Liron Pantanowitz
On Image Search in Histopathology
Jan 14, 2024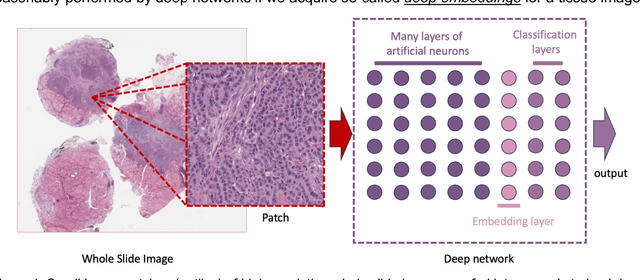
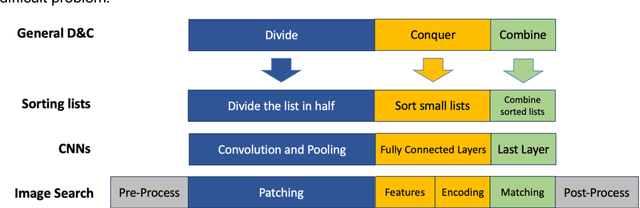
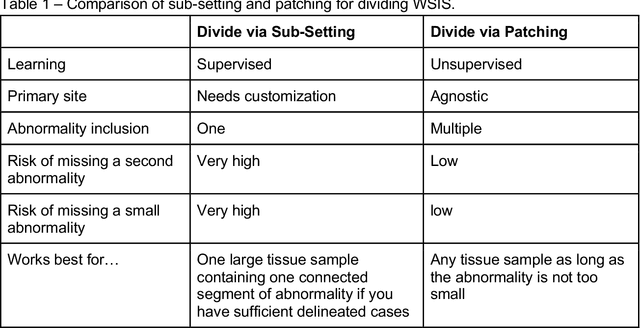
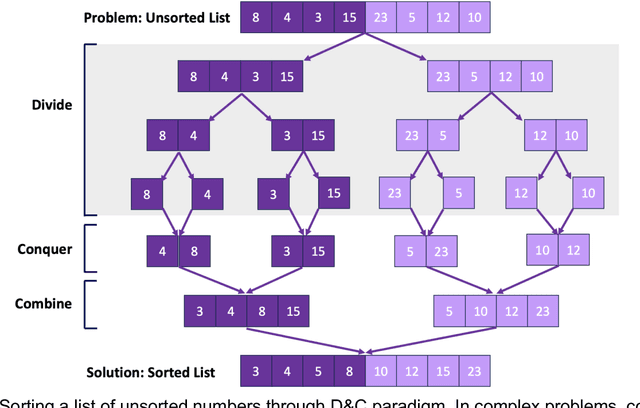
Abstract:Pathology images of histopathology can be acquired from camera-mounted microscopes or whole slide scanners. Utilizing similarity calculations to match patients based on these images holds significant potential in research and clinical contexts. Recent advancements in search technologies allow for nuanced quantification of cellular structures across diverse tissue types, facilitating comparisons and enabling inferences about diagnosis, prognosis, and predictions for new patients when compared against a curated database of diagnosed and treated cases. In this paper, we comprehensively review the latest developments in image search technologies for histopathology, offering a concise overview tailored for computational pathology researchers seeking effective, fast and efficient image search methods in their work.
A Preliminary Investigation into Search and Matching for Tumour Discrimination in WHO Breast Taxonomy Using Deep Networks
Aug 22, 2023Abstract:Breast cancer is one of the most common cancers affecting women worldwide. They include a group of malignant neoplasms with a variety of biological, clinical, and histopathological characteristics. There are more than 35 different histological forms of breast lesions that can be classified and diagnosed histologically according to cell morphology, growth, and architecture patterns. Recently, deep learning, in the field of artificial intelligence, has drawn a lot of attention for the computerized representation of medical images. Searchable digital atlases can provide pathologists with patch matching tools allowing them to search among evidently diagnosed and treated archival cases, a technology that may be regarded as computational second opinion. In this study, we indexed and analyzed the WHO breast taxonomy (Classification of Tumours 5th Ed.) spanning 35 tumour types. We visualized all tumour types using deep features extracted from a state-of-the-art deep learning model, pre-trained on millions of diagnostic histopathology images from the TCGA repository. Furthermore, we test the concept of a digital "atlas" as a reference for search and matching with rare test cases. The patch similarity search within the WHO breast taxonomy data reached over 88% accuracy when validating through "majority vote" and more than 91% accuracy when validating using top-n tumour types. These results show for the first time that complex relationships among common and rare breast lesions can be investigated using an indexed digital archive.
An Investigation into Glomeruli Detection in Kidney H&E and PAS Images using YOLO
Jul 25, 2023



Abstract:Context: Analyzing digital pathology images is necessary to draw diagnostic conclusions by investigating tissue patterns and cellular morphology. However, manual evaluation can be time-consuming, expensive, and prone to inter- and intra-observer variability. Objective: To assist pathologists using computerized solutions, automated tissue structure detection and segmentation must be proposed. Furthermore, generating pixel-level object annotations for histopathology images is expensive and time-consuming. As a result, detection models with bounding box labels may be a feasible solution. Design: This paper studies. YOLO-v4 (You-Only-Look-Once), a real-time object detector for microscopic images. YOLO uses a single neural network to predict several bounding boxes and class probabilities for objects of interest. YOLO can enhance detection performance by training on whole slide images. YOLO-v4 has been used in this paper. for glomeruli detection in human kidney images. Multiple experiments have been designed and conducted based on different training data of two public datasets and a private dataset from the University of Michigan for fine-tuning the model. The model was tested on the private dataset from the University of Michigan, serving as an external validation of two different stains, namely hematoxylin and eosin (H&E) and periodic acid-Schiff (PAS). Results: Average specificity and sensitivity for all experiments, and comparison of existing segmentation methods on the same datasets are discussed. Conclusions: Automated glomeruli detection in human kidney images is possible using modern AI models. The design and validation for different stains still depends on variability of public multi-stain datasets.
Fine-Tuning and Training of DenseNet for Histopathology Image Representation Using TCGA Diagnostic Slides
Jan 20, 2021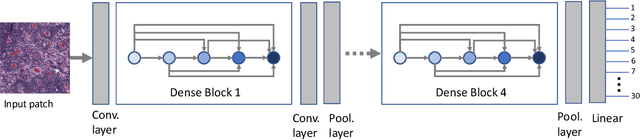
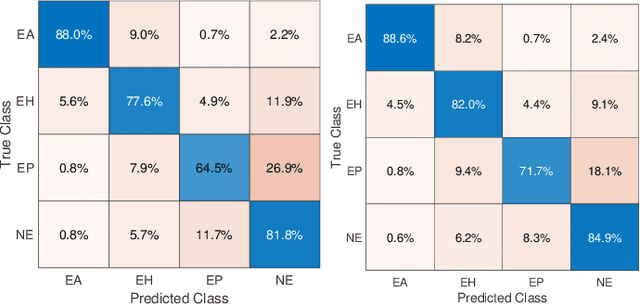
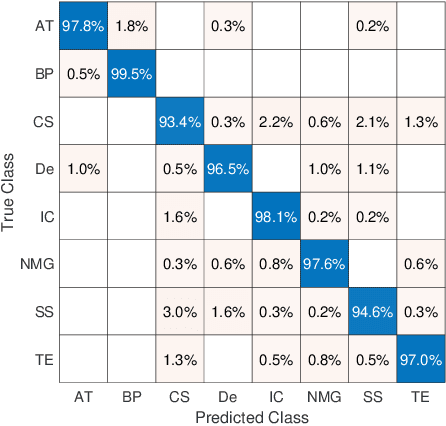
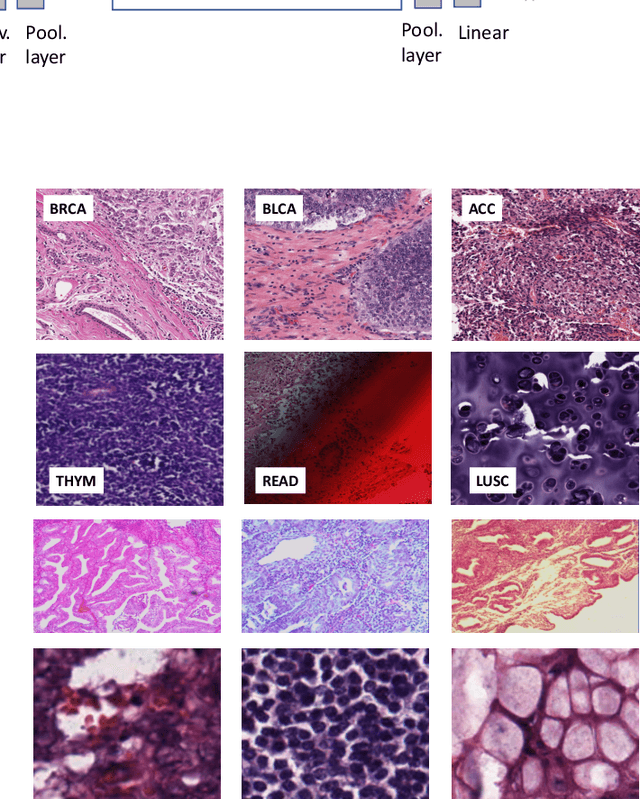
Abstract:Feature vectors provided by pre-trained deep artificial neural networks have become a dominant source for image representation in recent literature. Their contribution to the performance of image analysis can be improved through finetuning. As an ultimate solution, one might even train a deep network from scratch with the domain-relevant images, a highly desirable option which is generally impeded in pathology by lack of labeled images and the computational expense. In this study, we propose a new network, namely KimiaNet, that employs the topology of the DenseNet with four dense blocks, fine-tuned and trained with histopathology images in different configurations. We used more than 240,000 image patches with 1000x1000 pixels acquired at 20x magnification through our proposed "highcellularity mosaic" approach to enable the usage of weak labels of 7,126 whole slide images of formalin-fixed paraffin-embedded human pathology samples publicly available through the The Cancer Genome Atlas (TCGA) repository. We tested KimiaNet using three public datasets, namely TCGA, endometrial cancer images, and colorectal cancer images by evaluating the performance of search and classification when corresponding features of different networks are used for image representation. As well, we designed and trained multiple convolutional batch-normalized ReLU (CBR) networks. The results show that KimiaNet provides superior results compared to the original DenseNet and smaller CBR networks when used as feature extractor to represent histopathology images.
Pan-Cancer Diagnostic Consensus Through Searching Archival Histopathology Images Using Artificial Intelligence
Nov 20, 2019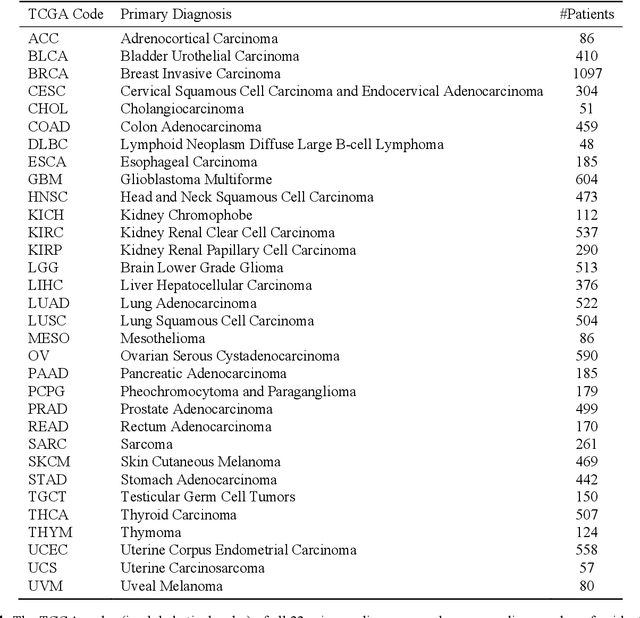
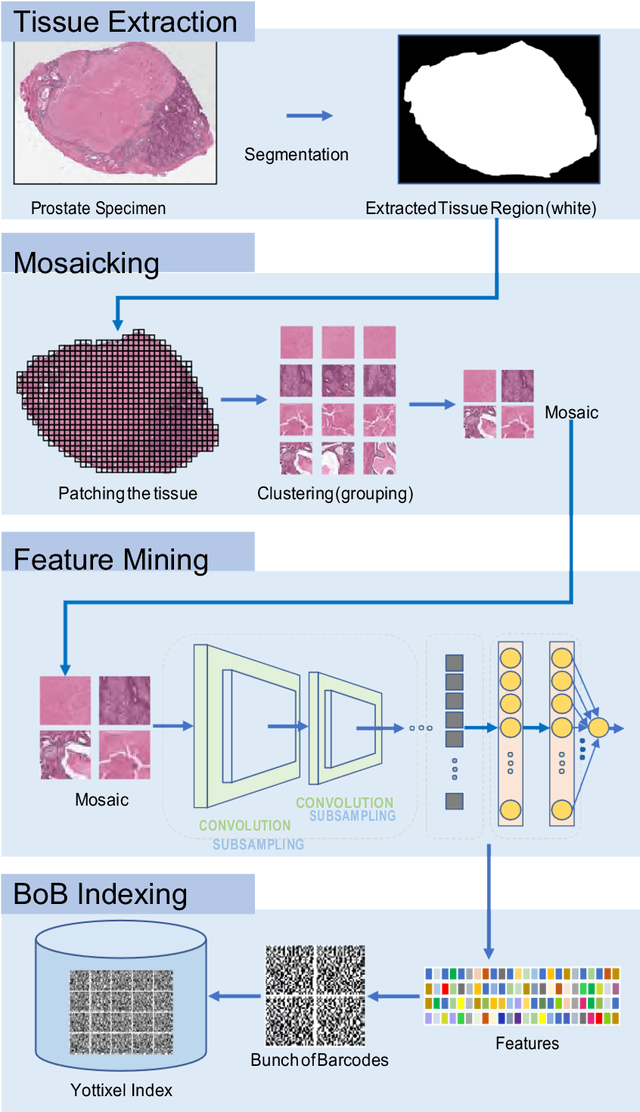

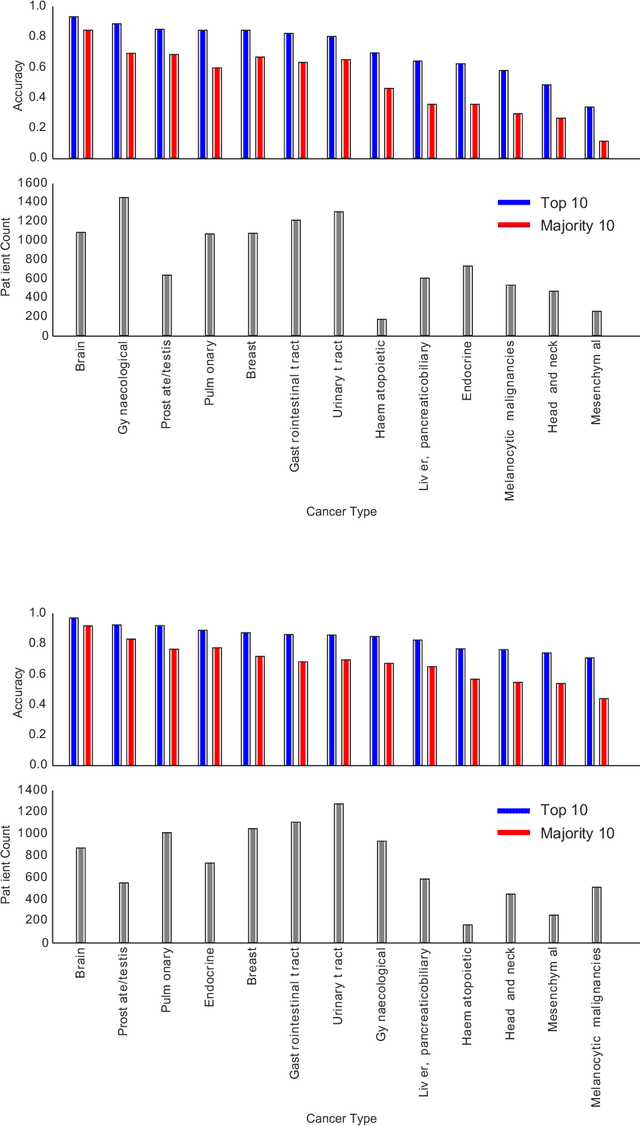
Abstract:The emergence of digital pathology has opened new horizons for histopathology and cytology. Artificial-intelligence algorithms are able to operate on digitized slides to assist pathologists with diagnostic tasks. Whereas machine learning involving classification and segmentation methods have obvious benefits for image analysis in pathology, image search represents a fundamental shift in computational pathology. Matching the pathology of new patients with already diagnosed and curated cases offers pathologist a novel approach to improve diagnostic accuracy through visual inspection of similar cases and computational majority vote for consensus building. In this study, we report the results from searching the largest public repository (The Cancer Genome Atlas [TCGA] program by National Cancer Institute, USA) of whole slide images from almost 11,000 patients depicting different types of malignancies. For the first time, we successfully indexed and searched almost 30,000 high-resolution digitized slides constituting 16 terabytes of data comprised of 20 million 1000x1000 pixels image patches. The TCGA image database covers 25 anatomic sites and contains 32 cancer subtypes. High-performance storage and GPU power were employed for experimentation. The results were assessed with conservative "majority voting" to build consensus for subtype diagnosis through vertical search and demonstrated high accuracy values for both frozen sections slides (e.g., bladder urothelial carcinoma 93%, kidney renal clear cell carcinoma 97%, and ovarian serous cystadenocarcinoma 99%) and permanent histopathology slides (e.g., prostate adenocarcinoma 98%, skin cutaneous melanoma 99%, and thymoma 100%). The key finding of this validation study was that computational consensus appears to be possible for rendering diagnoses if a sufficiently large number of searchable cases are available for each cancer subtype.
Aging display's effect on interpretation of digital pathology slides
Jun 30, 2015Abstract:It is our conjecture that the variability of colors in a pathology image effects the interpretation of pathology cases, whether it is diagnostic accuracy, diagnostic confidence, or workflow efficiency. In this paper, digital pathology images are analyzed to quantify the perceived difference in color that occurs due to display aging, in particular a change in the maximum luminance, white point, and color gamut. The digital pathology images studied include diagnostically important features, such as the conspicuity of nuclei. Three different display aging models are applied to images: aging of luminance & chrominance, aging of chrominance only, and a stabilized luminance & chrominance (i.e., no aging). These display models and images are then used to compare conspicuity of nuclei using CIE deltaE2000, a perceptual color difference metric. The effect of display aging using these display models and images is further analyzed through a human reader study designed to quantify the effects from a clinical perspective. Results from our reader study indicate significant impact of aged displays on workflow as well as diagnosis as follow. As compared to the originals (no-aging), slides with the effect of aging simulated were significantly more difficult to read (p-value of 0.0005) and took longer to score (p-value of 0.02). Moreover, luminance+chrominance aging significantly reduced inter-session percent agreement of diagnostic scores (p-value of 0.0418).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge