Karthik Viswanathan
A transformer architecture alteration to incentivise externalised reasoning
Mar 22, 2026Abstract:We propose a new architectural change, and post-training pipeline, for making LLMs more verbose reasoners by teaching a model to truncate forward passes early. We augment an existing transformer architecture with an early-exit mechanism at intermediate layers and train the model to exit at shallower layers when the next token can be predicted without deep computation. After a calibration stage, we incentivise the model to exit as early as possible while maintaining task performance using reinforcement learning. We provide preliminary results to this effect for small reasoning models, showing that they learn to adaptively reduce computations across tokens. We predict that, applied at the right scale, our approach can minimise the amount of excess computation that reasoning models have at their disposal to perform non-myopic planning using their internal activations, reserving this only for difficult-to-predict tokens.
Probing Geometry of Next Token Prediction Using Cumulant Expansion of the Softmax Entropy
Oct 05, 2025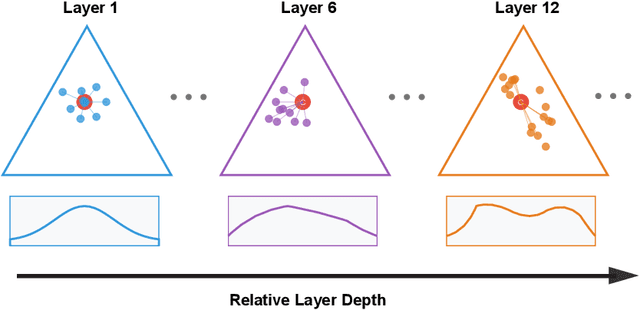
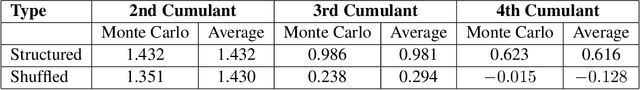
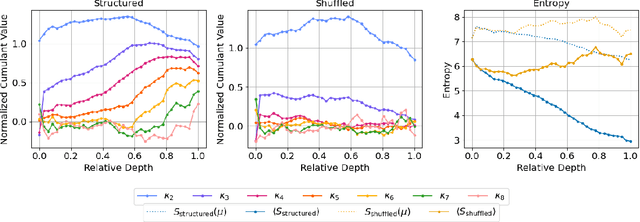
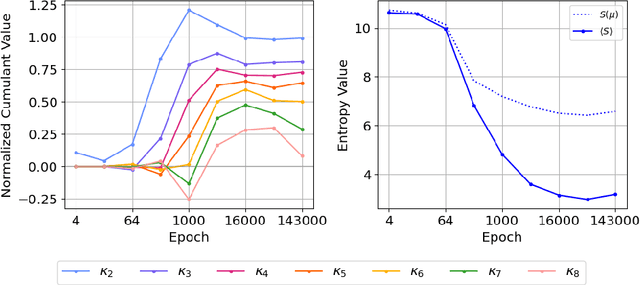
Abstract:We introduce a cumulant-expansion framework for quantifying how large language models (LLMs) internalize higher-order statistical structure during next-token prediction. By treating the softmax entropy of each layer's logit distribution as a perturbation around its "center" distribution, we derive closed-form cumulant observables that isolate successively higher-order correlations. Empirically, we track these cumulants in GPT-2 and Pythia models on Pile-10K prompts. (i) Structured prompts exhibit a characteristic rise-and-plateau profile across layers, whereas token-shuffled prompts remain flat, revealing the dependence of the cumulant profile on meaningful context. (ii) During training, all cumulants increase monotonically before saturating, directly visualizing the model's progression from capturing variance to learning skew, kurtosis, and higher-order statistical structures. (iii) Mathematical prompts show distinct cumulant signatures compared to general text, quantifying how models employ fundamentally different processing mechanisms for mathematical versus linguistic content. Together, these results establish cumulant analysis as a lightweight, mathematically grounded probe of feature-learning dynamics in high-dimensional neural networks.
Gene42: Long-Range Genomic Foundation Model With Dense Attention
Mar 20, 2025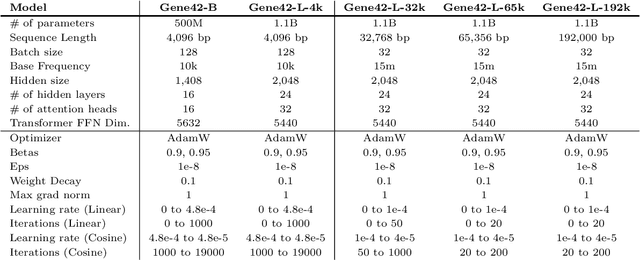
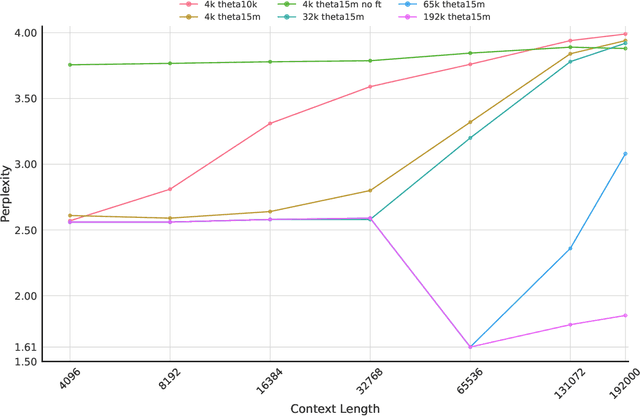
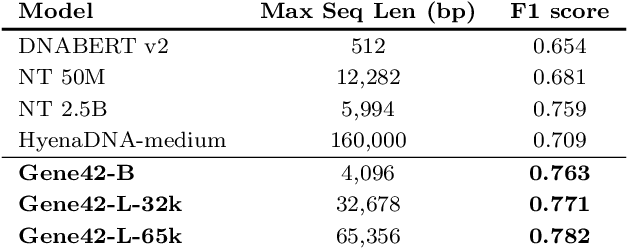
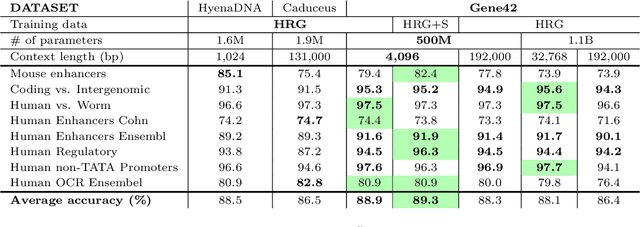
Abstract:We introduce Gene42, a novel family of Genomic Foundation Models (GFMs) designed to manage context lengths of up to 192,000 base pairs (bp) at a single-nucleotide resolution. Gene42 models utilize a decoder-only (LLaMA-style) architecture with a dense self-attention mechanism. Initially trained on fixed-length sequences of 4,096 bp, our models underwent continuous pretraining to extend the context length to 192,000 bp. This iterative extension allowed for the comprehensive processing of large-scale genomic data and the capture of intricate patterns and dependencies within the human genome. Gene42 is the first dense attention model capable of handling such extensive long context lengths in genomics, challenging state-space models that often rely on convolutional operators among other mechanisms. Our pretrained models exhibit notably low perplexity values and high reconstruction accuracy, highlighting their strong ability to model genomic data. Extensive experiments on various genomic benchmarks have demonstrated state-of-the-art performance across multiple tasks, including biotype classification, regulatory region identification, chromatin profiling prediction, variant pathogenicity prediction, and species classification. The models are publicly available at huggingface.co/inceptionai.
The Geometry of Tokens in Internal Representations of Large Language Models
Jan 17, 2025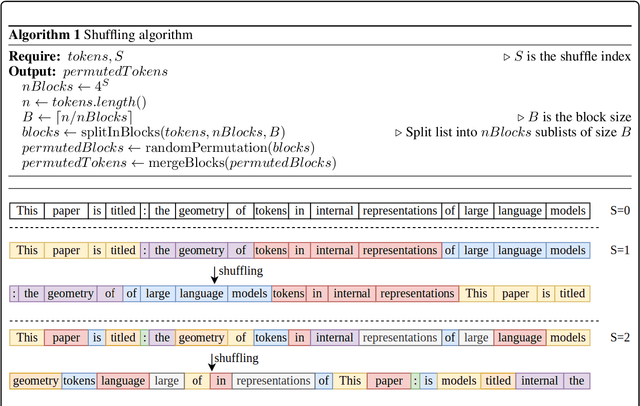
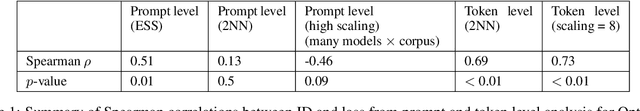
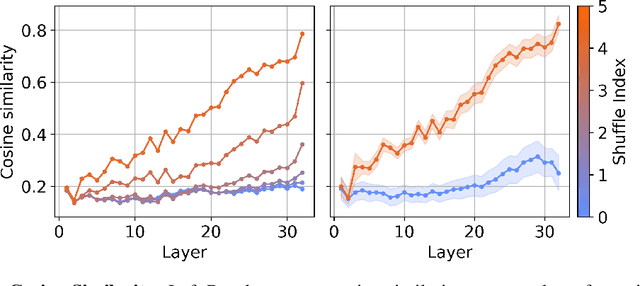
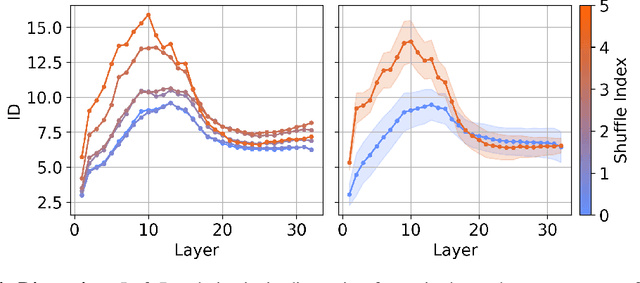
Abstract:We investigate the relationship between the geometry of token embeddings and their role in the next token prediction within transformer models. An important aspect of this connection uses the notion of empirical measure, which encodes the distribution of token point clouds across transformer layers and drives the evolution of token representations in the mean-field interacting picture. We use metrics such as intrinsic dimension, neighborhood overlap, and cosine similarity to observationally probe these empirical measures across layers. To validate our approach, we compare these metrics to a dataset where the tokens are shuffled, which disrupts the syntactic and semantic structure. Our findings reveal a correlation between the geometric properties of token embeddings and the cross-entropy loss of next token predictions, implying that prompts with higher loss values have tokens represented in higher-dimensional spaces.
Persistent Topological Features in Large Language Models
Oct 14, 2024Abstract:Understanding the decision-making processes of large language models (LLMs) is critical given their widespread applications. Towards this goal, describing the topological and geometrical properties of internal representations has recently provided valuable insights. For a more comprehensive characterization of these inherently complex spaces, we present a novel framework based on zigzag persistence, a method in topological data analysis (TDA) well-suited for describing data undergoing dynamic transformations across layers. Within this framework, we introduce persistence similarity, a new metric that quantifies the persistence and transformation of topological features such as $p$-cycles throughout the model layers. Unlike traditional similarity measures, our approach captures the entire evolutionary trajectory of these features, providing deeper insights into the internal workings of LLMs. As a practical application, we leverage persistence similarity to identify and prune redundant layers, demonstrating comparable performance to state-of-the-art methods across several benchmark datasets. Additionally, our analysis reveals consistent topological behaviors across various models and hyperparameter settings, suggesting a universal structure in LLM internal representations.
BraTS-PEDs: Results of the Multi-Consortium International Pediatric Brain Tumor Segmentation Challenge 2023
Jul 11, 2024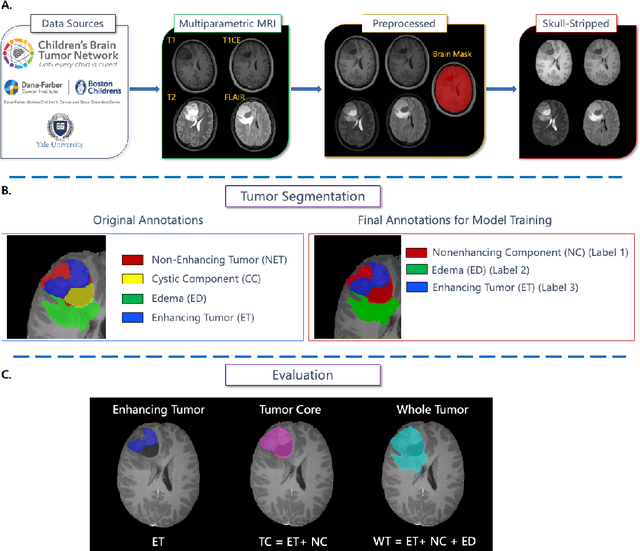

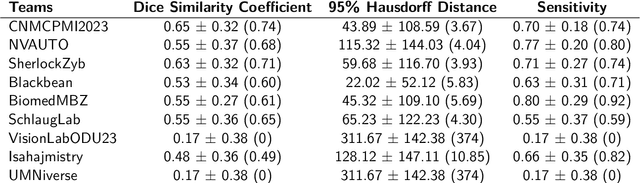
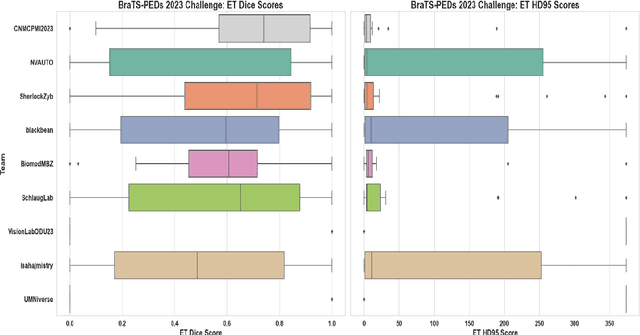
Abstract:Pediatric central nervous system tumors are the leading cause of cancer-related deaths in children. The five-year survival rate for high-grade glioma in children is less than 20%. The development of new treatments is dependent upon multi-institutional collaborative clinical trials requiring reproducible and accurate centralized response assessment. We present the results of the BraTS-PEDs 2023 challenge, the first Brain Tumor Segmentation (BraTS) challenge focused on pediatric brain tumors. This challenge utilized data acquired from multiple international consortia dedicated to pediatric neuro-oncology and clinical trials. BraTS-PEDs 2023 aimed to evaluate volumetric segmentation algorithms for pediatric brain gliomas from magnetic resonance imaging using standardized quantitative performance evaluation metrics employed across the BraTS 2023 challenges. The top-performing AI approaches for pediatric tumor analysis included ensembles of nnU-Net and Swin UNETR, Auto3DSeg, or nnU-Net with a self-supervised framework. The BraTSPEDs 2023 challenge fostered collaboration between clinicians (neuro-oncologists, neuroradiologists) and AI/imaging scientists, promoting faster data sharing and the development of automated volumetric analysis techniques. These advancements could significantly benefit clinical trials and improve the care of children with brain tumors.
The Brain Tumor Segmentation in Pediatrics (BraTS-PEDs) Challenge: Focus on Pediatrics (CBTN-CONNECT-DIPGR-ASNR-MICCAI BraTS-PEDs)
Apr 29, 2024Abstract:Pediatric tumors of the central nervous system are the most common cause of cancer-related death in children. The five-year survival rate for high-grade gliomas in children is less than 20%. Due to their rarity, the diagnosis of these entities is often delayed, their treatment is mainly based on historic treatment concepts, and clinical trials require multi-institutional collaborations. Here we present the CBTN-CONNECT-DIPGR-ASNR-MICCAI BraTS-PEDs challenge, focused on pediatric brain tumors with data acquired across multiple international consortia dedicated to pediatric neuro-oncology and clinical trials. The CBTN-CONNECT-DIPGR-ASNR-MICCAI BraTS-PEDs challenge brings together clinicians and AI/imaging scientists to lead to faster development of automated segmentation techniques that could benefit clinical trials, and ultimately the care of children with brain tumors.
Training and Comparison of nnU-Net and DeepMedic Methods for Autosegmentation of Pediatric Brain Tumors
Jan 30, 2024



Abstract:Brain tumors are the most common solid tumors and the leading cause of cancer-related death among children. Tumor segmentation is essential in surgical and treatment planning, and response assessment and monitoring. However, manual segmentation is time-consuming and has high inter-operator variability, underscoring the need for more efficient methods. We compared two deep learning-based 3D segmentation models, DeepMedic and nnU-Net, after training with pediatric-specific multi-institutional brain tumor data using based on multi-parametric MRI scans.Multi-parametric preoperative MRI scans of 339 pediatric patients (n=293 internal and n=46 external cohorts) with a variety of tumor subtypes, were preprocessed and manually segmented into four tumor subregions, i.e., enhancing tumor (ET), non-enhancing tumor (NET), cystic components (CC), and peritumoral edema (ED). After training, performance of the two models on internal and external test sets was evaluated using Dice scores, sensitivity, and Hausdorff distance with reference to ground truth manual segmentations. Dice score for nnU-Net internal test sets was (mean +/- SD (median)) 0.9+/-0.07 (0.94) for WT, 0.77+/-0.29 for ET, 0.66+/-0.32 for NET, 0.71+/-0.33 for CC, and 0.71+/-0.40 for ED, respectively. For DeepMedic the Dice scores were 0.82+/-0.16 for WT, 0.66+/-0.32 for ET, 0.48+/-0.27, for NET, 0.48+/-0.36 for CC, and 0.19+/-0.33 for ED, respectively. Dice scores were significantly higher for nnU-Net (p<=0.01). External validation of the trained nnU-Net model on the multi-institutional BraTS-PEDs 2023 dataset revealed high generalization capability in segmentation of whole tumor and tumor core with Dice scores of 0.87+/-0.13 (0.91) and 0.83+/-0.18 (0.89), respectively. Pediatric-specific data trained nnU-Net model is superior to DeepMedic for whole tumor and subregion segmentation of pediatric brain tumors.
A multi-institutional pediatric dataset of clinical radiology MRIs by the Children's Brain Tumor Network
Oct 02, 2023

Abstract:Pediatric brain and spinal cancers remain the leading cause of cancer-related death in children. Advancements in clinical decision-support in pediatric neuro-oncology utilizing the wealth of radiology imaging data collected through standard care, however, has significantly lagged other domains. Such data is ripe for use with predictive analytics such as artificial intelligence (AI) methods, which require large datasets. To address this unmet need, we provide a multi-institutional, large-scale pediatric dataset of 23,101 multi-parametric MRI exams acquired through routine care for 1,526 brain tumor patients, as part of the Children's Brain Tumor Network. This includes longitudinal MRIs across various cancer diagnoses, with associated patient-level clinical information, digital pathology slides, as well as tissue genotype and omics data. To facilitate downstream analysis, treatment-na\"ive images for 370 subjects were processed and released through the NCI Childhood Cancer Data Initiative via the Cancer Data Service. Through ongoing efforts to continuously build these imaging repositories, our aim is to accelerate discovery and translational AI models with real-world data, to ultimately empower precision medicine for children.
The Brain Tumor Segmentation Challenge 2023: Focus on Pediatrics
May 26, 2023Abstract:Pediatric tumors of the central nervous system are the most common cause of cancer-related death in children. The five-year survival rate for high-grade gliomas in children is less than 20\%. Due to their rarity, the diagnosis of these entities is often delayed, their treatment is mainly based on historic treatment concepts, and clinical trials require multi-institutional collaborations. The MICCAI Brain Tumor Segmentation (BraTS) Challenge is a landmark community benchmark event with a successful history of 12 years of resource creation for the segmentation and analysis of adult glioma. Here we present the CBTN-CONNECT-DIPGR-ASNR-MICCAI BraTS-PEDs 2023 challenge, which represents the first BraTS challenge focused on pediatric brain tumors with data acquired across multiple international consortia dedicated to pediatric neuro-oncology and clinical trials. The BraTS-PEDs 2023 challenge focuses on benchmarking the development of volumentric segmentation algorithms for pediatric brain glioma through standardized quantitative performance evaluation metrics utilized across the BraTS 2023 cluster of challenges. Models gaining knowledge from the BraTS-PEDs multi-parametric structural MRI (mpMRI) training data will be evaluated on separate validation and unseen test mpMRI dataof high-grade pediatric glioma. The CBTN-CONNECT-DIPGR-ASNR-MICCAI BraTS-PEDs 2023 challenge brings together clinicians and AI/imaging scientists to lead to faster development of automated segmentation techniques that could benefit clinical trials, and ultimately the care of children with brain tumors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge