James G. Terry
Pseudo-Label Guided Multi-Contrast Generalization for Non-Contrast Organ-Aware Segmentation
May 12, 2022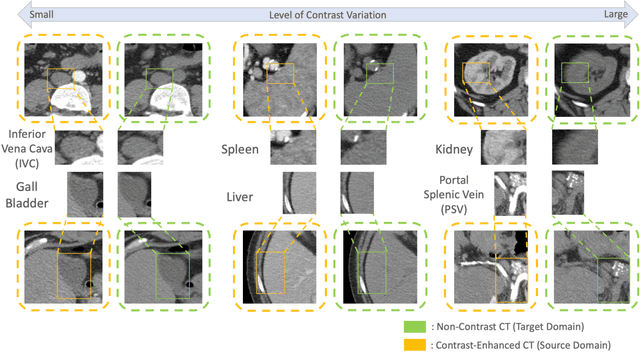

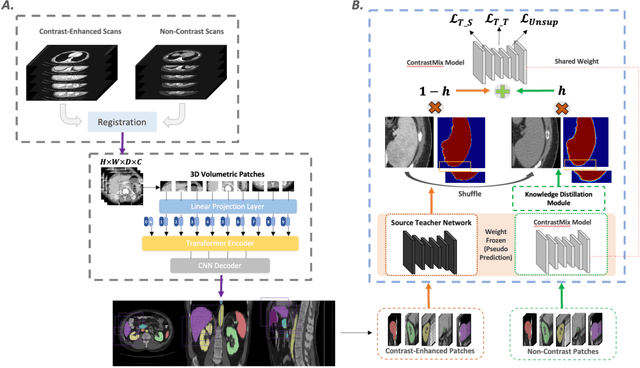

Abstract:Non-contrast computed tomography (NCCT) is commonly acquired for lung cancer screening, assessment of general abdominal pain or suspected renal stones, trauma evaluation, and many other indications. However, the absence of contrast limits distinguishing organ in-between boundaries. In this paper, we propose a novel unsupervised approach that leverages pairwise contrast-enhanced CT (CECT) context to compute non-contrast segmentation without ground-truth label. Unlike generative adversarial approaches, we compute the pairwise morphological context with CECT to provide teacher guidance instead of generating fake anatomical context. Additionally, we further augment the intensity correlations in 'organ-specific' settings and increase the sensitivity to organ-aware boundary. We validate our approach on multi-organ segmentation with paired non-contrast & contrast-enhanced CT scans using five-fold cross-validation. Full external validations are performed on an independent non-contrast cohort for aorta segmentation. Compared with current abdominal organs segmentation state-of-the-art in fully supervised setting, our proposed pipeline achieves a significantly higher Dice by 3.98% (internal multi-organ annotated), and 8.00% (external aorta annotated) for abdominal organs segmentation. The code and pretrained models are publicly available at https://github.com/MASILab/ContrastMix.
Stochastic tissue window normalization of deep learning on computed tomography
Dec 01, 2019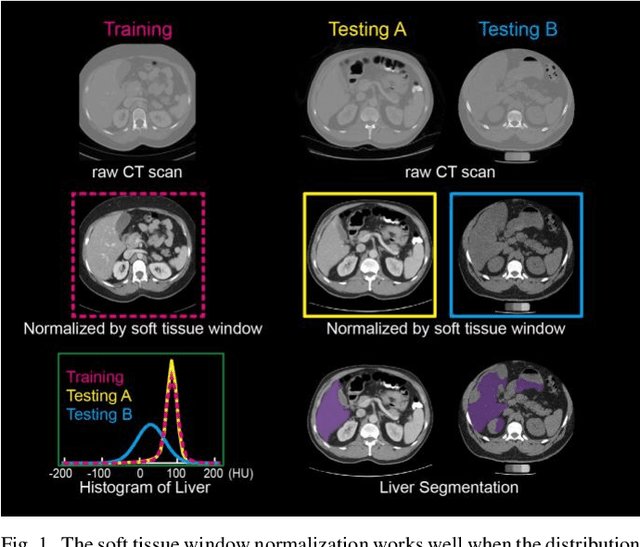
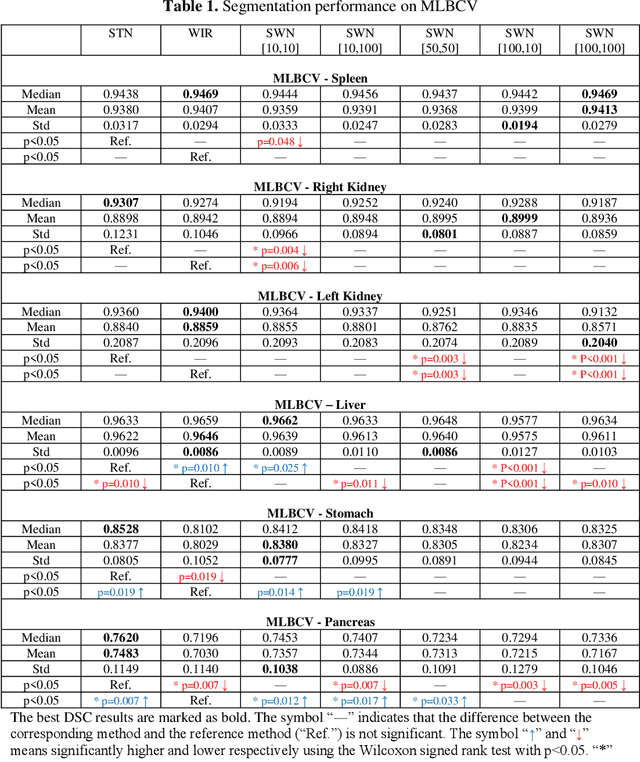
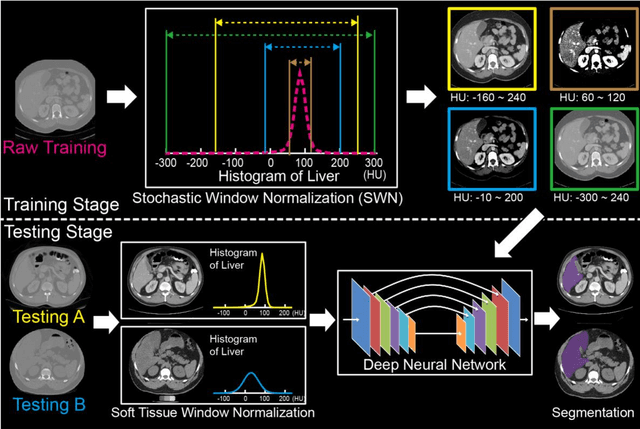
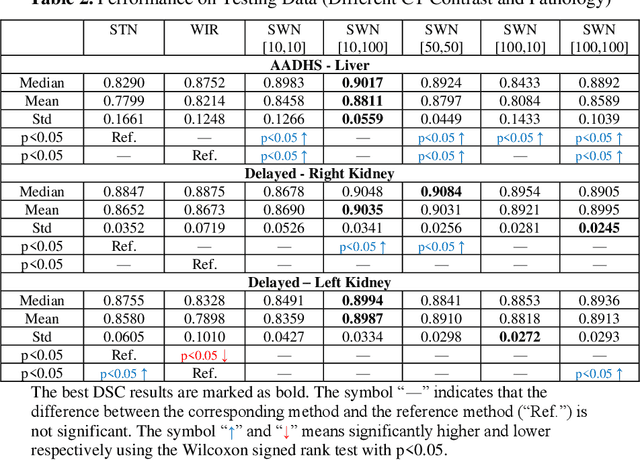
Abstract:Tissue window filtering has been widely used in deep learning for computed tomography (CT) image analyses to improve training performance (e.g., soft tissue windows for abdominal CT). However, the effectiveness of tissue window normalization is questionable since the generalizability of the trained model might be further harmed, especially when such models are applied to new cohorts with different CT reconstruction kernels, contrast mechanisms, dynamic variations in the acquisition, and physiological changes. We evaluate the effectiveness of both with and without using soft tissue window normalization on multisite CT cohorts. Moreover, we propose a stochastic tissue window normalization (SWN) method to improve the generalizability of tissue window normalization. Different from the random sampling, the SWN method centers the randomization around the soft tissue window to maintain the specificity for abdominal organs. To evaluate the performance of different strategies, 80 training and 453 validation and testing scans from six datasets are employed to perform multi-organ segmentation using standard 2D U-Net. The six datasets cover the scenarios, where the training and testing scans are from (1) same scanner and same population, (2) same CT contrast but different pathology, and (3) different CT contrast and pathology. The traditional soft tissue window and nonwindowed approaches achieved better performance on (1). The proposed SWN achieved general superior performance on (2) and (3) with statistical analyses, which offers better generalizability for a trained model.
Fully Automatic Liver Attenuation Estimation Combing CNN Segmentation and Morphological Operations
Jun 29, 2019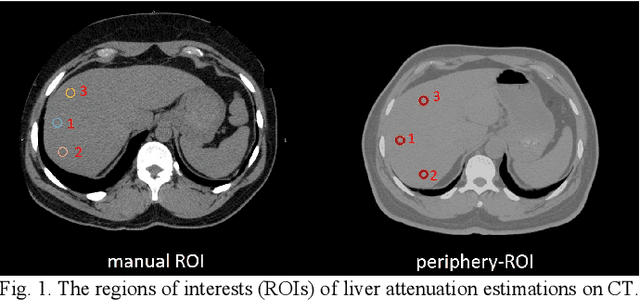
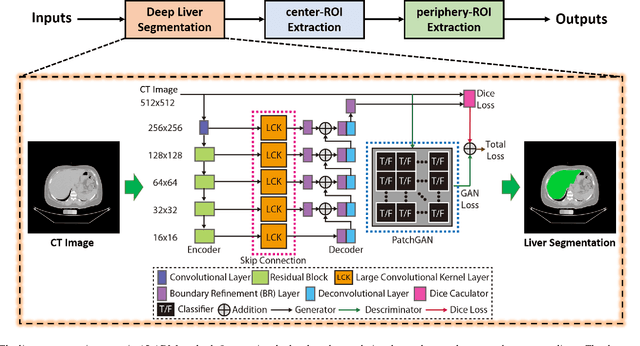
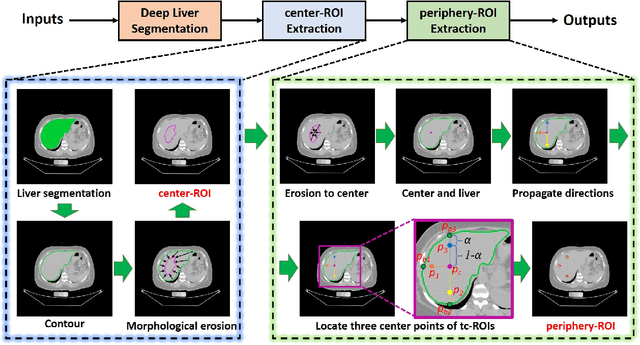
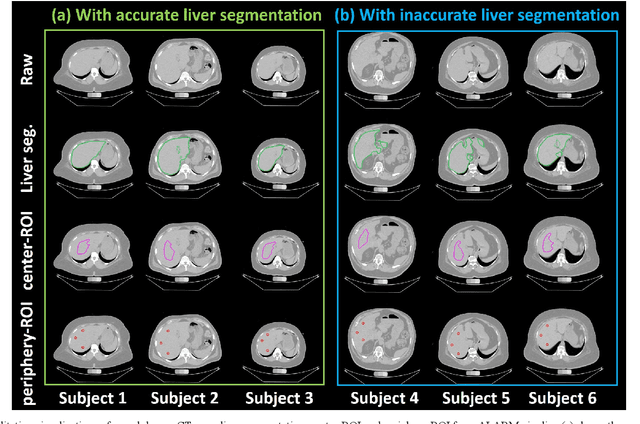
Abstract:Manually tracing regions of interest (ROIs) within the liver is the de facto standard method for measuring liver attenuation on computed tomography (CT) in diagnosing nonalcoholic fatty liver disease (NAFLD). However, manual tracing is resource intensive. To address these limitations and to expand the availability of a quantitative CT measure of hepatic steatosis, we propose the automatic liver attenuation ROI-based measurement (ALARM) method for automated liver attenuation estimation. The ALARM method consists of two major stages: (1) deep convolutional neural network (DCNN)-based liver segmentation and (2) automated ROI extraction. First, liver segmentation was achieved using our previously developed SS-Net. Then, a single central ROI (center-ROI) and three circles ROI (periphery-ROI) were computed based on liver segmentation and morphological operations. The ALARM method is available as an open source Docker container (https://github.com/MASILab/ALARM).246 subjects with 738 abdomen CT scans from the African American-Diabetes Heart Study (AA-DHS) were used for external validation (testing), independent from the training and validation cohort (100 clinically acquired CT abdominal scans).
Coronary Calcium Detection using 3D Attention Identical Dual Deep Network Based on Weakly Supervised Learning
Nov 10, 2018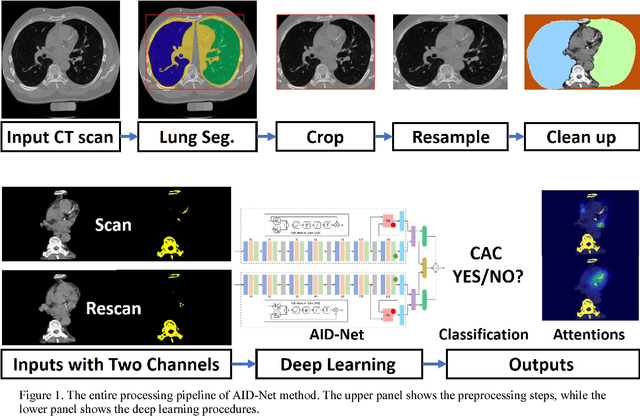
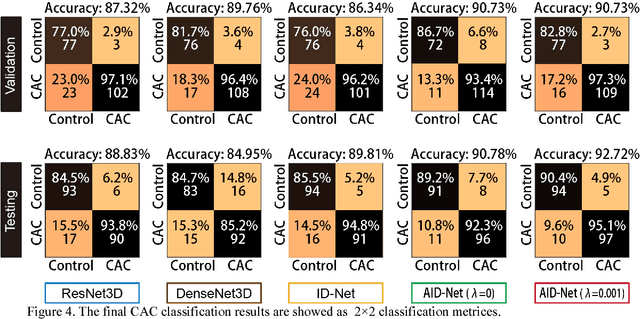
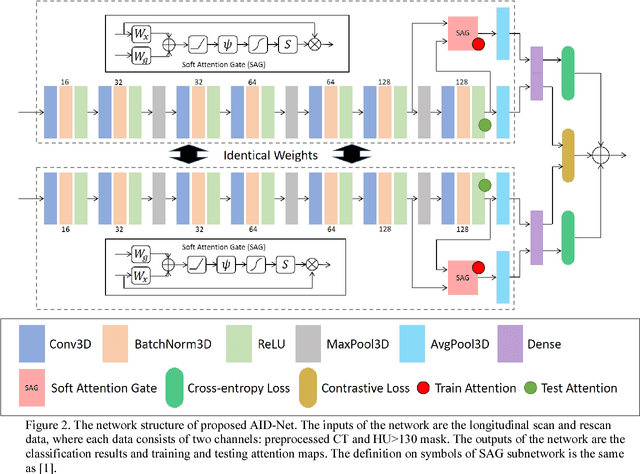


Abstract:Coronary artery calcium (CAC) is biomarker of advanced subclinical coronary artery disease and predicts myocardial infarction and death prior to age 60 years. The slice-wise manual delineation has been regarded as the gold standard of coronary calcium detection. However, manual efforts are time and resource consuming and even impracticable to be applied on large-scale cohorts. In this paper, we propose the attention identical dual network (AID-Net) to perform CAC detection using scan-rescan longitudinal non-contrast CT scans with weakly supervised attention by only using per scan level labels. To leverage the performance, 3D attention mechanisms were integrated into the AID-Net to provide complementary information for classification tasks. Moreover, the 3D Gradient-weighted Class Activation Mapping (Grad-CAM) was also proposed at the testing stage to interpret the behaviors of the deep neural network. 5075 non-contrast chest CT scans were used as training, validation and testing datasets. Baseline performance was assessed on the same cohort. From the results, the proposed AID-Net achieved the superior performance on classification accuracy (0.9272) and AUC (0.9627).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge