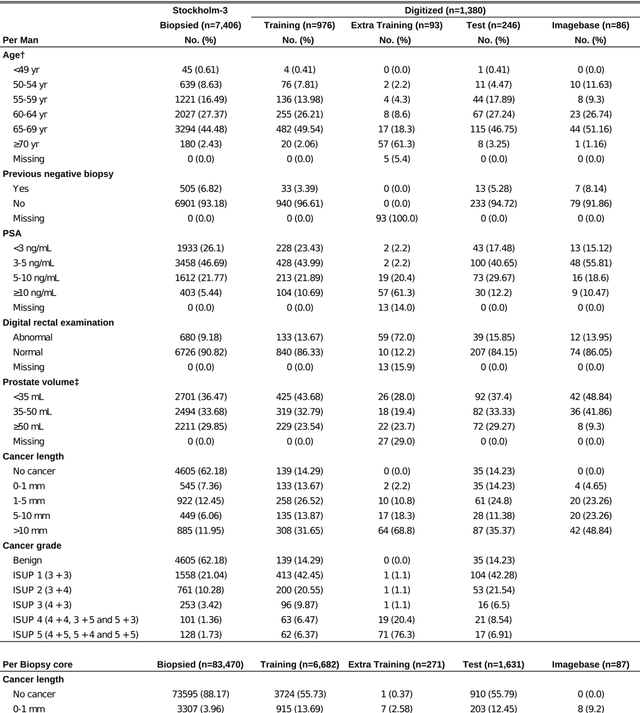
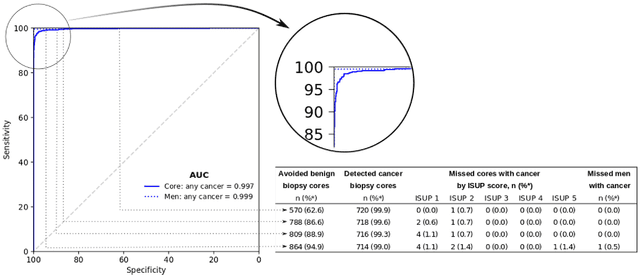
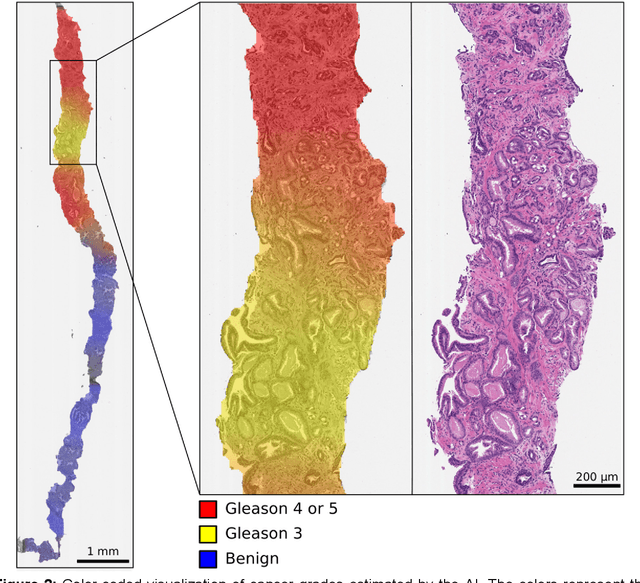
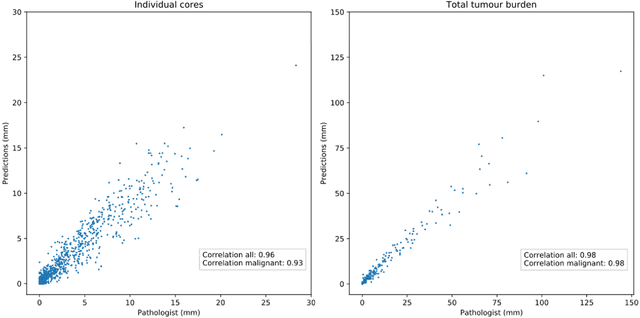
Abstract:Background: An increasing volume of prostate biopsies and a world-wide shortage of uro-pathologists puts a strain on pathology departments. Additionally, the high intra- and inter-observer variability in grading can result in over- and undertreatment of prostate cancer. Artificial intelligence (AI) methods may alleviate these problems by assisting pathologists to reduce workload and harmonize grading. Methods: We digitized 6,682 needle biopsies from 976 participants in the population based STHLM3 diagnostic study to train deep neural networks for assessing prostate biopsies. The networks were evaluated by predicting the presence, extent, and Gleason grade of malignant tissue for an independent test set comprising 1,631 biopsies from 245 men. We additionally evaluated grading performance on 87 biopsies individually graded by 23 experienced urological pathologists from the International Society of Urological Pathology. We assessed discriminatory performance by receiver operating characteristics (ROC) and tumor extent predictions by correlating predicted millimeter cancer length against measurements by the reporting pathologist. We quantified the concordance between grades assigned by the AI and the expert urological pathologists using Cohen's kappa. Results: The performance of the AI to detect and grade cancer in prostate needle biopsy samples was comparable to that of international experts in prostate pathology. The AI achieved an area under the ROC curve of 0.997 for distinguishing between benign and malignant biopsy cores, and 0.999 for distinguishing between men with or without prostate cancer. The correlation between millimeter cancer predicted by the AI and assigned by the reporting pathologist was 0.96. For assigning Gleason grades, the AI achieved an average pairwise kappa of 0.62. This was within the range of the corresponding values for the expert pathologists (0.60 to 0.73).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge