Joseph D. Viviano
A scalable gene network model of regulatory dynamics in single cells
Mar 25, 2025Abstract:Single-cell data provide high-dimensional measurements of the transcriptional states of cells, but extracting insights into the regulatory functions of genes, particularly identifying transcriptional mechanisms affected by biological perturbations, remains a challenge. Many perturbations induce compensatory cellular responses, making it difficult to distinguish direct from indirect effects on gene regulation. Modeling how gene regulatory functions shape the temporal dynamics of these responses is key to improving our understanding of biological perturbations. Dynamical models based on differential equations offer a principled way to capture transcriptional dynamics, but their application to single-cell data has been hindered by computational constraints, stochasticity, sparsity, and noise. Existing methods either rely on low-dimensional representations or make strong simplifying assumptions, limiting their ability to model transcriptional dynamics at scale. We introduce a Functional and Learnable model of Cell dynamicS, FLeCS, that incorporates gene network structure into coupled differential equations to model gene regulatory functions. Given (pseudo)time-series single-cell data, FLeCS accurately infers cell dynamics at scale, provides improved functional insights into transcriptional mechanisms perturbed by gene knockouts, both in myeloid differentiation and K562 Perturb-seq experiments, and simulates single-cell trajectories of A549 cells following small-molecule perturbations.
torchgfn: A PyTorch GFlowNet library
May 24, 2023
Abstract:The increasing popularity of generative flow networks (GFlowNets or GFNs) is accompanied with a proliferation of code sources. This hinders the implementation of new features, such as training losses, that can readily be compared to existing ones, on a set of common environments. In addition to slowing down research in the field of GFlowNets, different code bases use different conventions, that might be confusing for newcomers. `torchgfn` is a library built on top of PyTorch, that aims at addressing both problems. It provides user with a simple API for environments, and useful abstractions for samplers and losses. Multiple examples are provided, replicating published results. The code is available in https://github.com/saleml/torchgfn.
TorchXRayVision: A library of chest X-ray datasets and models
Oct 31, 2021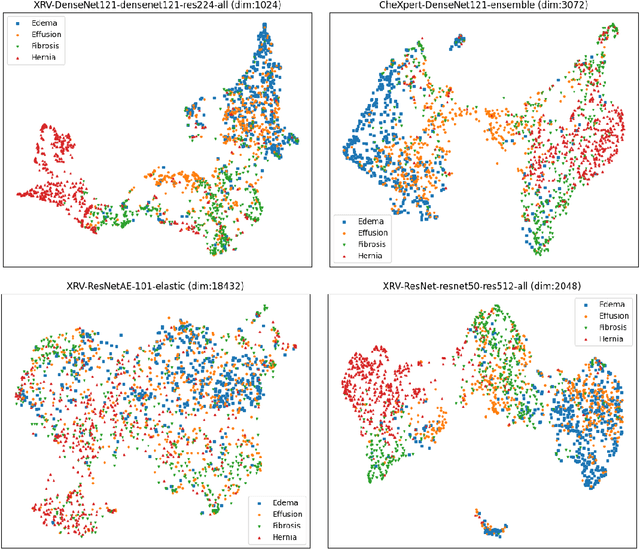
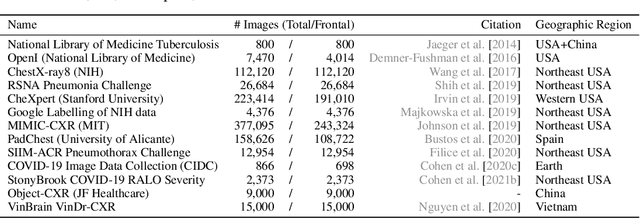
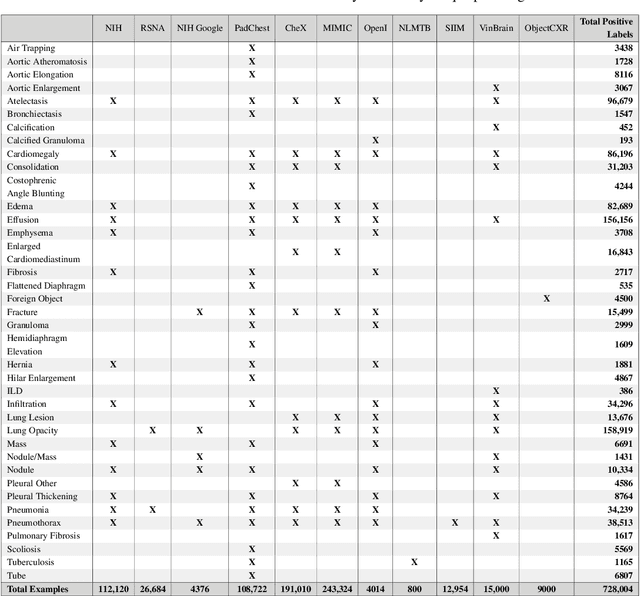
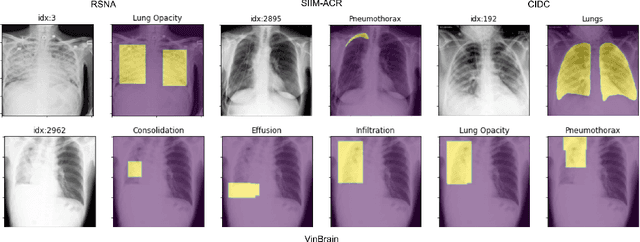
Abstract:TorchXRayVision is an open source software library for working with chest X-ray datasets and deep learning models. It provides a common interface and common pre-processing chain for a wide set of publicly available chest X-ray datasets. In addition, a number of classification and representation learning models with different architectures, trained on different data combinations, are available through the library to serve as baselines or feature extractors.
What's in the Box? A Preliminary Analysis of Undesirable Content in the Common Crawl Corpus
May 31, 2021
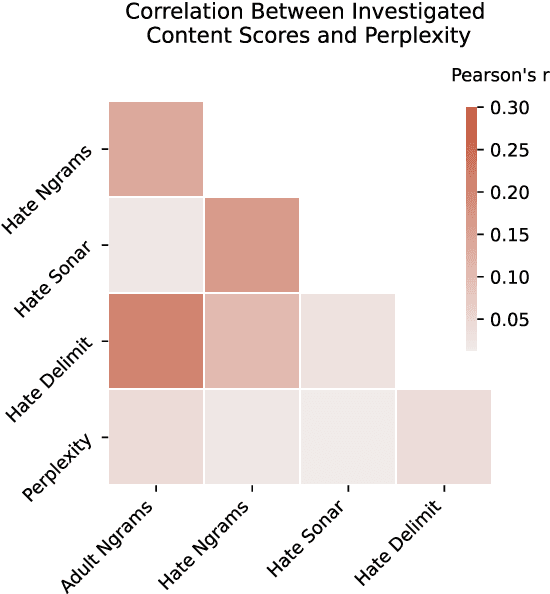

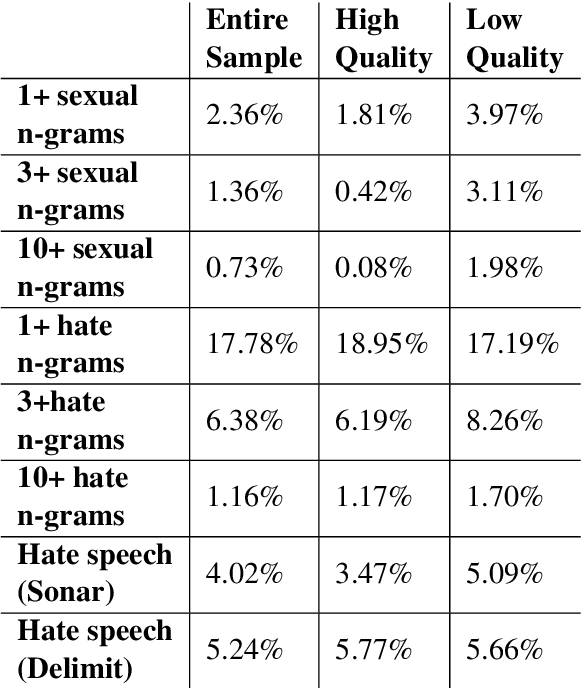
Abstract:Whereas much of the success of the current generation of neural language models has been driven by increasingly large training corpora, relatively little research has been dedicated to analyzing these massive sources of textual data. In this exploratory analysis, we delve deeper into the Common Crawl, a colossal web corpus that is extensively used for training language models. We find that it contains a significant amount of undesirable content, including hate speech and sexually explicit content, even after filtering procedures. We discuss the potential impacts of this content on language models and conclude with future research directions and a more mindful approach to corpus collection and analysis.
Underwhelming Generalization Improvements From Controlling Feature Attribution
Oct 01, 2019
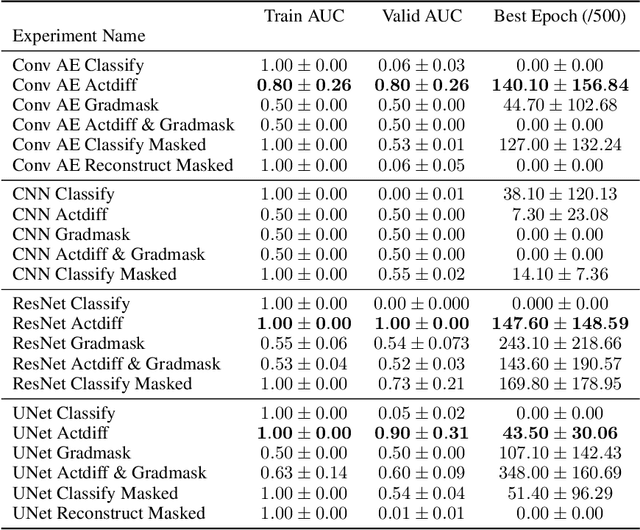
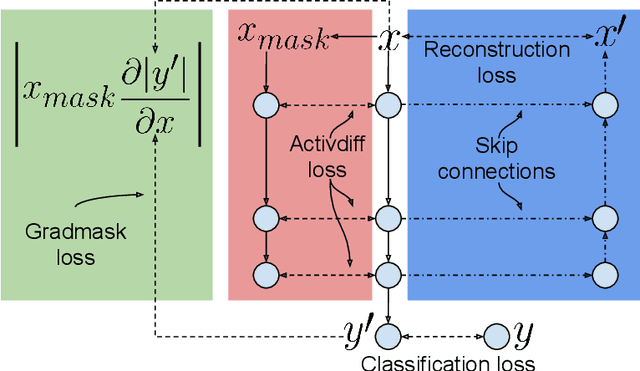
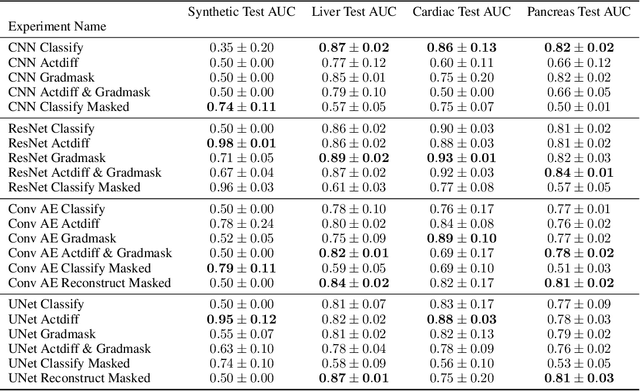
Abstract:Overfitting is a common issue in machine learning, which can arise when the model learns to predict class membership using convenient but spuriously-correlated image features instead of the true image features that denote a class. These are typically visualized using saliency maps. In some object classification tasks such as for medical images, one may have some images with masks, indicating a region of interest, i.e., which part of the image contains the most relevant information for the classification. We describe a simple method for taking advantage of such auxiliary labels, by training networks to ignore the distracting features which may be extracted outside of the region of interest, on the training images for which such masks are available. This mask information is only used during training and has an impact on generalization accuracy in a dataset-dependent way. We observe an underwhelming relationship between controlling saliency maps and improving generalization performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge