Abubakr Shafique
Kimia Lab, University of Waterloo, Waterloo, ON, Canada
Teacher-Student Model for Detecting and Classifying Mitosis in the MIDOG 2025 Challenge
Sep 03, 2025Abstract:Counting mitotic figures is time-intensive for pathologists and leads to inter-observer variability. Artificial intelligence (AI) promises a solution by automatically detecting mitotic figures while maintaining decision consistency. However, AI tools are susceptible to domain shift, where a significant drop in performance can occur due to differences in the training and testing sets, including morphological diversity between organs, species, and variations in staining protocols. Furthermore, the number of mitoses is much less than the count of normal nuclei, which introduces severely imbalanced data for the detection task. In this work, we formulate mitosis detection as a pixel-level segmentation and propose a teacher-student model that simultaneously addresses mitosis detection (Track 1) and atypical mitosis classification (Track 2). Our method is based on a UNet segmentation backbone that integrates domain generalization modules, namely contrastive representation learning and domain-adversarial training. A teacher-student strategy is employed to generate pixel-level pseudo-masks not only for annotated mitoses and hard negatives but also for normal nuclei, thereby enhancing feature discrimination and improving robustness against domain shift. For the classification task, we introduce a multi-scale CNN classifier that leverages feature maps from the segmentation model within a multi-task learning paradigm. On the preliminary test set, the algorithm achieved an F1 score of 0.7660 in Track 1 and balanced accuracy of 0.8414 in Track 2, demonstrating the effectiveness of integrating segmentation-based detection and classification into a unified framework for robust mitosis analysis.
SPLICE -- Streamlining Digital Pathology Image Processing
Apr 26, 2024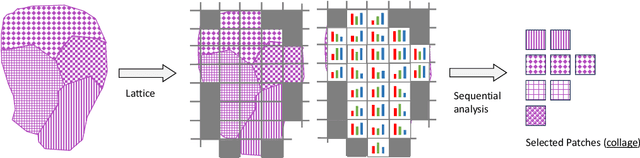
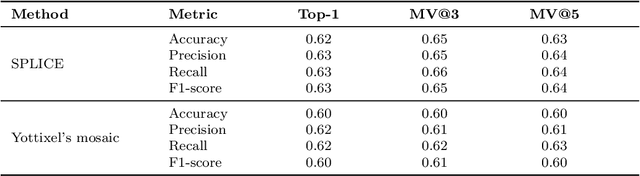
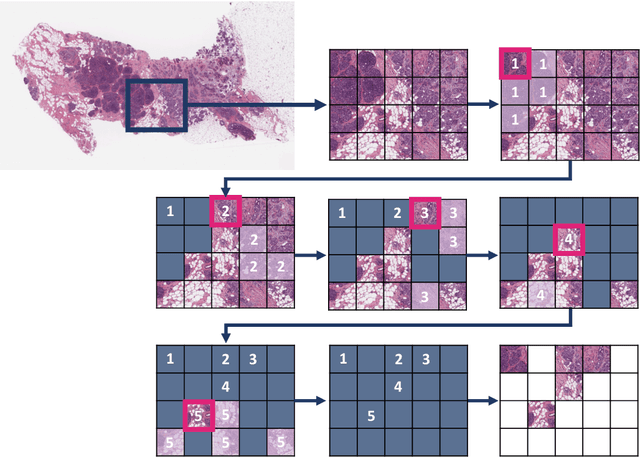
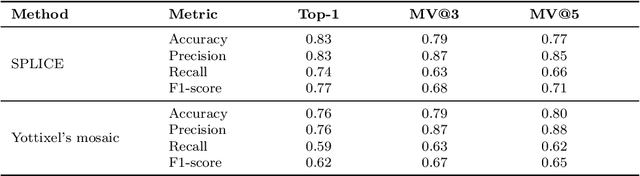
Abstract:Digital pathology and the integration of artificial intelligence (AI) models have revolutionized histopathology, opening new opportunities. With the increasing availability of Whole Slide Images (WSIs), there's a growing demand for efficient retrieval, processing, and analysis of relevant images from vast biomedical archives. However, processing WSIs presents challenges due to their large size and content complexity. Full computer digestion of WSIs is impractical, and processing all patches individually is prohibitively expensive. In this paper, we propose an unsupervised patching algorithm, Sequential Patching Lattice for Image Classification and Enquiry (SPLICE). This novel approach condenses a histopathology WSI into a compact set of representative patches, forming a "collage" of WSI while minimizing redundancy. SPLICE prioritizes patch quality and uniqueness by sequentially analyzing a WSI and selecting non-redundant representative features. We evaluated SPLICE for search and match applications, demonstrating improved accuracy, reduced computation time, and storage requirements compared to existing state-of-the-art methods. As an unsupervised method, SPLICE effectively reduces storage requirements for representing tissue images by 50%. This reduction enables numerous algorithms in computational pathology to operate much more efficiently, paving the way for accelerated adoption of digital pathology.
Analysis and Validation of Image Search Engines in Histopathology
Jan 06, 2024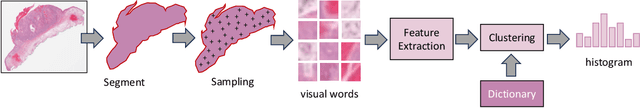
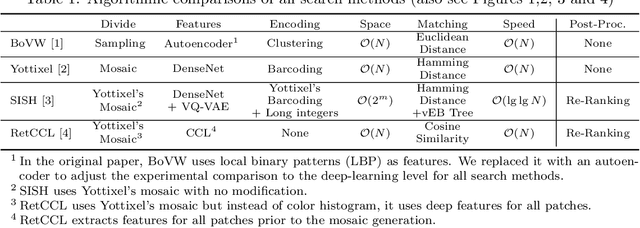
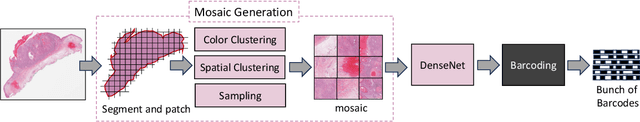
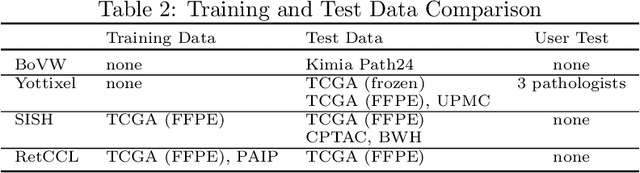
Abstract:Searching for similar images in archives of histology and histopathology images is a crucial task that may aid in patient matching for various purposes, ranging from triaging and diagnosis to prognosis and prediction. Whole slide images (WSIs) are highly detailed digital representations of tissue specimens mounted on glass slides. Matching WSI to WSI can serve as the critical method for patient matching. In this paper, we report extensive analysis and validation of four search methods bag of visual words (BoVW), Yottixel, SISH, RetCCL, and some of their potential variants. We analyze their algorithms and structures and assess their performance. For this evaluation, we utilized four internal datasets ($1269$ patients) and three public datasets ($1207$ patients), totaling more than $200,000$ patches from $38$ different classes/subtypes across five primary sites. Certain search engines, for example, BoVW, exhibit notable efficiency and speed but suffer from low accuracy. Conversely, search engines like Yottixel demonstrate efficiency and speed, providing moderately accurate results. Recent proposals, including SISH, display inefficiency and yield inconsistent outcomes, while alternatives like RetCCL prove inadequate in both accuracy and efficiency. Further research is imperative to address the dual aspects of accuracy and minimal storage requirements in histopathological image search.
Selection of Distinct Morphologies to Divide & Conquer Gigapixel Pathology Images
Nov 16, 2023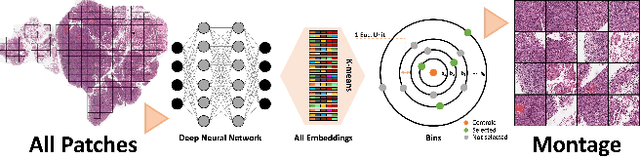
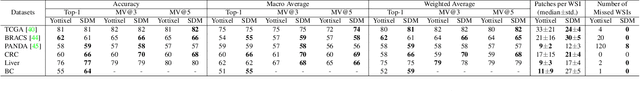
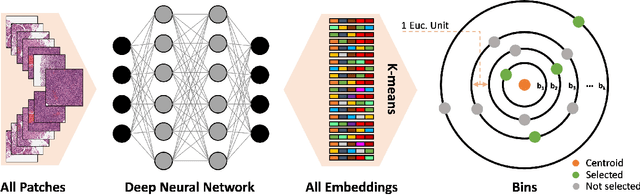
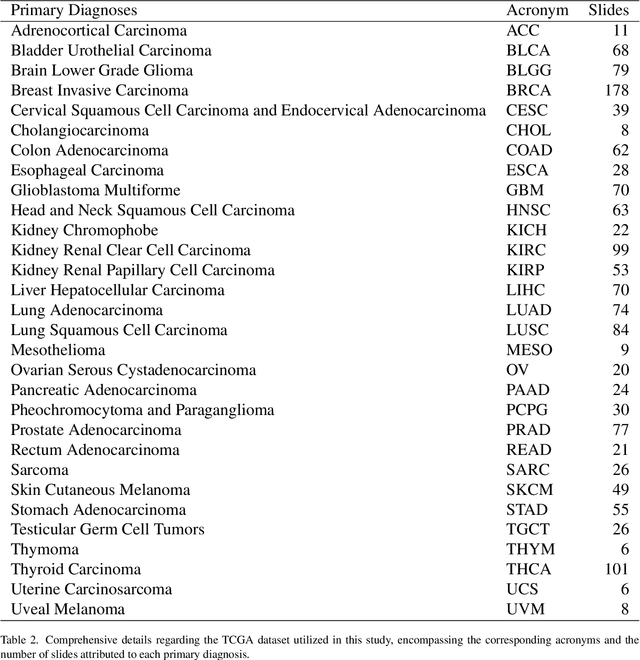
Abstract:Whole slide images (WSIs) are massive digital pathology files illustrating intricate tissue structures. Selecting a small, representative subset of patches from each WSI is essential yet challenging. Therefore, following the "Divide & Conquer" approach becomes essential to facilitate WSI analysis including the classification and the WSI matching in computational pathology. To this end, we propose a novel method termed "Selection of Distinct Morphologies" (SDM) to choose a subset of WSI patches. The aim is to encompass all inherent morphological variations within a given WSI while simultaneously minimizing the number of selected patches to represent these variations, ensuring a compact yet comprehensive set of patches. This systematically curated patch set forms what we term a "montage". We assess the representativeness of the SDM montage across various public and private histopathology datasets. This is conducted by using the leave-one-out WSI search and matching evaluation method, comparing it with the state-of-the-art Yottixel's mosaic. SDM demonstrates remarkable efficacy across all datasets during its evaluation. Furthermore, SDM eliminates the necessity for empirical parameterization, a crucial aspect of Yottixel's mosaic, by inherently optimizing the selection process to capture the distinct morphological features within the WSI.
Rotation-Agnostic Image Representation Learning for Digital Pathology
Nov 14, 2023

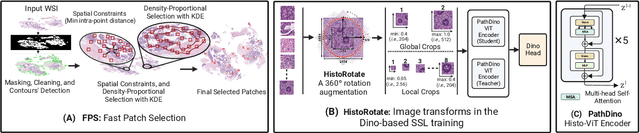
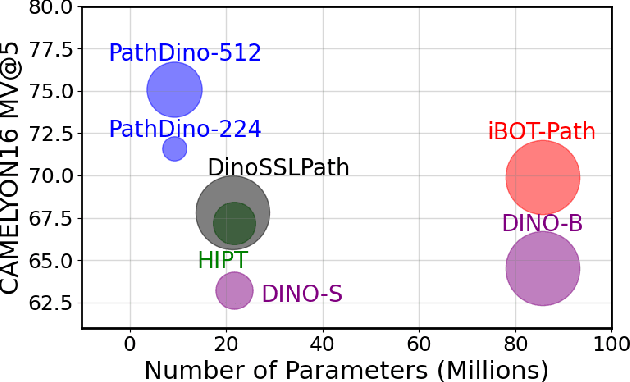
Abstract:This paper addresses complex challenges in histopathological image analysis through three key contributions. Firstly, it introduces a fast patch selection method, FPS, for whole-slide image (WSI) analysis, significantly reducing computational cost while maintaining accuracy. Secondly, it presents PathDino, a lightweight histopathology feature extractor with a minimal configuration of five Transformer blocks and only 9 million parameters, markedly fewer than alternatives. Thirdly, it introduces a rotation-agnostic representation learning paradigm using self-supervised learning, effectively mitigating overfitting. We also show that our compact model outperforms existing state-of-the-art histopathology-specific vision transformers on 12 diverse datasets, including both internal datasets spanning four sites (breast, liver, skin, and colorectal) and seven public datasets (PANDA, CAMELYON16, BRACS, DigestPath, Kather, PanNuke, and WSSS4LUAD). Notably, even with a training dataset of 6 million histopathology patches from The Cancer Genome Atlas (TCGA), our approach demonstrates an average 8.5% improvement in patch-level majority vote performance. These contributions provide a robust framework for enhancing image analysis in digital pathology, rigorously validated through extensive evaluation. Project Page: https://rhazeslab.github.io/PathDino-Page/
When is a Foundation Model a Foundation Model
Sep 14, 2023

Abstract:Recently, several studies have reported on the fine-tuning of foundation models for image-text modeling in the field of medicine, utilizing images from online data sources such as Twitter and PubMed. Foundation models are large, deep artificial neural networks capable of learning the context of a specific domain through training on exceptionally extensive datasets. Through validation, we have observed that the representations generated by such models exhibit inferior performance in retrieval tasks within digital pathology when compared to those generated by significantly smaller, conventional deep networks.
A Preliminary Investigation into Search and Matching for Tumour Discrimination in WHO Breast Taxonomy Using Deep Networks
Aug 22, 2023Abstract:Breast cancer is one of the most common cancers affecting women worldwide. They include a group of malignant neoplasms with a variety of biological, clinical, and histopathological characteristics. There are more than 35 different histological forms of breast lesions that can be classified and diagnosed histologically according to cell morphology, growth, and architecture patterns. Recently, deep learning, in the field of artificial intelligence, has drawn a lot of attention for the computerized representation of medical images. Searchable digital atlases can provide pathologists with patch matching tools allowing them to search among evidently diagnosed and treated archival cases, a technology that may be regarded as computational second opinion. In this study, we indexed and analyzed the WHO breast taxonomy (Classification of Tumours 5th Ed.) spanning 35 tumour types. We visualized all tumour types using deep features extracted from a state-of-the-art deep learning model, pre-trained on millions of diagnostic histopathology images from the TCGA repository. Furthermore, we test the concept of a digital "atlas" as a reference for search and matching with rare test cases. The patch similarity search within the WHO breast taxonomy data reached over 88% accuracy when validating through "majority vote" and more than 91% accuracy when validating using top-n tumour types. These results show for the first time that complex relationships among common and rare breast lesions can be investigated using an indexed digital archive.
Immunohistochemistry Biomarkers-Guided Image Search for Histopathology
Apr 24, 2023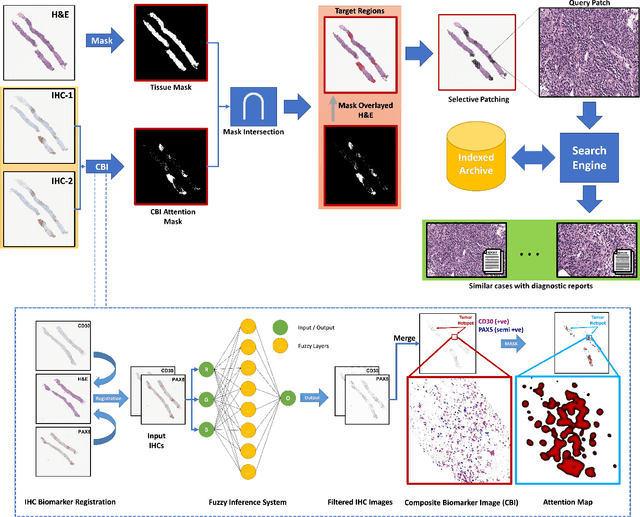
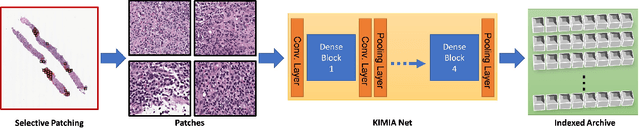
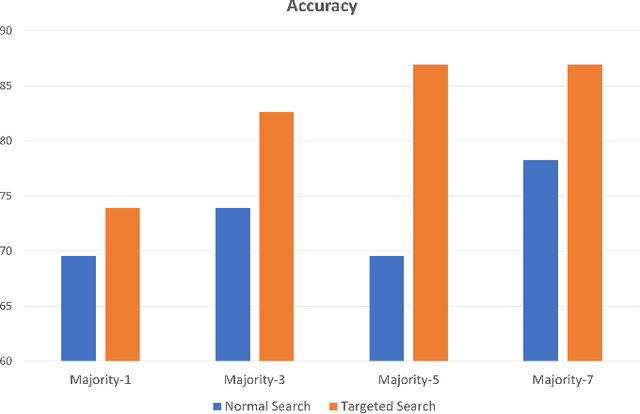
Abstract:Medical practitioners use a number of diagnostic tests to make a reliable diagnosis. Traditionally, Haematoxylin and Eosin (H&E) stained glass slides have been used for cancer diagnosis and tumor detection. However, recently a variety of immunohistochemistry (IHC) stained slides can be requested by pathologists to examine and confirm diagnoses for determining the subtype of a tumor when this is difficult using H&E slides only. Deep learning (DL) has received a lot of interest recently for image search engines to extract features from tissue regions, which may or may not be the target region for diagnosis. This approach generally fails to capture high-level patterns corresponding to the malignant or abnormal content of histopathology images. In this work, we are proposing a targeted image search approach, inspired by the pathologists workflow, which may use information from multiple IHC biomarker images when available. These IHC images could be aligned, filtered, and merged together to generate a composite biomarker image (CBI) that could eventually be used to generate an attention map to guide the search engine for localized search. In our experiments, we observed that an IHC-guided image search engine can retrieve relevant data more accurately than a conventional (i.e., H&E-only) search engine without IHC guidance. Moreover, such engines are also able to accurately conclude the subtypes through majority votes.
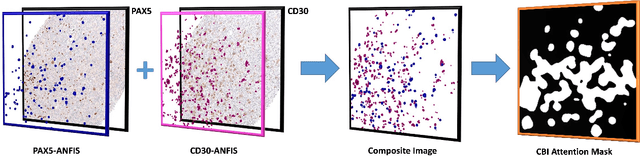
Composite Biomarker Image for Advanced Visualization in Histopathology
Apr 24, 2023



Abstract:Immunohistochemistry (IHC) biomarkers are essential tools for reliable cancer diagnosis and subtyping. It requires cross-staining comparison among Whole Slide Images (WSIs) of IHCs and hematoxylin and eosin (H&E) slides. Currently, pathologists examine the visually co-localized areas across IHC and H&E glass slides for a final diagnosis, which is a tedious and challenging task. Moreover, visually inspecting different IHC slides back and forth to analyze local co-expressions is inherently subjective and prone to error, even when carried out by experienced pathologists. Relying on digital pathology, we propose Composite Biomarker Image (CBI) in this work. CBI is a single image that can be composed using different filtered IHC biomarker images for better visualization. We present a CBI image produced in two steps by the proposed solution for better visualization and hence more efficient clinical workflow. In the first step, IHC biomarker images are aligned with the H&E images using one coordinate system and orientation. In the second step, the positive or negative IHC regions from each biomarker image (based on the pathologists recommendation) are filtered and combined into one image using a fuzzy inference system. For evaluation, the resulting CBI images, from the proposed system, were evaluated qualitatively by the expert pathologists. The CBI concept helps the pathologists to identify the suspected target tissues more easily, which could be further assessed by examining the actual WSIs at the same suspected regions.
Comments on 'Fast and scalable search of whole-slide images via self-supervised deep learning'
Apr 18, 2023Abstract:Chen et al. [Chen2022] recently published the article 'Fast and scalable search of whole-slide images via self-supervised deep learning' in Nature Biomedical Engineering. The authors call their method 'self-supervised image search for histology', short SISH. We express our concerns that SISH is an incremental modification of Yottixel, has used MinMax binarization but does not cite the original works, and is based on a misnomer 'self-supervised image search'. As well, we point to several other concerns regarding experiments and comparisons performed by Chen et al.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge