Gabriel Jimenez
ADNP-15: An Open-Source Histopathological Dataset for Neuritic Plaque Segmentation in Human Brain Whole Slide Images with Frequency Domain Image Enhancement for Stain Normalization
May 08, 2025Abstract:Alzheimer's Disease (AD) is a neurodegenerative disorder characterized by amyloid-beta plaques and tau neurofibrillary tangles, which serve as key histopathological features. The identification and segmentation of these lesions are crucial for understanding AD progression but remain challenging due to the lack of large-scale annotated datasets and the impact of staining variations on automated image analysis. Deep learning has emerged as a powerful tool for pathology image segmentation; however, model performance is significantly influenced by variations in staining characteristics, necessitating effective stain normalization and enhancement techniques. In this study, we address these challenges by introducing an open-source dataset (ADNP-15) of neuritic plaques (i.e., amyloid deposits combined with a crown of dystrophic tau-positive neurites) in human brain whole slide images. We establish a comprehensive benchmark by evaluating five widely adopted deep learning models across four stain normalization techniques, providing deeper insights into their influence on neuritic plaque segmentation. Additionally, we propose a novel image enhancement method that improves segmentation accuracy, particularly in complex tissue structures, by enhancing structural details and mitigating staining inconsistencies. Our experimental results demonstrate that this enhancement strategy significantly boosts model generalization and segmentation accuracy. All datasets and code are open-source, ensuring transparency and reproducibility while enabling further advancements in the field.
Performance Estimation for Supervised Medical Image Segmentation Models on Unlabeled Data Using UniverSeg
Apr 22, 2025Abstract:The performance of medical image segmentation models is usually evaluated using metrics like the Dice score and Hausdorff distance, which compare predicted masks to ground truth annotations. However, when applying the model to unseen data, such as in clinical settings, it is often impractical to annotate all the data, making the model's performance uncertain. To address this challenge, we propose the Segmentation Performance Evaluator (SPE), a framework for estimating segmentation models' performance on unlabeled data. This framework is adaptable to various evaluation metrics and model architectures. Experiments on six publicly available datasets across six evaluation metrics including pixel-based metrics such as Dice score and distance-based metrics like HD95, demonstrated the versatility and effectiveness of our approach, achieving a high correlation (0.956$\pm$0.046) and low MAE (0.025$\pm$0.019) compare with real Dice score on the independent test set. These results highlight its ability to reliably estimate model performance without requiring annotations. The SPE framework integrates seamlessly into any model training process without adding training overhead, enabling performance estimation and facilitating the real-world application of medical image segmentation algorithms. The source code is publicly available
Graph Theory and GNNs to Unravel the Topographical Organization of Brain Lesions in Variants of Alzheimer's Disease Progression
Mar 01, 2024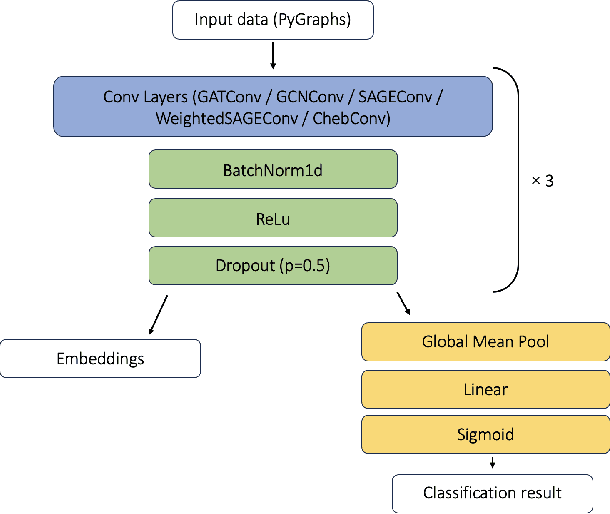
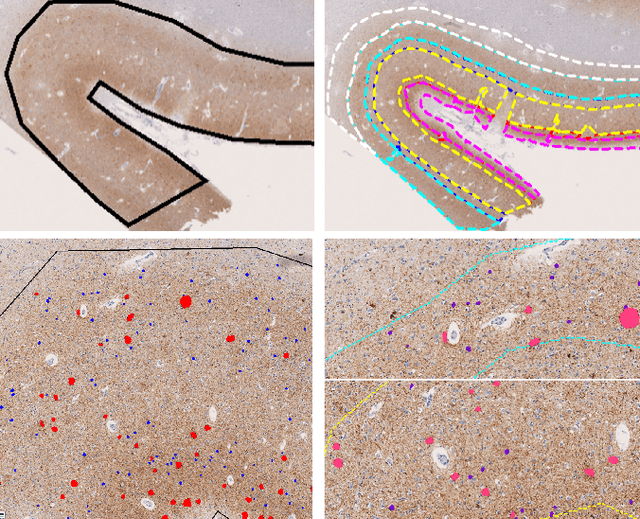
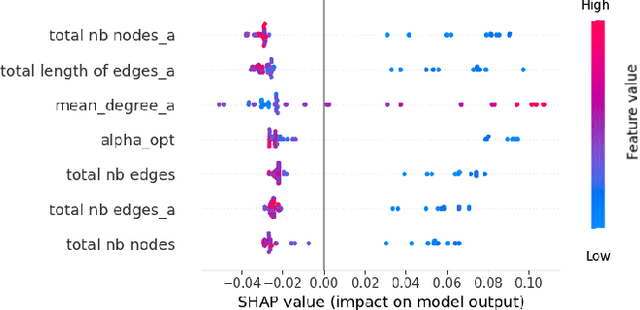
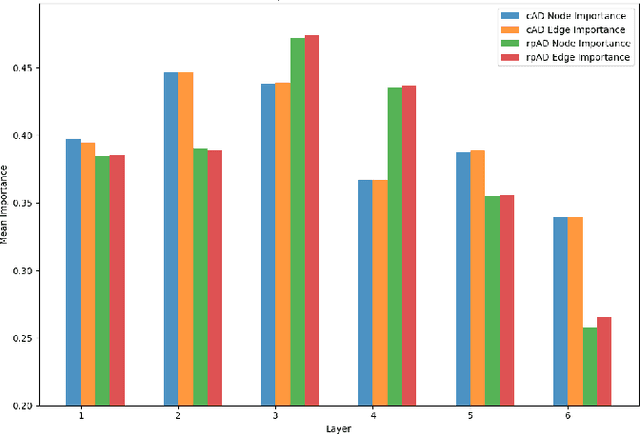
Abstract:This study utilizes graph theory and deep learning to assess variations in Alzheimer's disease (AD) neuropathologies, focusing on classic (cAD) and rapid (rpAD) progression forms. It analyses the distribution of amyloid plaques and tau tangles in postmortem brain tissues. Histopathological images are converted into tau-pathology-based graphs, and derived metrics are used for statistical analysis and in machine learning classifiers. These classifiers incorporate SHAP value explainability to differentiate between cAD and rpAD. Graph neural networks (GNNs) demonstrate greater efficiency than traditional CNN methods in analyzing this data, preserving spatial pathology context. Additionally, GNNs provide significant insights through explainable AI techniques. The analysis shows denser networks in rpAD and a distinctive impact on brain cortical layers: rpAD predominantly affects middle layers, whereas cAD influences both superficial and deep layers of the same cortical regions. These results suggest a unique neuropathological network organization for each AD variant.
Comparative Analysis of ImageNet Pre-Trained Deep Learning Models and DINOv2 in Medical Imaging Classification
Feb 13, 2024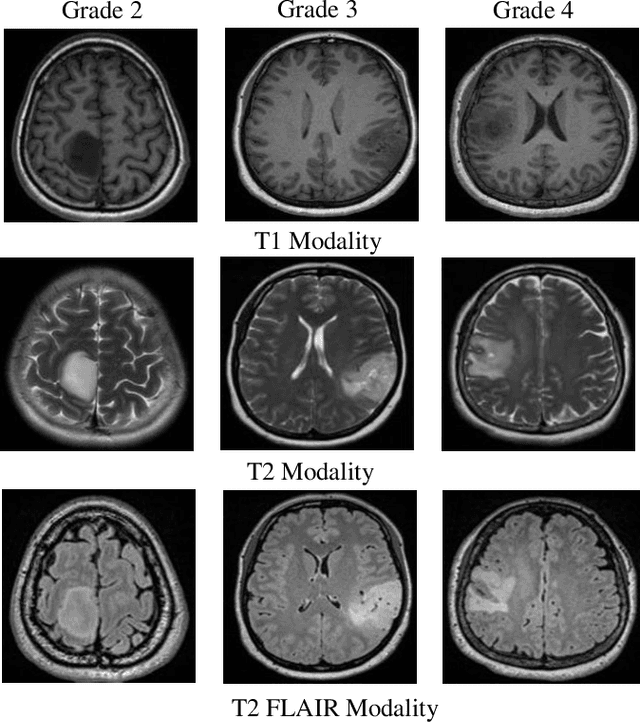
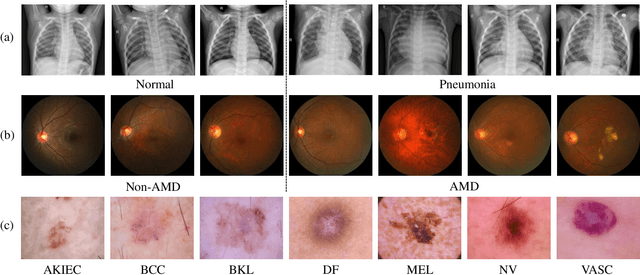
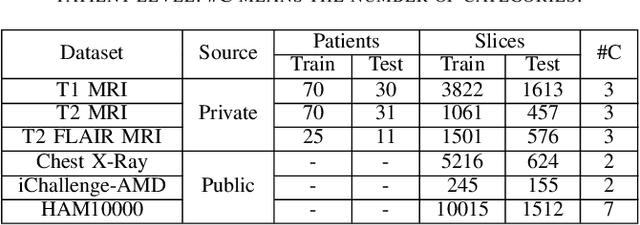
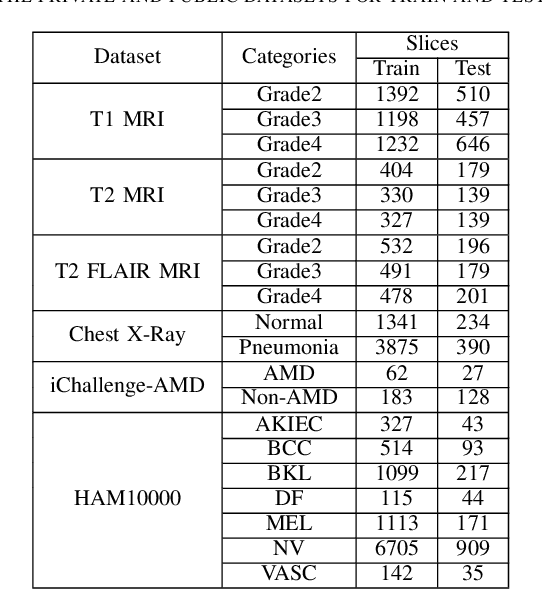
Abstract:Medical image analysis frequently encounters data scarcity challenges. Transfer learning has been effective in addressing this issue while conserving computational resources. The recent advent of foundational models like the DINOv2, which uses the vision transformer architecture, has opened new opportunities in the field and gathered significant interest. However, DINOv2's performance on clinical data still needs to be verified. In this paper, we performed a glioma grading task using three clinical modalities of brain MRI data. We compared the performance of various pre-trained deep learning models, including those based on ImageNet and DINOv2, in a transfer learning context. Our focus was on understanding the impact of the freezing mechanism on performance. We also validated our findings on three other types of public datasets: chest radiography, fundus radiography, and dermoscopy. Our findings indicate that in our clinical dataset, DINOv2's performance was not as strong as ImageNet-based pre-trained models, whereas in public datasets, DINOv2 generally outperformed other models, especially when using the frozen mechanism. Similar performance was observed with various sizes of DINOv2 models across different tasks. In summary, DINOv2 is viable for medical image classification tasks, particularly with data resembling natural images. However, its effectiveness may vary with data that significantly differs from natural images such as MRI. In addition, employing smaller versions of the model can be adequate for medical task, offering resource-saving benefits. Our codes are available at https://github.com/GuanghuiFU/medical_DINOv2_eval.
Frequency Disentangled Learning for Segmentation of Midbrain Structures from Quantitative Susceptibility Mapping Data
Feb 25, 2023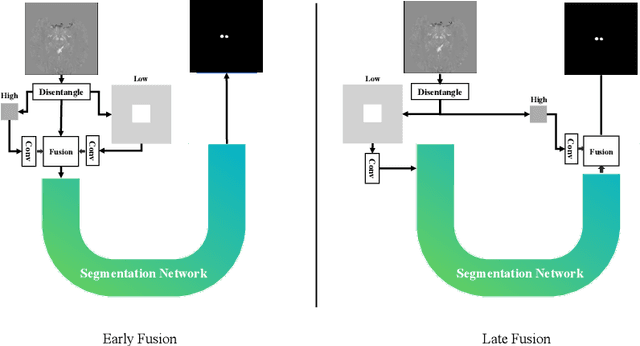

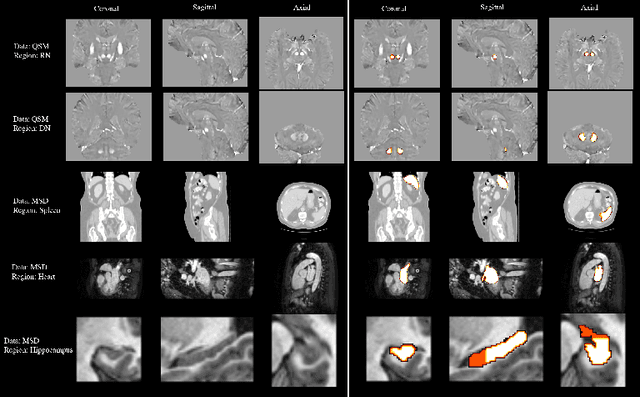
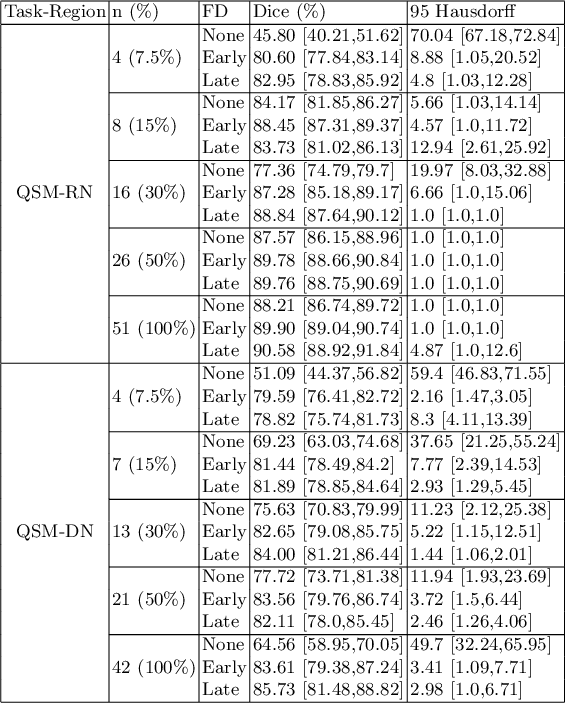
Abstract:One often lacks sufficient annotated samples for training deep segmentation models. This is in particular the case for less common imaging modalities such as Quantitative Susceptibility Mapping (QSM). It has been shown that deep models tend to fit the target function from low to high frequencies. One may hypothesize that such property can be leveraged for better training of deep learning models. In this paper, we exploit this property to propose a new training method based on frequency-domain disentanglement. It consists of two main steps: i) disentangling the image into high- and low-frequency parts and feature learning; ii) frequency-domain fusion to complete the task. The approach can be used with any backbone segmentation network. We apply the approach to the segmentation of the red and dentate nuclei from QSM data which is particularly relevant for the study of parkinsonian syndromes. We demonstrate that the proposed method provides considerable performance improvements for these tasks. We further applied it to three public datasets from the Medical Segmentation Decathlon (MSD) challenge. For two MSD tasks, it provided smaller but still substantial improvements (up to 7 points of Dice), especially under small training set situations.
Computational Pathology for Brain Disorders
Jan 13, 2023Abstract:Non-invasive brain imaging techniques allow understanding the behavior and macro changes in the brain to determine the progress of a disease. However, computational pathology provides a deeper understanding of brain disorders at cellular level, able to consolidate a diagnosis and make the bridge between the medical image and the omics analysis. In traditional histopathology, histology slides are visually inspected, under the microscope, by trained pathologists. This process is time-consuming and labor-intensive; therefore, the emergence of Computational Pathology has triggered great hope to ease this tedious task and make it more robust. This chapter focuses on understanding the state-of-the-art machine learning techniques used to analyze whole slide images within the context of brain disorders. We present a selective set of remarkable machine learning algorithms providing discriminative approaches and quality results on brain disorders. These methodologies are applied to different tasks, such as monitoring mechanisms contributing to disease progression and patient survival rates, analyzing morphological phenotypes for classification and quantitative assessment of disease, improving clinical care, diagnosing tumor specimens, and intraoperative interpretation. Thanks to the recent progress in machine learning algorithms for high-content image processing, computational pathology marks the rise of a new generation of medical discoveries and clinical protocols, including in brain disorders.
Fourier Disentangled Multimodal Prior Knowledge Fusion for Red Nucleus Segmentation in Brain MRI
Nov 02, 2022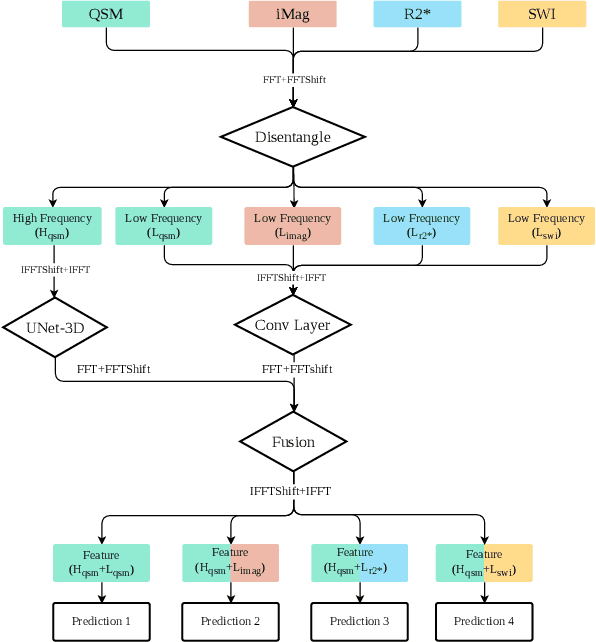
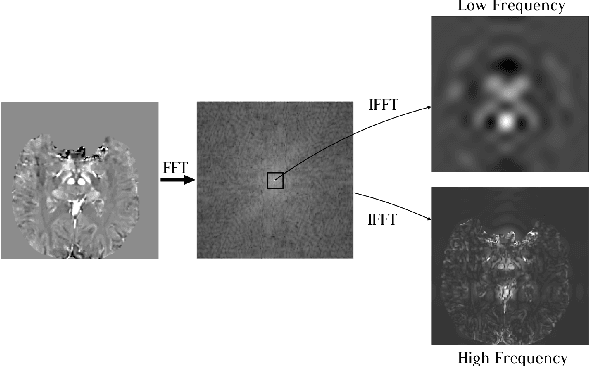
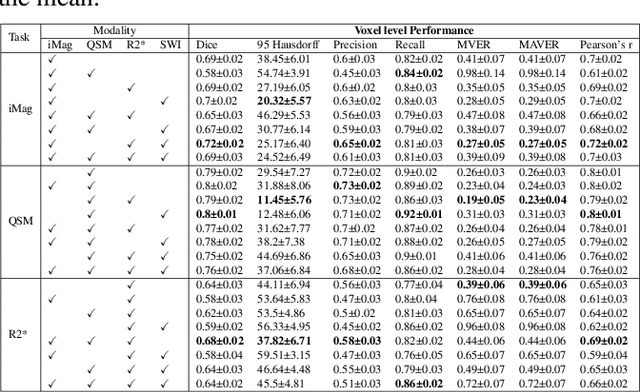

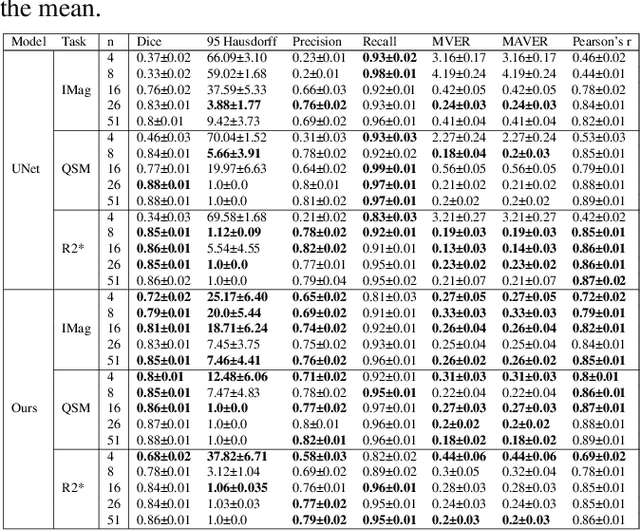
Abstract:Early and accurate diagnosis of parkinsonian syndromes is critical to provide appropriate care to patients and for inclusion in therapeutic trials. The red nucleus is a structure of the midbrain that plays an important role in these disorders. It can be visualized using iron-sensitive magnetic resonance imaging (MRI) sequences. Different iron-sensitive contrasts can be produced with MRI. Combining such multimodal data has the potential to improve segmentation of the red nucleus. Current multimodal segmentation algorithms are computationally consuming, cannot deal with missing modalities and need annotations for all modalities. In this paper, we propose a new model that integrates prior knowledge from different contrasts for red nucleus segmentation. The method consists of three main stages. First, it disentangles the image into high-level information representing the brain structure, and low-frequency information representing the contrast. The high-frequency information is then fed into a network to learn anatomical features, while the list of multimodal low-frequency information is processed by another module. Finally, feature fusion is performed to complete the segmentation task. The proposed method was used with several iron-sensitive contrasts (iMag, QSM, R2*, SWI). Experiments demonstrate that our proposed model substantially outperforms a baseline UNet model when the training set size is very small.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge