João Pedrosa
MedShapeNet -- A Large-Scale Dataset of 3D Medical Shapes for Computer Vision
Sep 12, 2023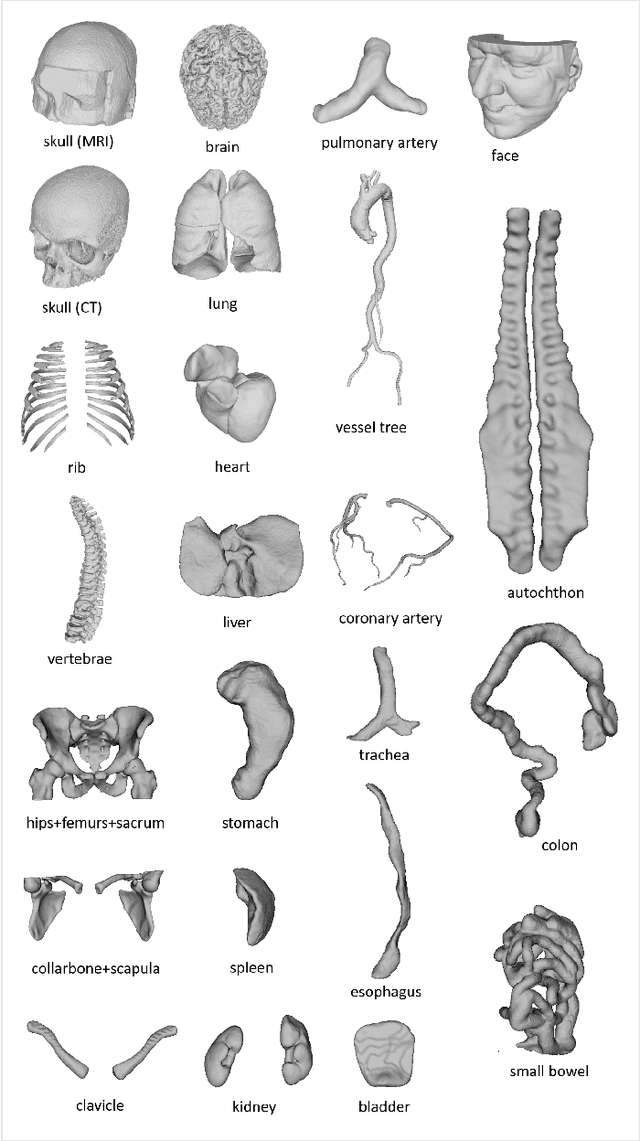

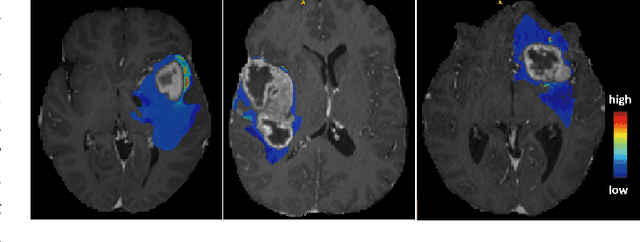
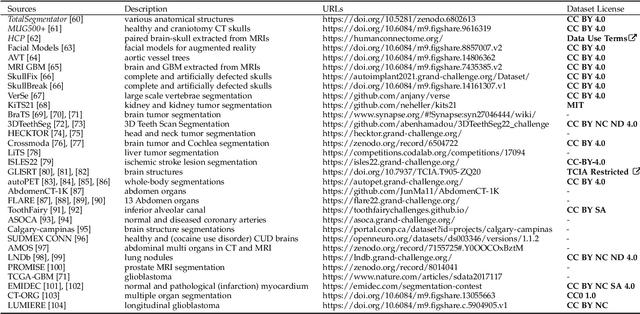
Abstract:We present MedShapeNet, a large collection of anatomical shapes (e.g., bones, organs, vessels) and 3D surgical instrument models. Prior to the deep learning era, the broad application of statistical shape models (SSMs) in medical image analysis is evidence that shapes have been commonly used to describe medical data. Nowadays, however, state-of-the-art (SOTA) deep learning algorithms in medical imaging are predominantly voxel-based. In computer vision, on the contrary, shapes (including, voxel occupancy grids, meshes, point clouds and implicit surface models) are preferred data representations in 3D, as seen from the numerous shape-related publications in premier vision conferences, such as the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), as well as the increasing popularity of ShapeNet (about 51,300 models) and Princeton ModelNet (127,915 models) in computer vision research. MedShapeNet is created as an alternative to these commonly used shape benchmarks to facilitate the translation of data-driven vision algorithms to medical applications, and it extends the opportunities to adapt SOTA vision algorithms to solve critical medical problems. Besides, the majority of the medical shapes in MedShapeNet are modeled directly on the imaging data of real patients, and therefore it complements well existing shape benchmarks comprising of computer-aided design (CAD) models. MedShapeNet currently includes more than 100,000 medical shapes, and provides annotations in the form of paired data. It is therefore also a freely available repository of 3D models for extended reality (virtual reality - VR, augmented reality - AR, mixed reality - MR) and medical 3D printing. This white paper describes in detail the motivations behind MedShapeNet, the shape acquisition procedures, the use cases, as well as the usage of the online shape search portal: https://medshapenet.ikim.nrw/
DEEPBEAS3D: Deep Learning and B-Spline Explicit Active Surfaces
Sep 05, 2023Abstract:Deep learning-based automatic segmentation methods have become state-of-the-art. However, they are often not robust enough for direct clinical application, as domain shifts between training and testing data affect their performance. Failure in automatic segmentation can cause sub-optimal results that require correction. To address these problems, we propose a novel 3D extension of an interactive segmentation framework that represents a segmentation from a convolutional neural network (CNN) as a B-spline explicit active surface (BEAS). BEAS ensures segmentations are smooth in 3D space, increasing anatomical plausibility, while allowing the user to precisely edit the 3D surface. We apply this framework to the task of 3D segmentation of the anal sphincter complex (AS) from transperineal ultrasound (TPUS) images, and compare it to the clinical tool used in the pelvic floor disorder clinic (4D View VOCAL, GE Healthcare; Zipf, Austria). Experimental results show that: 1) the proposed framework gives the user explicit control of the surface contour; 2) the perceived workload calculated via the NASA-TLX index was reduced by 30% compared to VOCAL; and 3) it required 7 0% (170 seconds) less user time than VOCAL (p< 0.00001)
Interactive Segmentation via Deep Learning and B-Spline Explicit Active Surfaces
Oct 25, 2021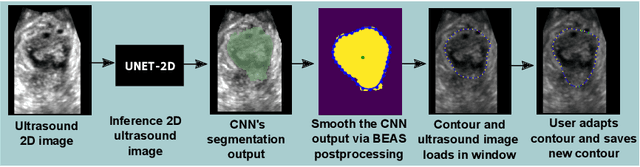
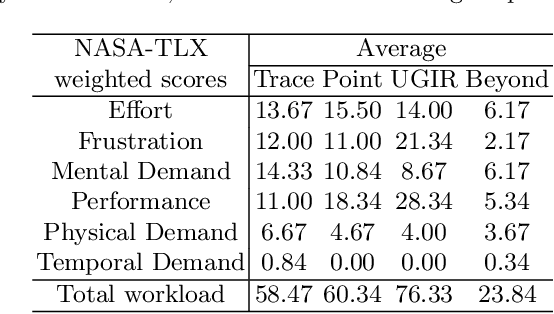
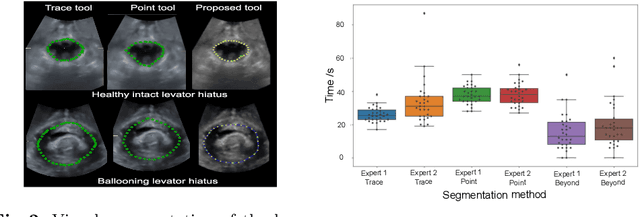
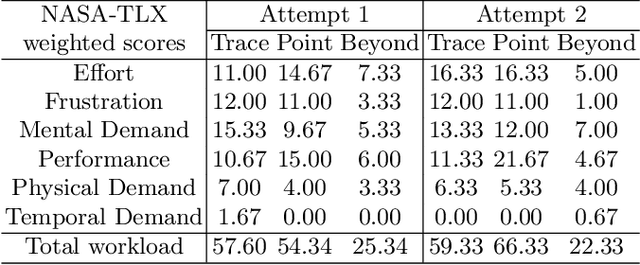
Abstract:Automatic medical image segmentation via convolutional neural networks (CNNs) has shown promising results. However, they may not always be robust enough for clinical use. Sub-optimal segmentation would require clinician's to manually delineate the target object, causing frustration. To address this problem, a novel interactive CNN-based segmentation framework is proposed in this work. The aim is to represent the CNN segmentation contour as B-splines by utilising B-spline explicit active surfaces (BEAS). The interactive element of the framework allows the user to precisely edit the contour in real-time, and by utilising BEAS it ensures the final contour is smooth and anatomically plausible. This framework was applied to the task of 2D segmentation of the levator hiatus from 2D ultrasound (US) images, and compared to the current clinical tools used in pelvic floor disorder clinic (4DView, GE Healthcare; Zipf, Austria). Experimental results show that: 1) the proposed framework is more robust than current state-of-the-art CNNs; 2) the perceived workload calculated via the NASA-TLX index was reduced more than half for the proposed approach in comparison to current clinical tools; and 3) the proposed tool requires at least 13 seconds less user time than the clinical tools, which was significant (p=0.001).
* 11 pages, 3 figures, 2 tables
LNDb: A Lung Nodule Database on Computed Tomography
Dec 19, 2019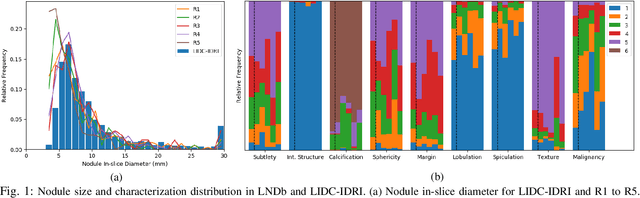
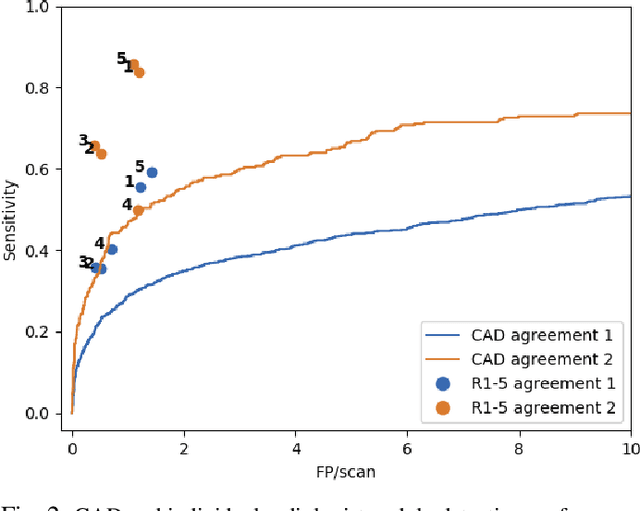
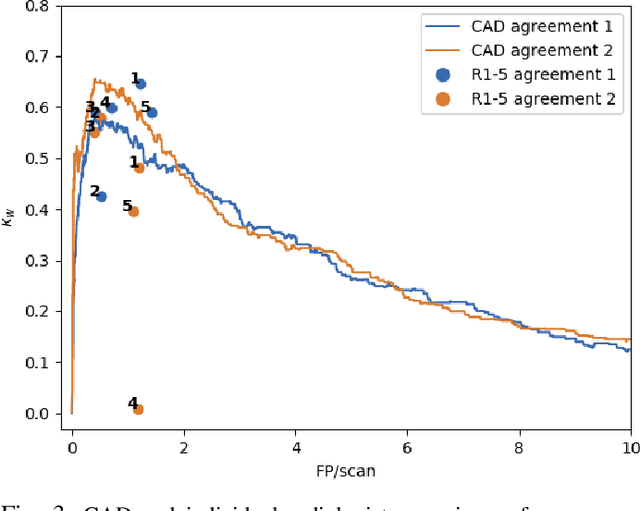
Abstract:Lung cancer is the deadliest type of cancer worldwide and late detection is the major factor for the low survival rate of patients. Low dose computed tomography has been suggested as a potential screening tool but manual screening is costly, time-consuming and prone to variability. This has fueled the development of automatic methods for the detection, segmentation and characterisation of pulmonary nodules but its application to clinical routine is challenging. In this study, a new database for the development and testing of pulmonary nodule computer-aided strategies is presented which intends to complement current databases by giving additional focus to radiologist variability and local clinical reality. State-of-the-art nodule detection, segmentation and characterization methods are tested and compared to manual annotations as well as collaborative strategies combining multiple radiologists and radiologists and computer-aided systems. It is shown that state-of-the-art methodologies can determine a patient's follow-up recommendation as accurately as a radiologist, though the nodule detection method used shows decreased performance in this database.
Did you miss it? Automatic lung nodule detection combined with gaze information improves radiologists' screening performance
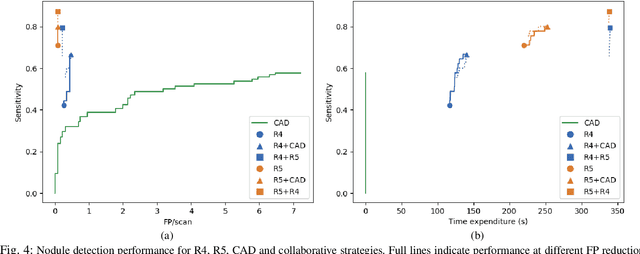
Oct 09, 2019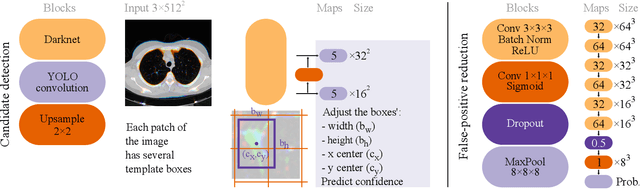
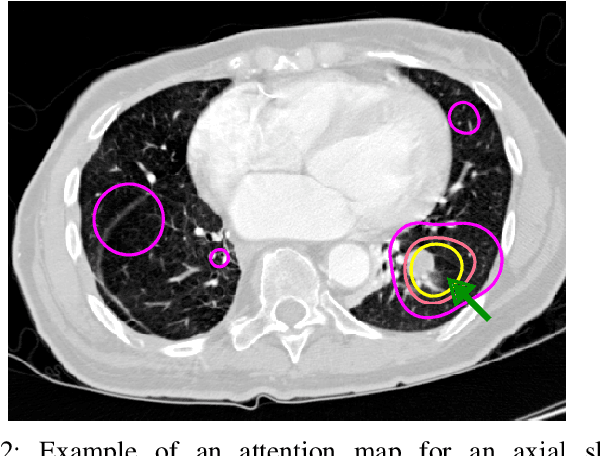
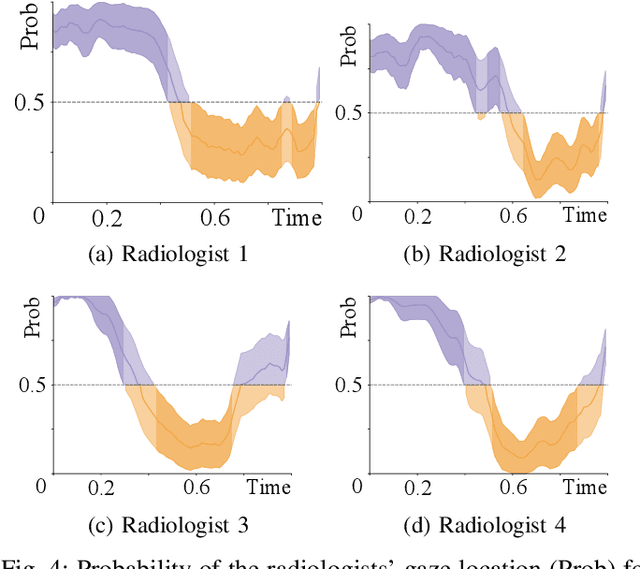
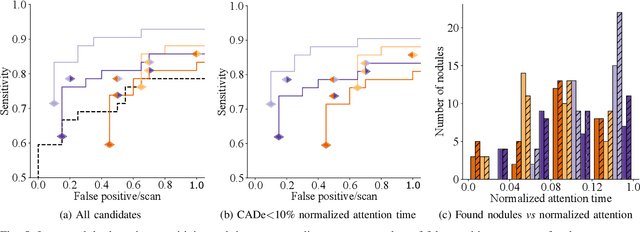
Abstract:Early diagnosis of lung cancer via computed tomography can significantly reduce the morbidity and mortality rates associated with the pathology. However, search lung nodules is a high complexity task, which affects the success of screening programs. Whilst computer-aided detection systems can be used as second observers, they may bias radiologists and introduce significant time overheads. With this in mind, this study assesses the potential of using gaze information for integrating automatic detection systems in the clinical practice. For that purpose, 4 radiologists were asked to annotate 20 scans from a public dataset while being monitored by an eye tracker device and an automatic lung nodule detection system was developed. Our results show that radiologists follow a similar search routine and tend to have lower fixation periods in regions where finding errors occur. The overall detection sensitivity of the specialists was 0.67$\pm$0.07, whereas the system achieved 0.69. Combining the annotations of one radiologist with the automatic system significantly improves the detection performance to similar levels of two annotators. Likewise, combining the findings of radiologist with the detection algorithm only for low fixation regions still significantly improves the detection sensitivity without increasing the number of false-positives. The combination of the automatic system with the gaze information allows to mitigate possible errors of the radiologist without some of the issues usually associated with automatic detection system.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge