Hideaki Suzuki
Division of Brain Sciences, Dept. of Medicine, Imperial College London
Automated Quality Control in Image Segmentation: Application to the UK Biobank Cardiac MR Imaging Study
Jan 27, 2019
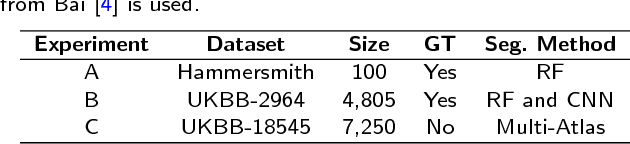
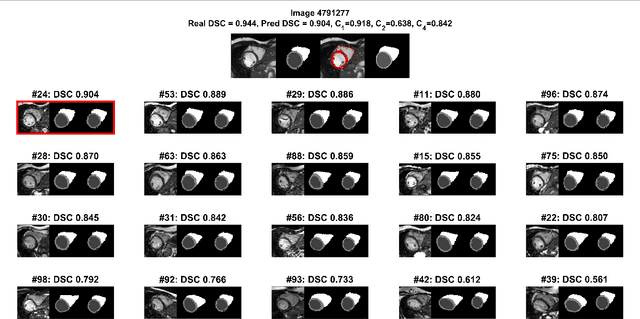
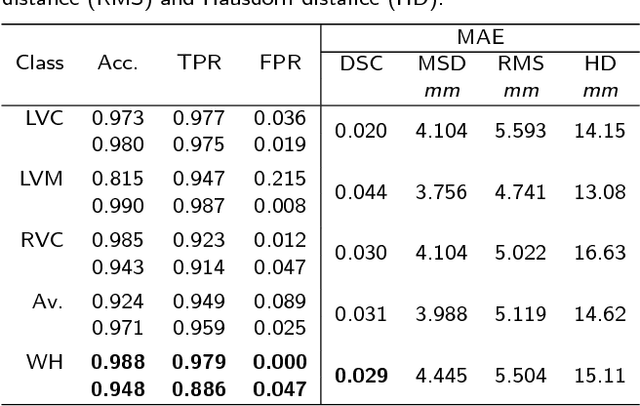
Abstract:Background: The trend towards large-scale studies including population imaging poses new challenges in terms of quality control (QC). This is a particular issue when automatic processing tools, e.g. image segmentation methods, are employed to derive quantitative measures or biomarkers for later analyses. Manual inspection and visual QC of each segmentation isn't feasible at large scale. However, it's important to be able to automatically detect when a segmentation method fails so as to avoid inclusion of wrong measurements into subsequent analyses which could lead to incorrect conclusions. Methods: To overcome this challenge, we explore an approach for predicting segmentation quality based on Reverse Classification Accuracy, which enables us to discriminate between successful and failed segmentations on a per-cases basis. We validate this approach on a new, large-scale manually-annotated set of 4,800 cardiac magnetic resonance scans. We then apply our method to a large cohort of 7,250 cardiac MRI on which we have performed manual QC. Results: We report results used for predicting segmentation quality metrics including Dice Similarity Coefficient (DSC) and surface-distance measures. As initial validation, we present data for 400 scans demonstrating 99% accuracy for classifying low and high quality segmentations using predicted DSC scores. As further validation we show high correlation between real and predicted scores and 95% classification accuracy on 4,800 scans for which manual segmentations were available. We mimic real-world application of the method on 7,250 cardiac MRI where we show good agreement between predicted quality metrics and manual visual QC scores. Conclusions: We show that RCA has the potential for accurate and fully automatic segmentation QC on a per-case basis in the context of large-scale population imaging as in the UK Biobank Imaging Study.
A Comprehensive Approach for Learning-based Fully-Automated Inter-slice Motion Correction for Short-Axis Cine Cardiac MR Image Stacks
Oct 03, 2018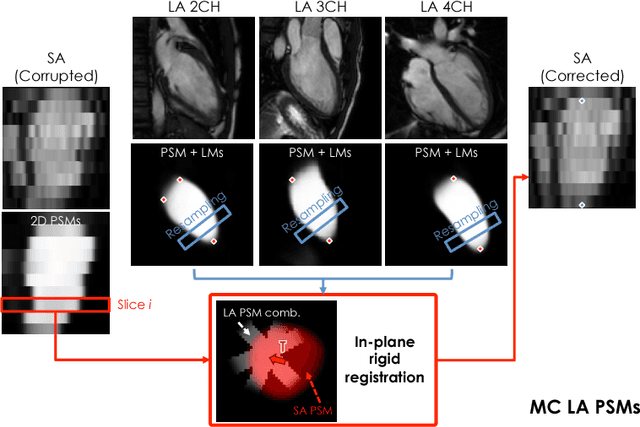
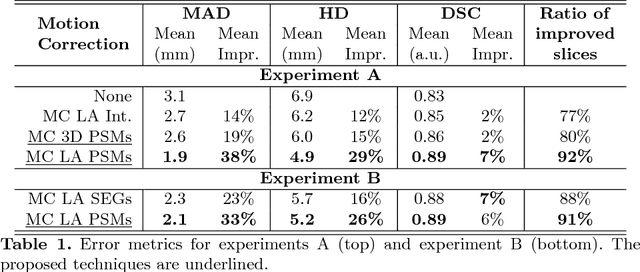
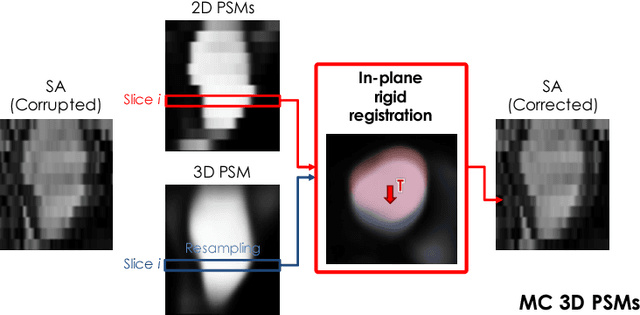
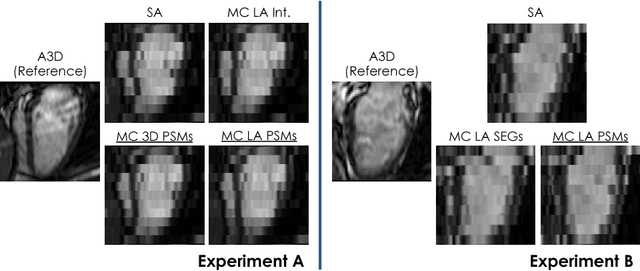
Abstract:In the clinical routine, short axis (SA) cine cardiac MR (CMR) image stacks are acquired during multiple subsequent breath-holds. If the patient cannot consistently hold the breath at the same position, the acquired image stack will be affected by inter-slice respiratory motion and will not correctly represent the cardiac volume, introducing potential errors in the following analyses and visualisations. We propose an approach to automatically correct inter-slice respiratory motion in SA CMR image stacks. Our approach makes use of probabilistic segmentation maps (PSMs) of the left ventricular (LV) cavity generated with decision forests. PSMs are generated for each slice of the SA stack and rigidly registered in-plane to a target PSM. If long axis (LA) images are available, PSMs are generated for them and combined to create the target PSM; if not, the target PSM is produced from the same stack using a 3D model trained from motion-free stacks. The proposed approach was tested on a dataset of SA stacks acquired from 24 healthy subjects (for which anatomical 3D cardiac images were also available as reference) and compared to two techniques which use LA intensity images and LA segmentations as targets, respectively. The results show the accuracy and robustness of the proposed approach in motion compensation.
Learning-Based Quality Control for Cardiac MR Images
Sep 15, 2018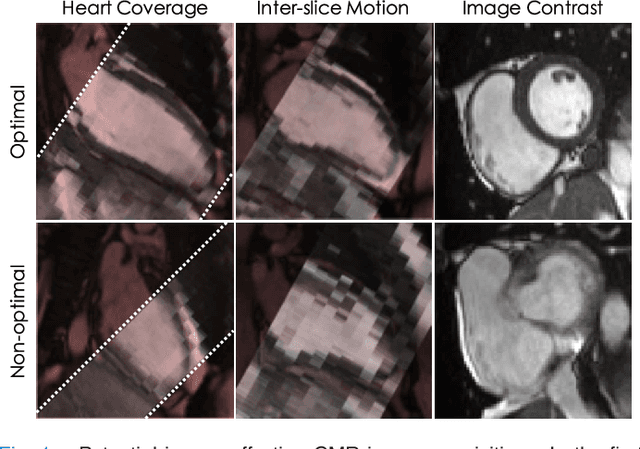
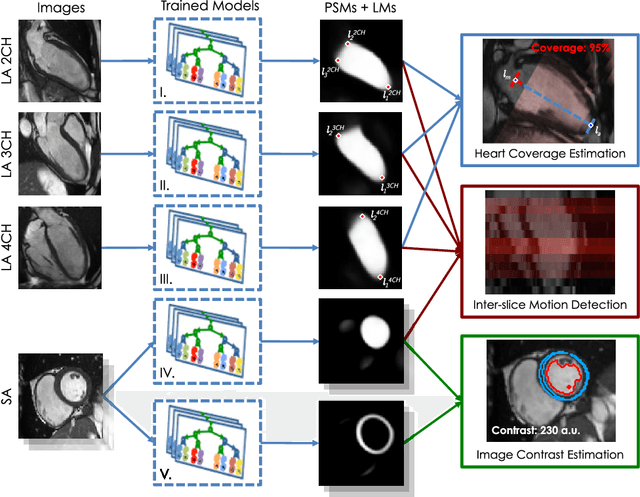
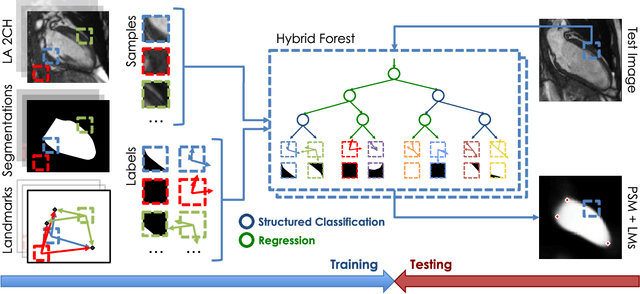
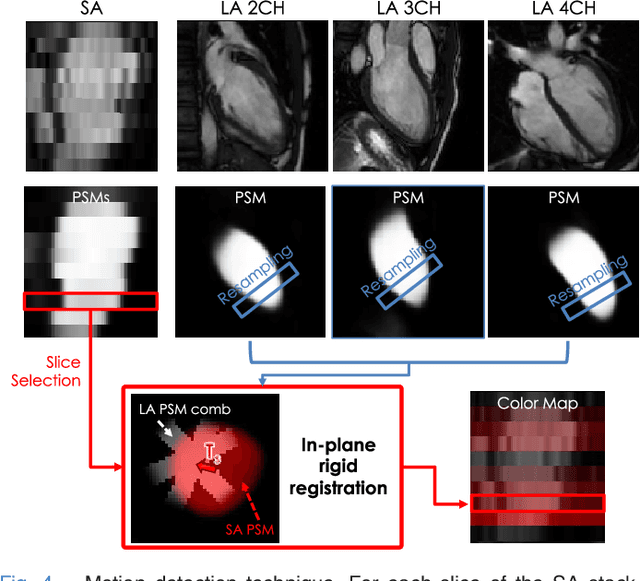
Abstract:The effectiveness of a cardiovascular magnetic resonance (CMR) scan depends on the ability of the operator to correctly tune the acquisition parameters to the subject being scanned and on the potential occurrence of imaging artefacts such as cardiac and respiratory motion. In the clinical practice, a quality control step is performed by visual assessment of the acquired images: however, this procedure is strongly operator-dependent, cumbersome and sometimes incompatible with the time constraints in clinical settings and large-scale studies. We propose a fast, fully-automated, learning-based quality control pipeline for CMR images, specifically for short-axis image stacks. Our pipeline performs three important quality checks: 1) heart coverage estimation, 2) inter-slice motion detection, 3) image contrast estimation in the cardiac region. The pipeline uses a hybrid decision forest method - integrating both regression and structured classification models - to extract landmarks as well as probabilistic segmentation maps from both long- and short-axis images as a basis to perform the quality checks. The technique was tested on up to 3000 cases from the UK Biobank as well as on 100 cases from the UK Digital Heart Project, and validated against manual annotations and visual inspections performed by expert interpreters. The results show the capability of the proposed pipeline to correctly detect incomplete or corrupted scans (e.g. on UK Biobank, sensitivity and specificity respectively 88% and 99% for heart coverage estimation, 85% and 95% for motion detection), allowing their exclusion from the analysed dataset or the triggering of a new acquisition.
Recurrent neural networks for aortic image sequence segmentation with sparse annotations
Aug 01, 2018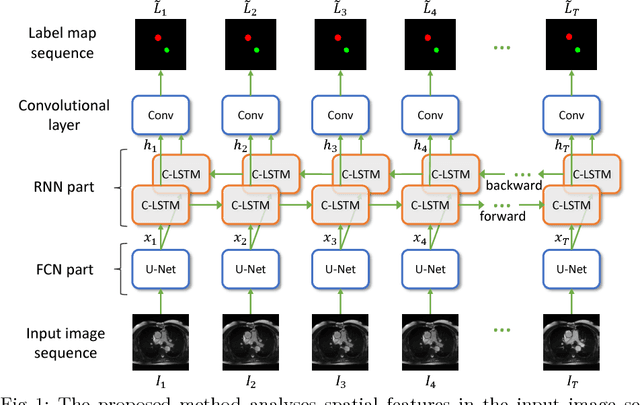


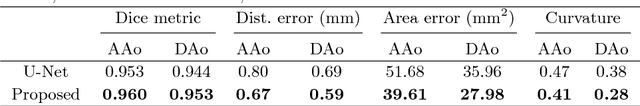
Abstract:Segmentation of image sequences is an important task in medical image analysis, which enables clinicians to assess the anatomy and function of moving organs. However, direct application of a segmentation algorithm to each time frame of a sequence may ignore the temporal continuity inherent in the sequence. In this work, we propose an image sequence segmentation algorithm by combining a fully convolutional network with a recurrent neural network, which incorporates both spatial and temporal information into the segmentation task. A key challenge in training this network is that the available manual annotations are temporally sparse, which forbids end-to-end training. We address this challenge by performing non-rigid label propagation on the annotations and introducing an exponentially weighted loss function for training. Experiments on aortic MR image sequences demonstrate that the proposed method significantly improves both accuracy and temporal smoothness of segmentation, compared to a baseline method that utilises spatial information only. It achieves an average Dice metric of 0.960 for the ascending aorta and 0.953 for the descending aorta.
Automated cardiovascular magnetic resonance image analysis with fully convolutional networks
May 22, 2018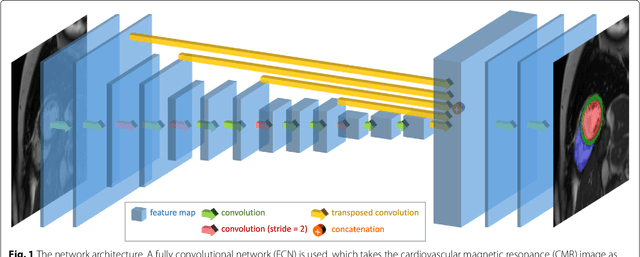

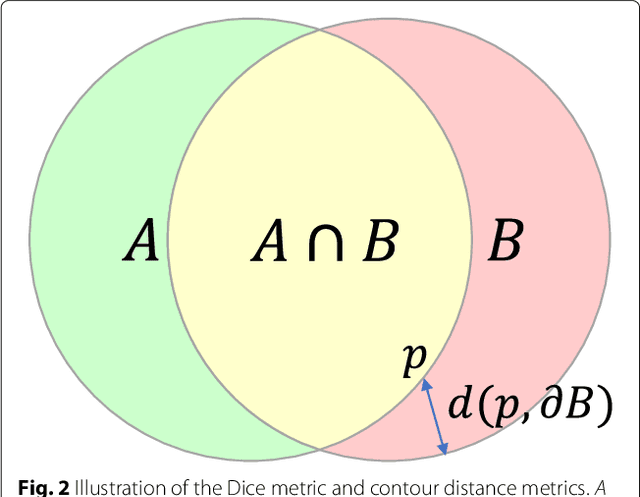
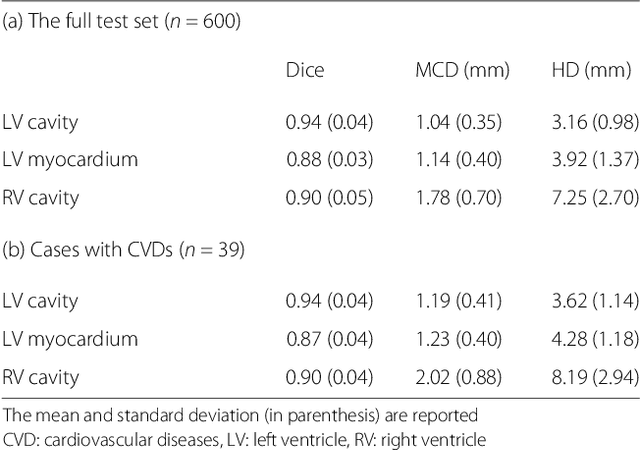
Abstract:Cardiovascular magnetic resonance (CMR) imaging is a standard imaging modality for assessing cardiovascular diseases (CVDs), the leading cause of death globally. CMR enables accurate quantification of the cardiac chamber volume, ejection fraction and myocardial mass, providing information for diagnosis and monitoring of CVDs. However, for years, clinicians have been relying on manual approaches for CMR image analysis, which is time consuming and prone to subjective errors. It is a major clinical challenge to automatically derive quantitative and clinically relevant information from CMR images. Deep neural networks have shown a great potential in image pattern recognition and segmentation for a variety of tasks. Here we demonstrate an automated analysis method for CMR images, which is based on a fully convolutional network (FCN). The network is trained and evaluated on a large-scale dataset from the UK Biobank, consisting of 4,875 subjects with 93,500 pixelwise annotated images. The performance of the method has been evaluated using a number of technical metrics, including the Dice metric, mean contour distance and Hausdorff distance, as well as clinically relevant measures, including left ventricle (LV) end-diastolic volume (LVEDV) and end-systolic volume (LVESV), LV mass (LVM); right ventricle (RV) end-diastolic volume (RVEDV) and end-systolic volume (RVESV). By combining FCN with a large-scale annotated dataset, the proposed automated method achieves a high performance on par with human experts in segmenting the LV and RV on short-axis CMR images and the left atrium (LA) and right atrium (RA) on long-axis CMR images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge