Lubaina Ehsan
The intersection of video capsule endoscopy and artificial intelligence: addressing unique challenges using machine learning
Aug 24, 2023



Abstract:Introduction: Technical burdens and time-intensive review processes limit the practical utility of video capsule endoscopy (VCE). Artificial intelligence (AI) is poised to address these limitations, but the intersection of AI and VCE reveals challenges that must first be overcome. We identified five challenges to address. Challenge #1: VCE data are stochastic and contains significant artifact. Challenge #2: VCE interpretation is cost-intensive. Challenge #3: VCE data are inherently imbalanced. Challenge #4: Existing VCE AIMLT are computationally cumbersome. Challenge #5: Clinicians are hesitant to accept AIMLT that cannot explain their process. Methods: An anatomic landmark detection model was used to test the application of convolutional neural networks (CNNs) to the task of classifying VCE data. We also created a tool that assists in expert annotation of VCE data. We then created more elaborate models using different approaches including a multi-frame approach, a CNN based on graph representation, and a few-shot approach based on meta-learning. Results: When used on full-length VCE footage, CNNs accurately identified anatomic landmarks (99.1%), with gradient weighted-class activation mapping showing the parts of each frame that the CNN used to make its decision. The graph CNN with weakly supervised learning (accuracy 89.9%, sensitivity of 91.1%), the few-shot model (accuracy 90.8%, precision 91.4%, sensitivity 90.9%), and the multi-frame model (accuracy 97.5%, precision 91.5%, sensitivity 94.8%) performed well. Discussion: Each of these five challenges is addressed, in part, by one of our AI-based models. Our goal of producing high performance using lightweight models that aim to improve clinician confidence was achieved.
HistoTransfer: Understanding Transfer Learning for Histopathology
Jun 13, 2021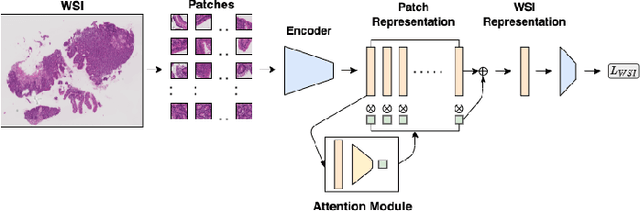
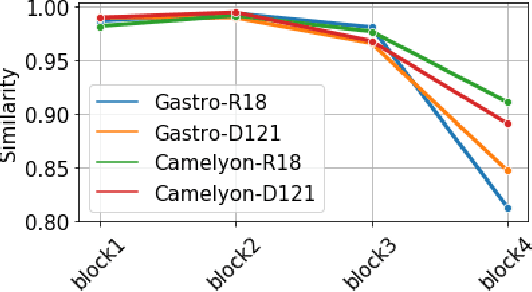
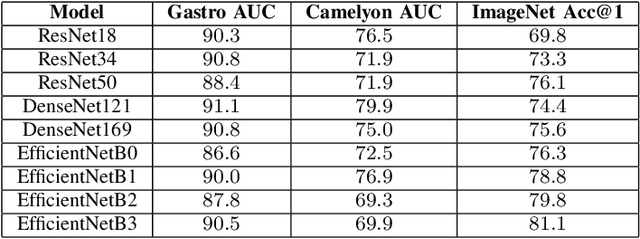

Abstract:Advancement in digital pathology and artificial intelligence has enabled deep learning-based computer vision techniques for automated disease diagnosis and prognosis. However, WSIs present unique computational and algorithmic challenges. WSIs are gigapixel-sized, making them infeasible to be used directly for training deep neural networks. Hence, for modeling, a two-stage approach is adopted: Patch representations are extracted first, followed by the aggregation for WSI prediction. These approaches require detailed pixel-level annotations for training the patch encoder. However, obtaining these annotations is time-consuming and tedious for medical experts. Transfer learning is used to address this gap and deep learning architectures pre-trained on ImageNet are used for generating patch-level representation. Even though ImageNet differs significantly from histopathology data, pre-trained networks have been shown to perform impressively on histopathology data. Also, progress in self-supervised and multi-task learning coupled with the release of multiple histopathology data has led to the release of histopathology-specific networks. In this work, we compare the performance of features extracted from networks trained on ImageNet and histopathology data. We use an attention pooling network over these extracted features for slide-level aggregation. We investigate if features learned using more complex networks lead to gain in performance. We use a simple top-k sampling approach for fine-tuning framework and study the representation similarity between frozen and fine-tuned networks using Centered Kernel Alignment. Further, to examine if intermediate block representation is better suited for feature extraction and ImageNet architectures are unnecessarily large for histopathology, we truncate the blocks of ResNet18 and DenseNet121 and examine the performance.
Cluster-to-Conquer: A Framework for End-to-End Multi-Instance Learning for Whole Slide Image Classification
Mar 19, 2021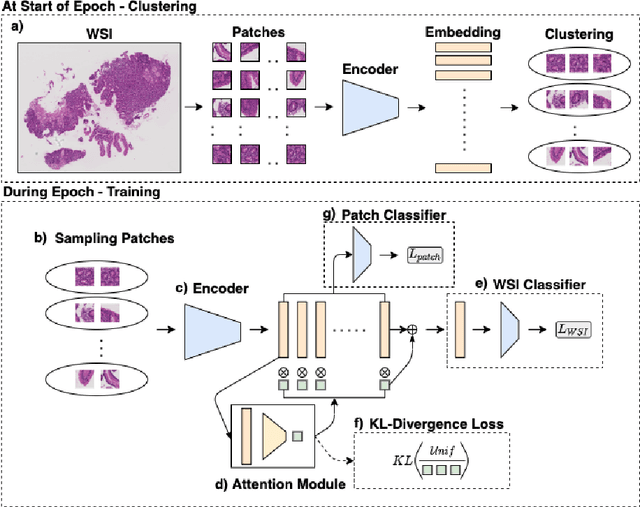
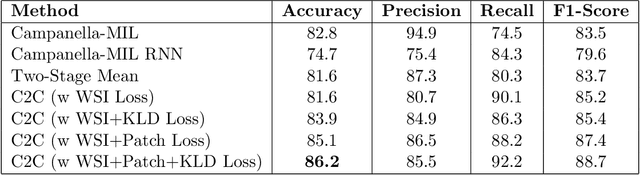
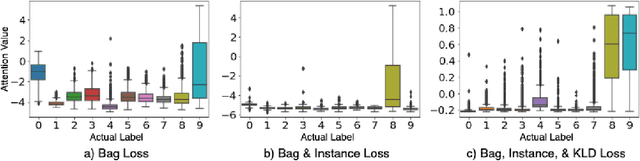
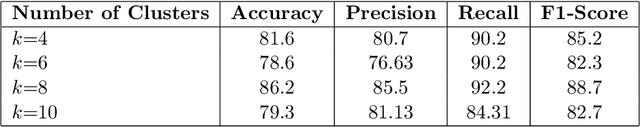
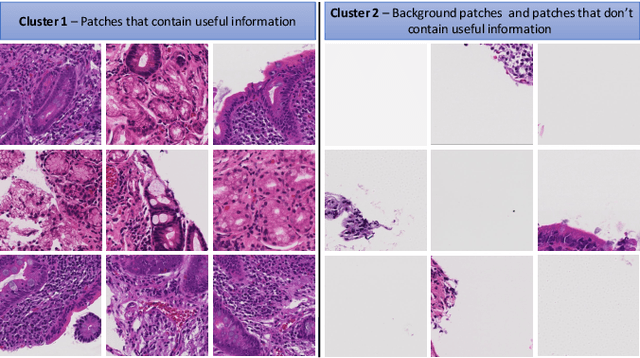
Abstract:In recent years, the availability of digitized Whole Slide Images (WSIs) has enabled the use of deep learning-based computer vision techniques for automated disease diagnosis. However, WSIs present unique computational and algorithmic challenges. WSIs are gigapixel-sized ($\sim$100K pixels), making them infeasible to be used directly for training deep neural networks. Also, often only slide-level labels are available for training as detailed annotations are tedious and can be time-consuming for experts. Approaches using multiple-instance learning (MIL) frameworks have been shown to overcome these challenges. Current state-of-the-art approaches divide the learning framework into two decoupled parts: a convolutional neural network (CNN) for encoding the patches followed by an independent aggregation approach for slide-level prediction. In this approach, the aggregation step has no bearing on the representations learned by the CNN encoder. We have proposed an end-to-end framework that clusters the patches from a WSI into ${k}$-groups, samples ${k}'$ patches from each group for training, and uses an adaptive attention mechanism for slide level prediction; Cluster-to-Conquer (C2C). We have demonstrated that dividing a WSI into clusters can improve the model training by exposing it to diverse discriminative features extracted from the patches. We regularized the clustering mechanism by introducing a KL-divergence loss between the attention weights of patches in a cluster and the uniform distribution. The framework is optimized end-to-end on slide-level cross-entropy, patch-level cross-entropy, and KL-divergence loss (Implementation: https://github.com/YashSharma/C2C).
Advancing Eosinophilic Esophagitis Diagnosis and Phenotype Assessment with Deep Learning Computer Vision
Jan 13, 2021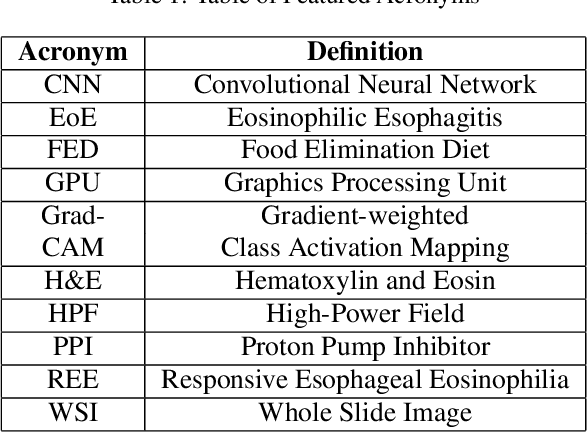

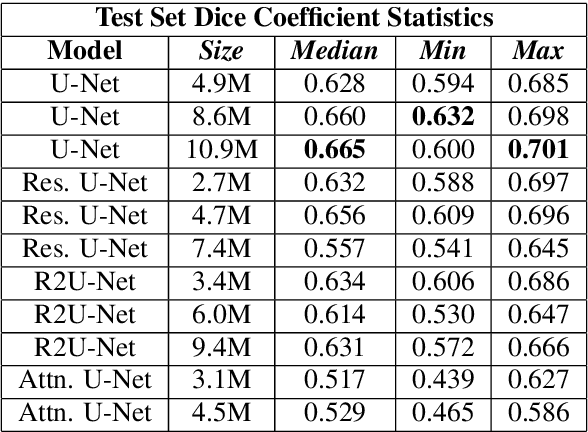
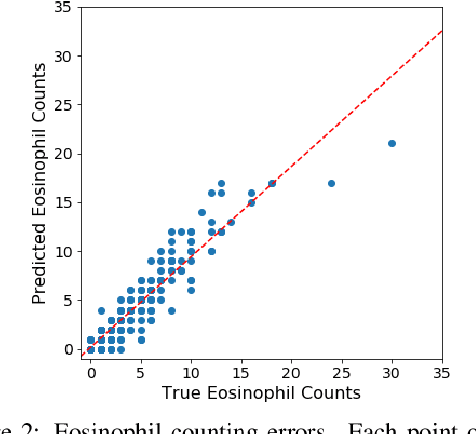
Abstract:Eosinophilic Esophagitis (EoE) is an inflammatory esophageal disease which is increasing in prevalence. The diagnostic gold-standard involves manual review of a patient's biopsy tissue sample by a clinical pathologist for the presence of 15 or greater eosinophils within a single high-power field (400x magnification). Diagnosing EoE can be a cumbersome process with added difficulty for assessing the severity and progression of disease. We propose an automated approach for quantifying eosinophils using deep image segmentation. A U-Net model and post-processing system are applied to generate eosinophil-based statistics that can diagnose EoE as well as describe disease severity and progression. These statistics are captured in biopsies at the initial EoE diagnosis and are then compared with patient metadata: clinical and treatment phenotypes. The goal is to find linkages that could potentially guide treatment plans for new patients at their initial disease diagnosis. A deep image classification model is further applied to discover features other than eosinophils that can be used to diagnose EoE. This is the first study to utilize a deep learning computer vision approach for EoE diagnosis and to provide an automated process for tracking disease severity and progression.
HMIC: Hierarchical Medical Image Classification, A Deep Learning Approach
Jun 23, 2020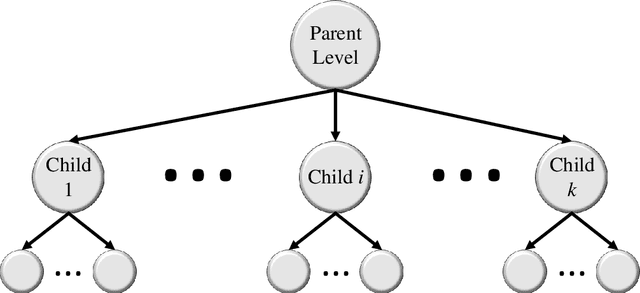
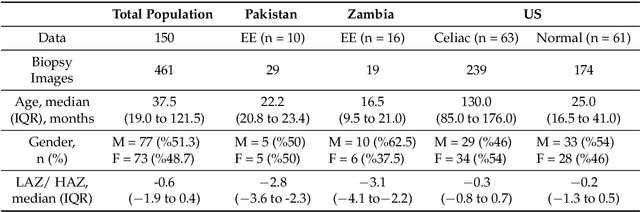
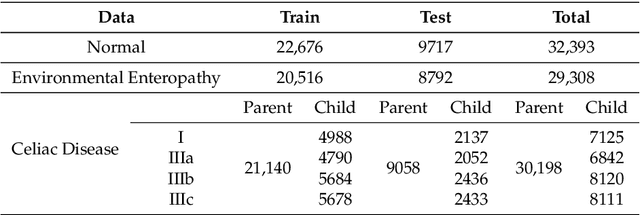
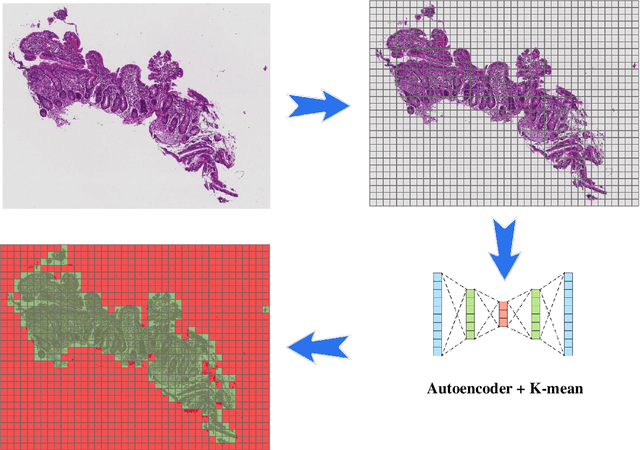
Abstract:Image classification is central to the big data revolution in medicine. Improved information processing methods for diagnosis and classification of digital medical images have shown to be successful via deep learning approaches. As this field is explored, there are limitations to the performance of traditional supervised classifiers. This paper outlines an approach that is different from the current medical image classification tasks that view the issue as multi-class classification. We performed a hierarchical classification using our Hierarchical Medical Image classification (HMIC) approach. HMIC uses stacks of deep learning models to give particular comprehension at each level of the clinical picture hierarchy. For testing our performance, we use biopsy of the small bowel images that contain three categories in the parent level (Celiac Disease, Environmental Enteropathy, and histologically normal controls). For the child level, Celiac Disease Severity is classified into 4 classes (I, IIIa, IIIb, and IIIC).
Hierarchical Deep Convolutional Neural Networks for Multi-category Diagnosis of Gastrointestinal Disorders on Histopathological Images
May 08, 2020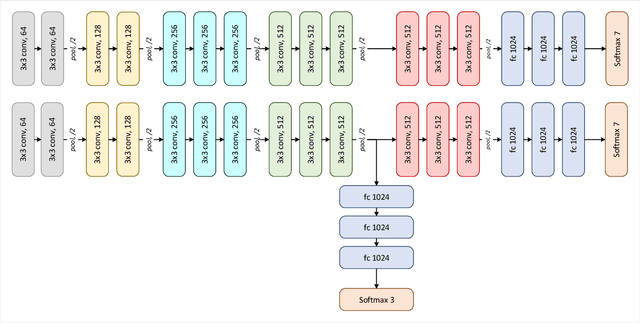
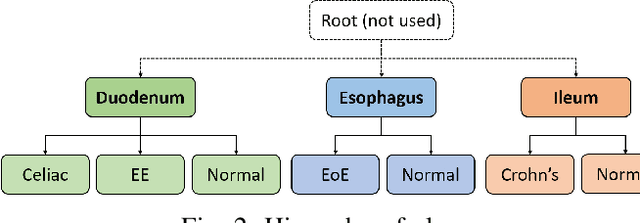
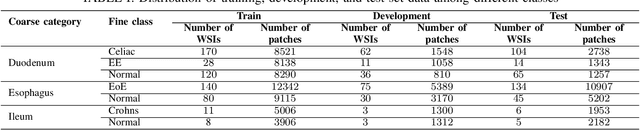
Abstract:Deep convolutional neural networks (CNNs) have been successful for a wide range of computer vision tasks including image classification. A specific area of application lies in digital pathology for pattern recognition in tissue-based diagnosis of gastrointestinal (GI) diseases. This domain can utilize CNNs to translate histopathological images into precise diagnostics. This is challenging since these complex biopsies are heterogeneous and require multiple levels of assessment. This is mainly due to structural similarities in different parts of the GI tract and shared features among different gut diseases. Addressing this problem with a flat model which assumes all classes (parts of the gut and their diseases) are equally difficult to distinguish leads to an inadequate assessment of each class. Since hierarchical model restricts classification error to each sub-class, it leads to a more informative model compared to a flat model. In this paper we propose to apply hierarchical classification of biopsy images from different parts of the GI tract and the receptive diseases within each. We embedded a class hierarchy into the plain VGGNet to take advantage of the hierarchical structure of its layers. The proposed model was evaluated using an independent set of image patches from 373 whole slide images. The results indicate that hierarchical model can achieve better results compared to the flat model for multi-category diagnosis of GI disorders using histopathological images.
CeliacNet: Celiac Disease Severity Diagnosis on Duodenal Histopathological Images Using Deep Residual Networks
Oct 07, 2019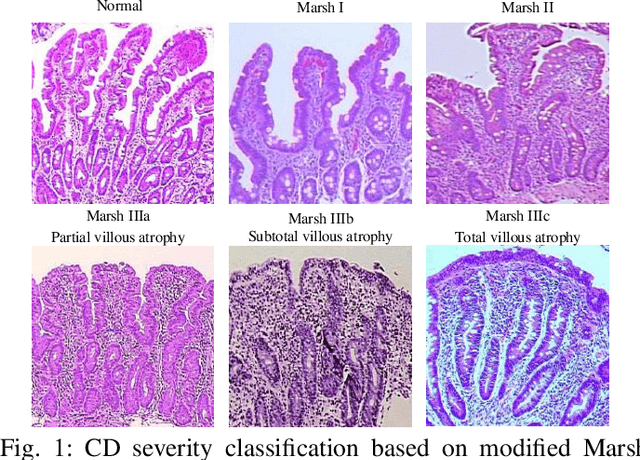
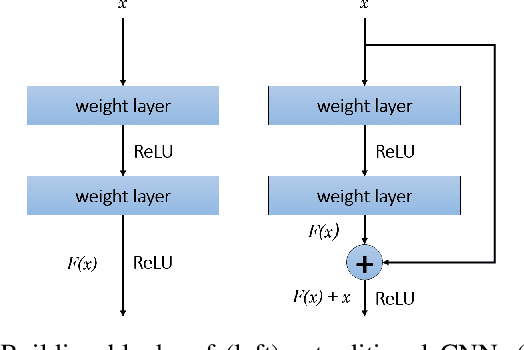
Abstract:Celiac Disease (CD) is a chronic autoimmune disease that affects the small intestine in genetically predisposed children and adults. Gluten exposure triggers an inflammatory cascade which leads to compromised intestinal barrier function. If this enteropathy is unrecognized, this can lead to anemia, decreased bone density, and, in longstanding cases, intestinal cancer. The prevalence of the disorder is 1% in the United States. An intestinal (duodenal) biopsy is considered the "gold standard" for diagnosis. The mild CD might go unnoticed due to non-specific clinical symptoms or mild histologic features. In our current work, we trained a model based on deep residual networks to diagnose CD severity using a histological scoring system called the modified Marsh score. The proposed model was evaluated using an independent set of 120 whole slide images from 15 CD patients and achieved an AUC greater than 0.96 in all classes. These results demonstrate the diagnostic power of the proposed model for CD severity classification using histological images.
Self-Attentive Adversarial Stain Normalization
Sep 04, 2019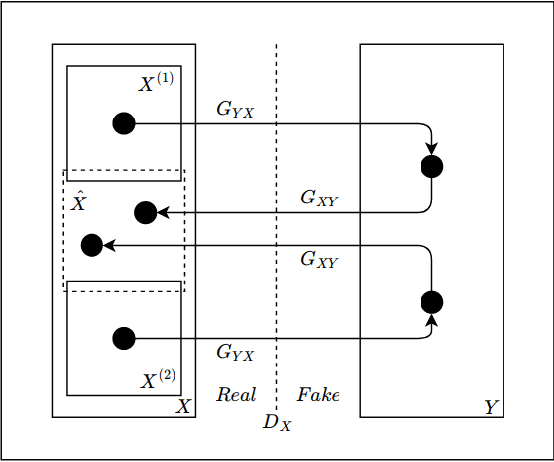
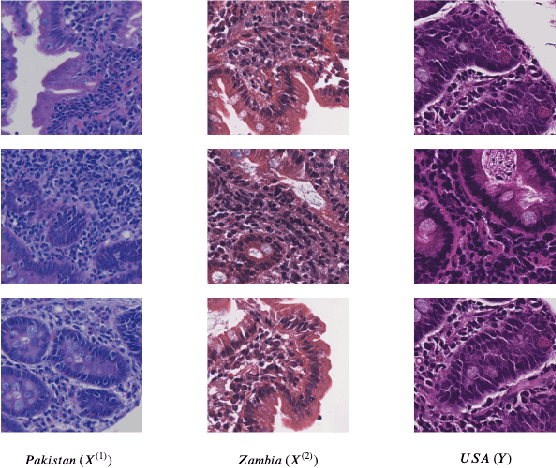
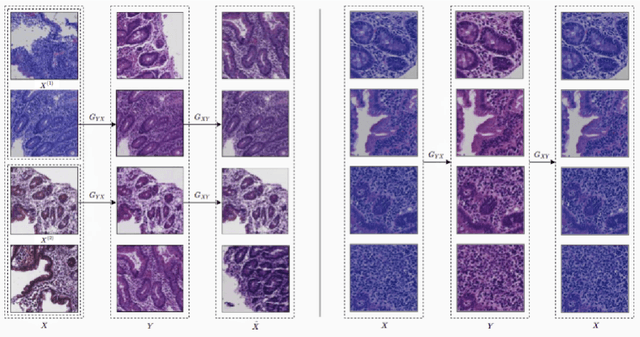
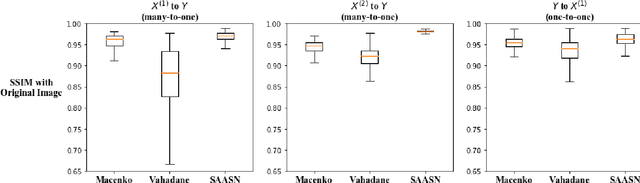
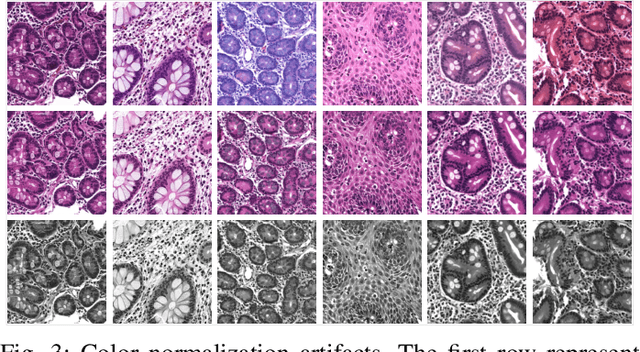
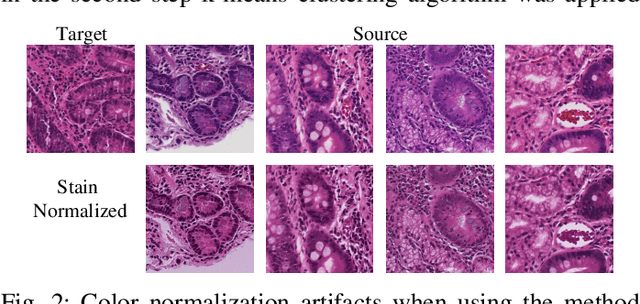
Abstract:Hematoxylin and Eosin (H&E) stained Whole Slide Images (WSIs) are utilized for biopsy visualization-based diagnostic and prognostic assessment of diseases. Variation in the H&E staining process across different lab sites can lead to significant variations in biopsy image appearance. These variations introduce an undesirable bias when the slides are examined by pathologists or used for training deep learning models. To reduce this bias, slides need to be translated to a common domain of stain appearance before analysis. We propose a Self-Attentive Adversarial Stain Normalization (SAASN) approach for the normalization of multiple stain appearances to a common domain. This unsupervised generative adversarial approach includes self-attention mechanism for synthesizing images with finer detail while preserving the structural consistency of the biopsy features during translation. SAASN demonstrates consistent and superior performance compared to other popular stain normalization techniques on H&E stained duodenal biopsy image data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge