Lucilio Cordero-Grande
Meta-learning Slice-to-Volume Reconstruction in Fetal Brain MRI using Implicit Neural Representations
May 14, 2025



Abstract:High-resolution slice-to-volume reconstruction (SVR) from multiple motion-corrupted low-resolution 2D slices constitutes a critical step in image-based diagnostics of moving subjects, such as fetal brain Magnetic Resonance Imaging (MRI). Existing solutions struggle with image artifacts and severe subject motion or require slice pre-alignment to achieve satisfying reconstruction performance. We propose a novel SVR method to enable fast and accurate MRI reconstruction even in cases of severe image and motion corruption. Our approach performs motion correction, outlier handling, and super-resolution reconstruction with all operations being entirely based on implicit neural representations. The model can be initialized with task-specific priors through fully self-supervised meta-learning on either simulated or real-world data. In extensive experiments including over 480 reconstructions of simulated and clinical MRI brain data from different centers, we prove the utility of our method in cases of severe subject motion and image artifacts. Our results demonstrate improvements in reconstruction quality, especially in the presence of severe motion, compared to state-of-the-art methods, and up to 50% reduction in reconstruction time.
CINA: Conditional Implicit Neural Atlas for Spatio-Temporal Representation of Fetal Brains
Mar 13, 2024



Abstract:We introduce a conditional implicit neural atlas (CINA) for spatio-temporal atlas generation from Magnetic Resonance Images (MRI) of the neurotypical and pathological fetal brain, that is fully independent of affine or non-rigid registration. During training, CINA learns a general representation of the fetal brain and encodes subject specific information into latent code. After training, CINA can construct a faithful atlas with tissue probability maps of the fetal brain for any gestational age (GA) and anatomical variation covered within the training domain. Thus, CINA is competent to represent both, neurotypical and pathological brains. Furthermore, a trained CINA model can be fit to brain MRI of unseen subjects via test-time optimization of the latent code. CINA can then produce probabilistic tissue maps tailored to a particular subject. We evaluate our method on a total of 198 T2 weighted MRI of normal and abnormal fetal brains from the dHCP and FeTA datasets. We demonstrate CINA's capability to represent a fetal brain atlas that can be flexibly conditioned on GA and on anatomical variations like ventricular volume or degree of cortical folding, making it a suitable tool for modeling both neurotypical and pathological brains. We quantify the fidelity of our atlas by means of tissue segmentation and age prediction and compare it to an established baseline. CINA demonstrates superior accuracy for neurotypical brains and pathological brains with ventriculomegaly. Moreover, CINA scores a mean absolute error of 0.23 weeks in fetal brain age prediction, further confirming an accurate representation of fetal brain development.
Fetal MRI by robust deep generative prior reconstruction and diffeomorphic registration: application to gestational age prediction
Oct 29, 2021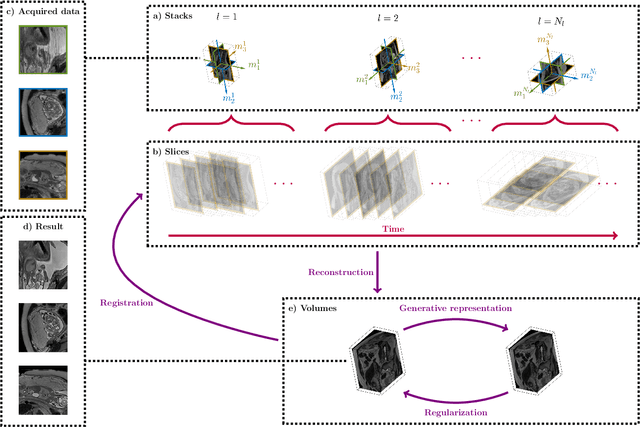
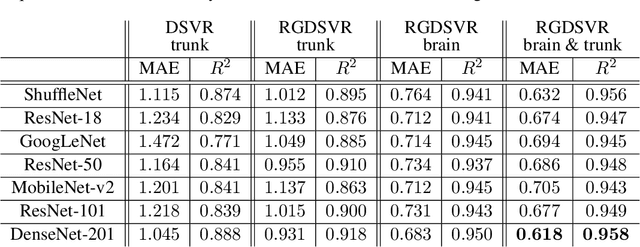

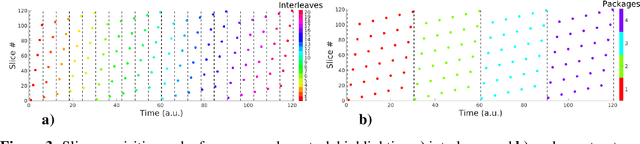
Abstract:Magnetic resonance imaging of whole fetal body and placenta is limited by different sources of motion affecting the womb. Usual scanning techniques employ single-shot multi-slice sequences where anatomical information in different slices may be subject to different deformations, contrast variations or artifacts. Volumetric reconstruction formulations have been proposed to correct for these factors, but they must accommodate a non-homogeneous and non-isotropic sampling, so regularization becomes necessary. Thus, in this paper we propose a deep generative prior for robust volumetric reconstructions integrated with a diffeomorphic volume to slice registration method. Experiments are performed to validate our contributions and compare with a state of the art method in a cohort of $72$ fetal datasets in the range of $20-36$ weeks gestational age. Results suggest improved image resolution and more accurate prediction of gestational age at scan when comparing to a state of the art reconstruction method. In addition, gestational age prediction results from our volumetric reconstructions compare favourably with existing brain-based approaches, with boosted accuracy when integrating information of organs other than the brain. Namely, a mean absolute error of $0.618$ weeks ($R^2=0.958$) is achieved when combining fetal brain and trunk information.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge