Anthony N. Price
Meta-learning Slice-to-Volume Reconstruction in Fetal Brain MRI using Implicit Neural Representations
May 14, 2025



Abstract:High-resolution slice-to-volume reconstruction (SVR) from multiple motion-corrupted low-resolution 2D slices constitutes a critical step in image-based diagnostics of moving subjects, such as fetal brain Magnetic Resonance Imaging (MRI). Existing solutions struggle with image artifacts and severe subject motion or require slice pre-alignment to achieve satisfying reconstruction performance. We propose a novel SVR method to enable fast and accurate MRI reconstruction even in cases of severe image and motion corruption. Our approach performs motion correction, outlier handling, and super-resolution reconstruction with all operations being entirely based on implicit neural representations. The model can be initialized with task-specific priors through fully self-supervised meta-learning on either simulated or real-world data. In extensive experiments including over 480 reconstructions of simulated and clinical MRI brain data from different centers, we prove the utility of our method in cases of severe subject motion and image artifacts. Our results demonstrate improvements in reconstruction quality, especially in the presence of severe motion, compared to state-of-the-art methods, and up to 50% reduction in reconstruction time.
CINA: Conditional Implicit Neural Atlas for Spatio-Temporal Representation of Fetal Brains
Mar 13, 2024



Abstract:We introduce a conditional implicit neural atlas (CINA) for spatio-temporal atlas generation from Magnetic Resonance Images (MRI) of the neurotypical and pathological fetal brain, that is fully independent of affine or non-rigid registration. During training, CINA learns a general representation of the fetal brain and encodes subject specific information into latent code. After training, CINA can construct a faithful atlas with tissue probability maps of the fetal brain for any gestational age (GA) and anatomical variation covered within the training domain. Thus, CINA is competent to represent both, neurotypical and pathological brains. Furthermore, a trained CINA model can be fit to brain MRI of unseen subjects via test-time optimization of the latent code. CINA can then produce probabilistic tissue maps tailored to a particular subject. We evaluate our method on a total of 198 T2 weighted MRI of normal and abnormal fetal brains from the dHCP and FeTA datasets. We demonstrate CINA's capability to represent a fetal brain atlas that can be flexibly conditioned on GA and on anatomical variations like ventricular volume or degree of cortical folding, making it a suitable tool for modeling both neurotypical and pathological brains. We quantify the fidelity of our atlas by means of tissue segmentation and age prediction and compare it to an established baseline. CINA demonstrates superior accuracy for neurotypical brains and pathological brains with ventriculomegaly. Moreover, CINA scores a mean absolute error of 0.23 weeks in fetal brain age prediction, further confirming an accurate representation of fetal brain development.
Complementary Time-Frequency Domain Networks for Dynamic Parallel MR Image Reconstruction
Dec 22, 2020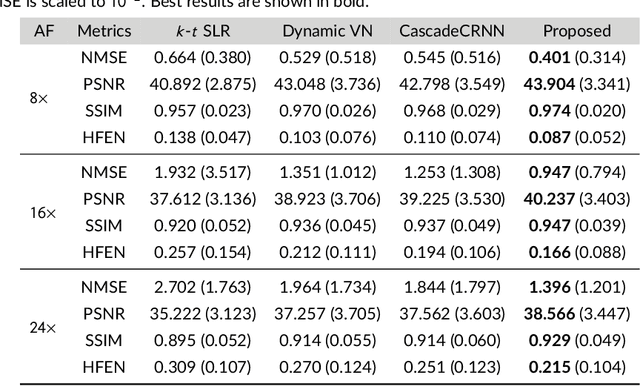
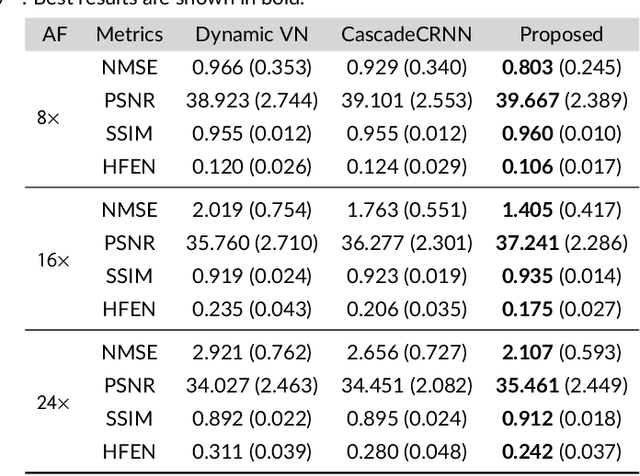
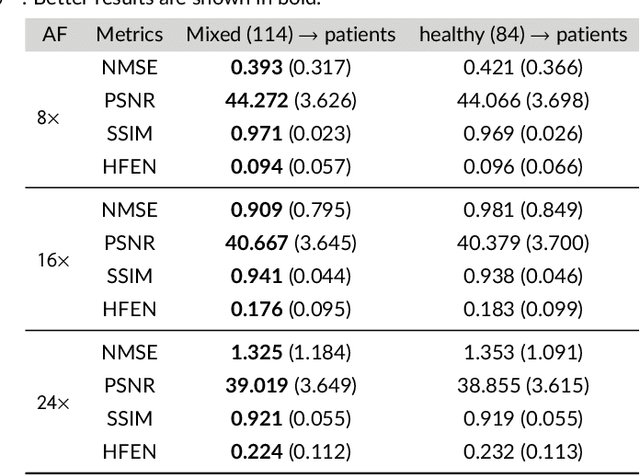
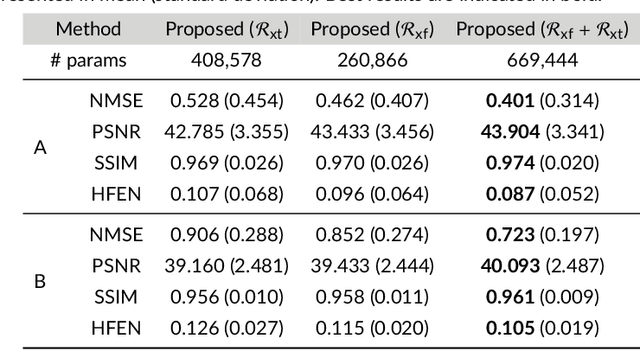
Abstract:Purpose: To introduce a novel deep learning based approach for fast and high-quality dynamic multi-coil MR reconstruction by learning a complementary time-frequency domain network that exploits spatio-temporal correlations simultaneously from complementary domains. Theory and Methods: Dynamic parallel MR image reconstruction is formulated as a multi-variable minimisation problem, where the data is regularised in combined temporal Fourier and spatial (x-f) domain as well as in spatio-temporal image (x-t) domain. An iterative algorithm based on variable splitting technique is derived, which alternates among signal de-aliasing steps in x-f and x-t spaces, a closed-form point-wise data consistency step and a weighted coupling step. The iterative model is embedded into a deep recurrent neural network which learns to recover the image via exploiting spatio-temporal redundancies in complementary domains. Results: Experiments were performed on two datasets of highly undersampled multi-coil short-axis cardiac cine MRI scans. Results demonstrate that our proposed method outperforms the current state-of-the-art approaches both quantitatively and qualitatively. The proposed model can also generalise well to data acquired from a different scanner and data with pathologies that were not seen in the training set. Conclusion: The work shows the benefit of reconstructing dynamic parallel MRI in complementary time-frequency domains with deep neural networks. The method can effectively and robustly reconstruct high-quality images from highly undersampled dynamic multi-coil data ($16 \times$ and $24 \times$ yielding 15s and 10s scan times respectively) with fast reconstruction speed (2.8s). This could potentially facilitate achieving fast single-breath-hold clinical 2D cardiac cine imaging.
Generalising Deep Learning MRI Reconstruction across Different Domains
Jan 31, 2019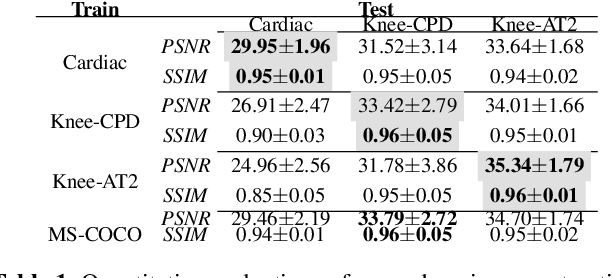
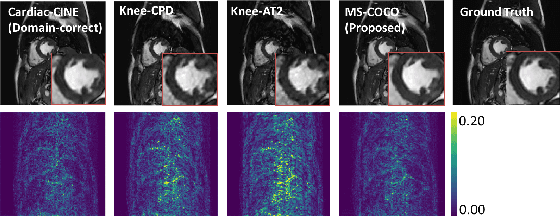

Abstract:We look into robustness of deep learning based MRI reconstruction when tested on unseen contrasts and organs. We then propose to generalise the network by training with large publicly-available natural image datasets with synthesised phase information to achieve high cross-domain reconstruction performance which is competitive with domain-specific training. To explain its generalisation mechanism, we have also analysed patch sets for different training datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge