Fabio Barth
SciLaD: A Large-Scale, Transparent, Reproducible Dataset for Natural Scientific Language Processing
Dec 12, 2025Abstract:SciLaD is a novel, large-scale dataset of scientific language constructed entirely using open-source frameworks and publicly available data sources. It comprises a curated English split containing over 10 million scientific publications and a multilingual, unfiltered TEI XML split including more than 35 million publications. We also publish the extensible pipeline for generating SciLaD. The dataset construction and processing workflow demonstrates how open-source tools can enable large-scale, scientific data curation while maintaining high data quality. Finally, we pre-train a RoBERTa model on our dataset and evaluate it across a comprehensive set of benchmarks, achieving performance comparable to other scientific language models of similar size, validating the quality and utility of SciLaD. We publish the dataset and evaluation pipeline to promote reproducibility, transparency, and further research in natural scientific language processing and understanding including scholarly document processing.
MastermindEval: A Simple But Scalable Reasoning Benchmark
Mar 11, 2025



Abstract:Recent advancements in large language models (LLMs) have led to remarkable performance across a wide range of language understanding and mathematical tasks. As a result, increasing attention has been given to assessing the true reasoning capabilities of LLMs, driving research into commonsense, numerical, logical, and qualitative reasoning. However, with the rapid progress of reasoning-focused models such as OpenAI's o1 and DeepSeek's R1, there has been a growing demand for reasoning benchmarks that can keep pace with ongoing model developments. In this paper, we introduce MastermindEval, a simple, scalable, and interpretable deductive reasoning benchmark inspired by the board game Mastermind. Our benchmark supports two evaluation paradigms: (1) agentic evaluation, in which the model autonomously plays the game, and (2) deductive reasoning evaluation, in which the model is given a pre-played game state with only one possible valid code to infer. In our experimental results we (1) find that even easy Mastermind instances are difficult for current models and (2) demonstrate that the benchmark is scalable to possibly more advanced models in the future Furthermore, we investigate possible reasons why models cannot deduce the final solution and find that current models are limited in deducing the concealed code as the number of statement to combine information from is increasing.
Are Multilingual Language Models an Off-ramp for Under-resourced Languages? Will we arrive at Digital Language Equality in Europe in 2030?
Feb 18, 2025
Abstract:Large language models (LLMs) demonstrate unprecedented capabilities and define the state of the art for almost all natural language processing (NLP) tasks and also for essentially all Language Technology (LT) applications. LLMs can only be trained for languages for which a sufficient amount of pre-training data is available, effectively excluding many languages that are typically characterised as under-resourced. However, there is both circumstantial and empirical evidence that multilingual LLMs, which have been trained using data sets that cover multiple languages (including under-resourced ones), do exhibit strong capabilities for some of these under-resourced languages. Eventually, this approach may have the potential to be a technological off-ramp for those under-resourced languages for which "native" LLMs, and LLM-based technologies, cannot be developed due to a lack of training data. This paper, which concentrates on European languages, examines this idea, analyses the current situation in terms of technology support and summarises related work. The article concludes by focusing on the key open questions that need to be answered for the approach to be put into practice in a systematic way.
Multilingual European Language Models: Benchmarking Approaches and Challenges
Feb 18, 2025Abstract:The breakthrough of generative large language models (LLMs) that can solve different tasks through chat interaction has led to a significant increase in the use of general benchmarks to assess the quality or performance of these models beyond individual applications. There is also a need for better methods to evaluate and also to compare models due to the ever increasing number of new models published. However, most of the established benchmarks revolve around the English language. This paper analyses the benefits and limitations of current evaluation datasets, focusing on multilingual European benchmarks. We analyse seven multilingual benchmarks and identify four major challenges. Furthermore, we discuss potential solutions to enhance translation quality and mitigate cultural biases, including human-in-the-loop verification and iterative translation ranking. Our analysis highlights the need for culturally aware and rigorously validated benchmarks to assess the reasoning and question-answering capabilities of multilingual LLMs accurately.
Familiarity: Better Evaluation of Zero-Shot Named Entity Recognition by Quantifying Label Shifts in Synthetic Training Data
Dec 13, 2024



Abstract:Zero-shot named entity recognition (NER) is the task of detecting named entities of specific types (such as 'Person' or 'Medicine') without any training examples. Current research increasingly relies on large synthetic datasets, automatically generated to cover tens of thousands of distinct entity types, to train zero-shot NER models. However, in this paper, we find that these synthetic datasets often contain entity types that are semantically highly similar to (or even the same as) those in standard evaluation benchmarks. Because of this overlap, we argue that reported F1 scores for zero-shot NER overestimate the true capabilities of these approaches. Further, we argue that current evaluation setups provide an incomplete picture of zero-shot abilities since they do not quantify the label shift (i.e., the similarity of labels) between training and evaluation datasets. To address these issues, we propose Familiarity, a novel metric that captures both the semantic similarity between entity types in training and evaluation, as well as their frequency in the training data, to provide an estimate of label shift. It allows researchers to contextualize reported zero-shot NER scores when using custom synthetic training datasets. Further, it enables researchers to generate evaluation setups of various transfer difficulties for fine-grained analysis of zero-shot NER.
Towards Multilingual LLM Evaluation for European Languages
Oct 17, 2024



Abstract:The rise of Large Language Models (LLMs) has revolutionized natural language processing across numerous languages and tasks. However, evaluating LLM performance in a consistent and meaningful way across multiple European languages remains challenging, especially due to the scarcity of language-parallel multilingual benchmarks. We introduce a multilingual evaluation approach tailored for European languages. We employ translated versions of five widely-used benchmarks to assess the capabilities of 40 LLMs across 21 European languages. Our contributions include examining the effectiveness of translated benchmarks, assessing the impact of different translation services, and offering a multilingual evaluation framework for LLMs that includes newly created datasets: EU20-MMLU, EU20-HellaSwag, EU20-ARC, EU20-TruthfulQA, and EU20-GSM8K. The benchmarks and results are made publicly available to encourage further research in multilingual LLM evaluation.
Towards Cross-Lingual LLM Evaluation for European Languages
Oct 11, 2024



Abstract:The rise of Large Language Models (LLMs) has revolutionized natural language processing across numerous languages and tasks. However, evaluating LLM performance in a consistent and meaningful way across multiple European languages remains challenging, especially due to the scarcity of multilingual benchmarks. We introduce a cross-lingual evaluation approach tailored for European languages. We employ translated versions of five widely-used benchmarks to assess the capabilities of 40 LLMs across 21 European languages. Our contributions include examining the effectiveness of translated benchmarks, assessing the impact of different translation services, and offering a multilingual evaluation framework for LLMs that includes newly created datasets: EU20-MMLU, EU20-HellaSwag, EU20-ARC, EU20-TruthfulQA, and EU20-GSM8K. The benchmarks and results are made publicly available to encourage further research in multilingual LLM evaluation.
BLOOM: A 176B-Parameter Open-Access Multilingual Language Model
Nov 09, 2022Abstract:Large language models (LLMs) have been shown to be able to perform new tasks based on a few demonstrations or natural language instructions. While these capabilities have led to widespread adoption, most LLMs are developed by resource-rich organizations and are frequently kept from the public. As a step towards democratizing this powerful technology, we present BLOOM, a 176B-parameter open-access language model designed and built thanks to a collaboration of hundreds of researchers. BLOOM is a decoder-only Transformer language model that was trained on the ROOTS corpus, a dataset comprising hundreds of sources in 46 natural and 13 programming languages (59 in total). We find that BLOOM achieves competitive performance on a wide variety of benchmarks, with stronger results after undergoing multitask prompted finetuning. To facilitate future research and applications using LLMs, we publicly release our models and code under the Responsible AI License.
BigBIO: A Framework for Data-Centric Biomedical Natural Language Processing
Jun 30, 2022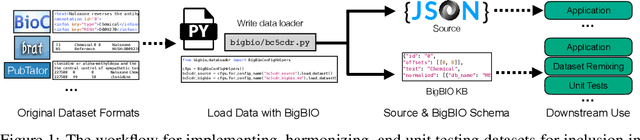
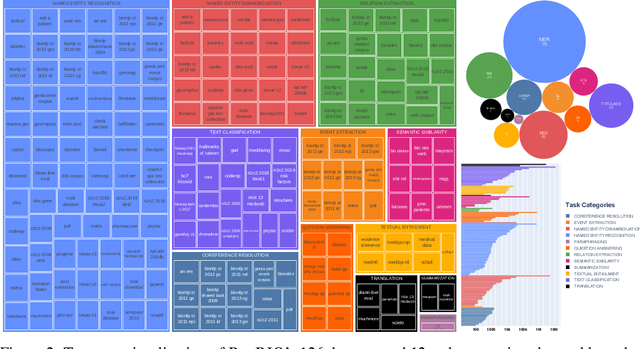
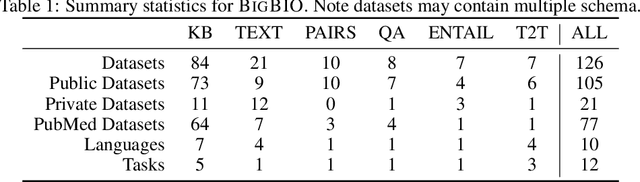

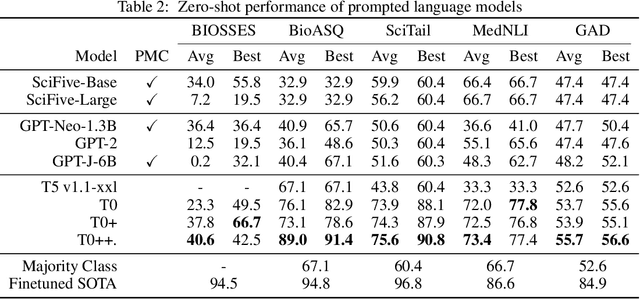
Abstract:Training and evaluating language models increasingly requires the construction of meta-datasets --diverse collections of curated data with clear provenance. Natural language prompting has recently lead to improved zero-shot generalization by transforming existing, supervised datasets into a diversity of novel pretraining tasks, highlighting the benefits of meta-dataset curation. While successful in general-domain text, translating these data-centric approaches to biomedical language modeling remains challenging, as labeled biomedical datasets are significantly underrepresented in popular data hubs. To address this challenge, we introduce BigBIO a community library of 126+ biomedical NLP datasets, currently covering 12 task categories and 10+ languages. BigBIO facilitates reproducible meta-dataset curation via programmatic access to datasets and their metadata, and is compatible with current platforms for prompt engineering and end-to-end few/zero shot language model evaluation. We discuss our process for task schema harmonization, data auditing, contribution guidelines, and outline two illustrative use cases: zero-shot evaluation of biomedical prompts and large-scale, multi-task learning. BigBIO is an ongoing community effort and is available at https://github.com/bigscience-workshop/biomedical
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge