Zheyu Zhu
Omni-Seg+: A Scale-aware Dynamic Network for Pathological Image Segmentation
Jun 27, 2022
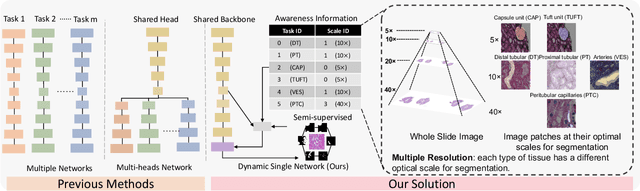
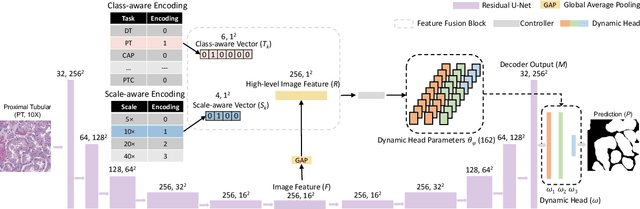
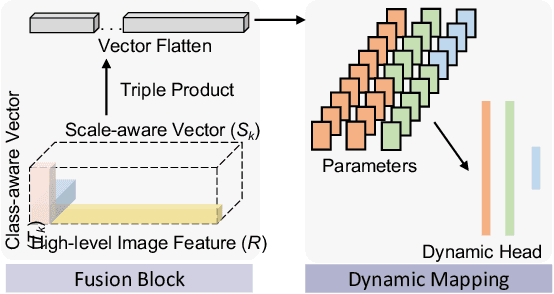
Abstract:Comprehensive semantic segmentation on renal pathological images is challenging due to the heterogeneous scales of the objects. For example, on a whole slide image (WSI), the cross-sectional areas of glomeruli can be 64 times larger than that of the peritubular capillaries, making it impractical to segment both objects on the same patch, at the same scale. To handle this scaling issue, prior studies have typically trained multiple segmentation networks in order to match the optimal pixel resolution of heterogeneous tissue types. This multi-network solution is resource-intensive and fails to model the spatial relationship between tissue types. In this paper, we propose the Omni-Seg+ network, a scale-aware dynamic neural network that achieves multi-object (six tissue types) and multi-scale (5X to 40X scale) pathological image segmentation via a single neural network. The contribution of this paper is three-fold: (1) a novel scale-aware controller is proposed to generalize the dynamic neural network from single-scale to multi-scale; (2) semi-supervised consistency regularization of pseudo-labels is introduced to model the inter-scale correlation of unannotated tissue types into a single end-to-end learning paradigm; and (3) superior scale-aware generalization is evidenced by directly applying a model trained on human kidney images to mouse kidney images, without retraining. By learning from ~150,000 human pathological image patches from six tissue types at three different resolutions, our approach achieved superior segmentation performance according to human visual assessment and evaluation of image-omics (i.e., spatial transcriptomics). The official implementation is available at https://github.com/ddrrnn123/Omni-Seg.
Glo-In-One: Holistic Glomerular Detection, Segmentation, and Lesion Characterization with Large-scale Web Image Mining
May 31, 2022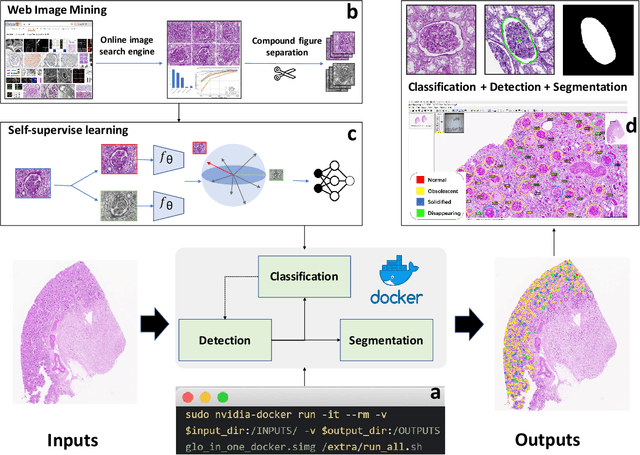
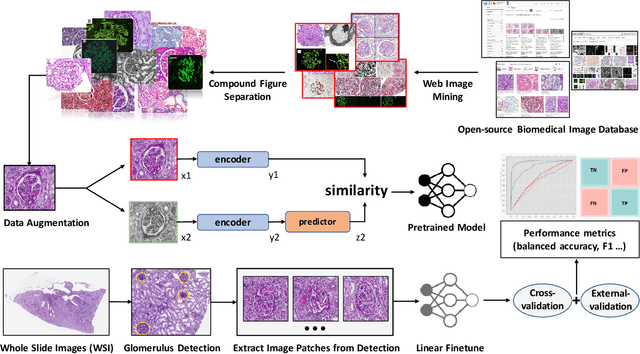
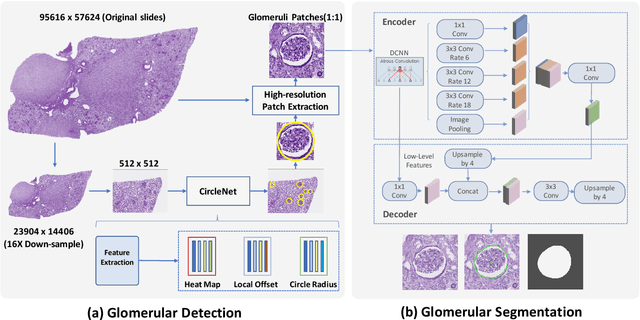
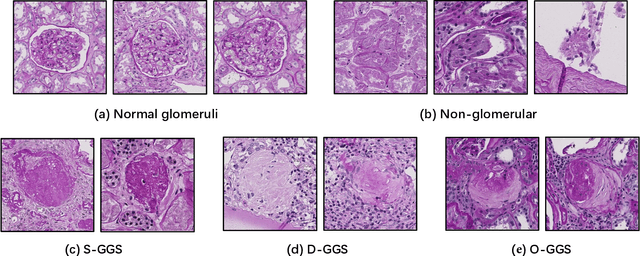
Abstract:The quantitative detection, segmentation, and characterization of glomeruli from high-resolution whole slide imaging (WSI) play essential roles in the computer-assisted diagnosis and scientific research in digital renal pathology. Historically, such comprehensive quantification requires extensive programming skills in order to be able to handle heterogeneous and customized computational tools. To bridge the gap of performing glomerular quantification for non-technical users, we develop the Glo-In-One toolkit to achieve holistic glomerular detection, segmentation, and characterization via a single line of command. Additionally, we release a large-scale collection of 30,000 unlabeled glomerular images to further facilitate the algorithmic development of self-supervised deep learning. The inputs of the Glo-In-One toolkit are WSIs, while the outputs are (1) WSI-level multi-class circle glomerular detection results (which can be directly manipulated with ImageScope), (2) glomerular image patches with segmentation masks, and (3) different lesion types. To leverage the performance of the Glo-In-One toolkit, we introduce self-supervised deep learning to glomerular quantification via large-scale web image mining. The GGS fine-grained classification model achieved a decent performance compared with baseline supervised methods while only using 10% of the annotated data. The glomerular detection achieved an average precision of 0.627 with circle representations, while the glomerular segmentation achieved a 0.955 patch-wise Dice Similarity Coefficient (DSC).
Holistic Fine-grained GGS Characterization: From Detection to Unbalanced Classification
Jan 31, 2022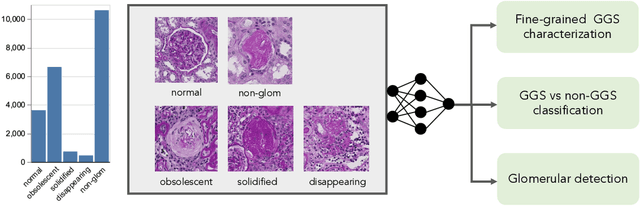



Abstract:Recent studies have demonstrated the diagnostic and prognostic values of global glomerulosclerosis (GGS) in IgA nephropathy, aging, and end-stage renal disease. However, the fine-grained quantitative analysis of multiple GGS subtypes (e.g., obsolescent, solidified, and disappearing glomerulosclerosis) is typically a resource extensive manual process. Very few automatic methods, if any, have been developed to bridge this gap for such analytics. In this paper, we present a holistic pipeline to quantify GGS (with both detection and classification) from a whole slide image in a fully automatic manner. In addition, we conduct the fine-grained classification for the sub-types of GGS. Our study releases the open-source quantitative analytical tool for fine-grained GGS characterization while tackling the technical challenges in unbalanced classification and integrating detection and classification.
Circle Representation for Medical Object Detection
Oct 22, 2021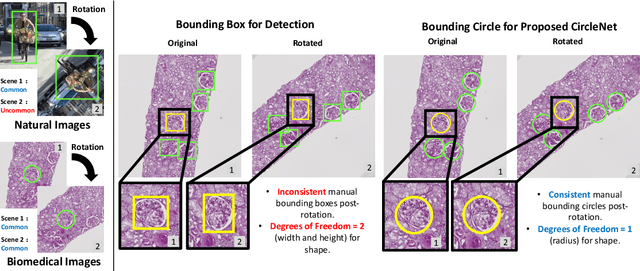
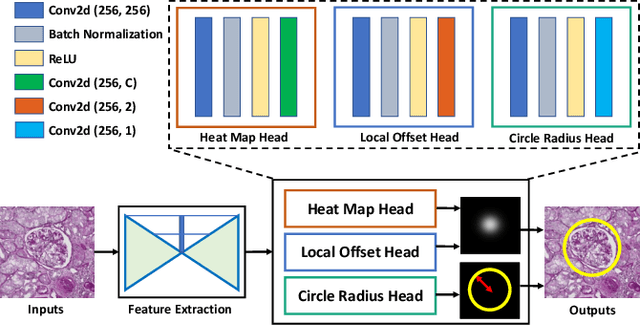
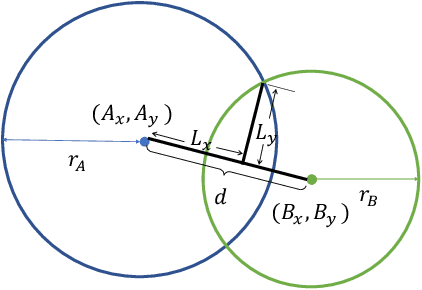
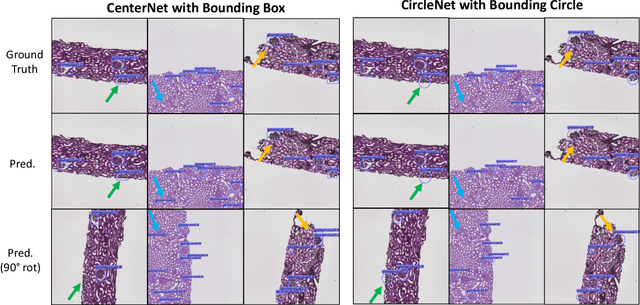
Abstract:Box representation has been extensively used for object detection in computer vision. Such representation is efficacious but not necessarily optimized for biomedical objects (e.g., glomeruli), which play an essential role in renal pathology. In this paper, we propose a simple circle representation for medical object detection and introduce CircleNet, an anchor-free detection framework. Compared with the conventional bounding box representation, the proposed bounding circle representation innovates in three-fold: (1) it is optimized for ball-shaped biomedical objects; (2) The circle representation reduced the degree of freedom compared with box representation; (3) It is naturally more rotation invariant. When detecting glomeruli and nuclei on pathological images, the proposed circle representation achieved superior detection performance and be more rotation-invariant, compared with the bounding box. The code has been made publicly available: https://github.com/hrlblab/CircleNet
Improve Global Glomerulosclerosis Classification with Imbalanced Data using CircleMix Augmentation
Jan 16, 2021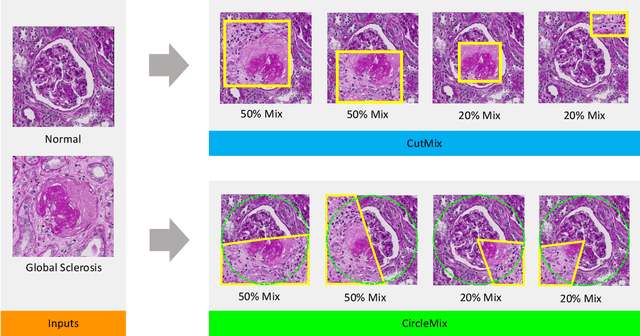
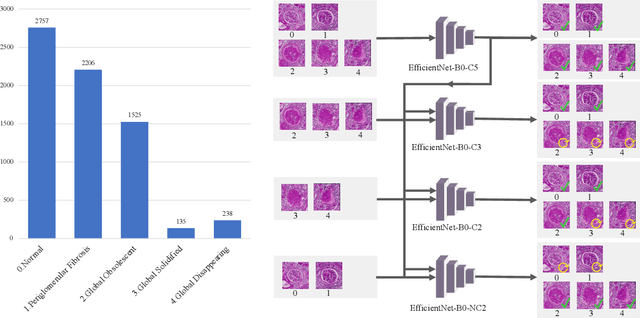
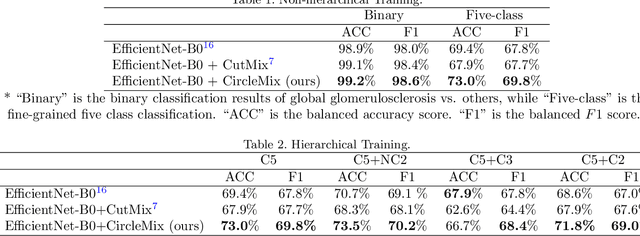
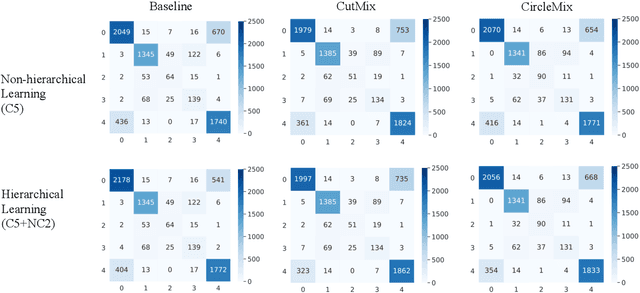
Abstract:The classification of glomerular lesions is a routine and essential task in renal pathology. Recently, machine learning approaches, especially deep learning algorithms, have been used to perform computer-aided lesion characterization of glomeruli. However, one major challenge of developing such methods is the naturally imbalanced distribution of different lesions. In this paper, we propose CircleMix, a novel data augmentation technique, to improve the accuracy of classifying globally sclerotic glomeruli with a hierarchical learning strategy. Different from the recently proposed CutMix method, the CircleMix augmentation is optimized for the ball-shaped biomedical objects, such as glomeruli. 6,861 glomeruli with five classes (normal, periglomerular fibrosis, obsolescent glomerulosclerosis, solidified glomerulosclerosis, and disappearing glomerulosclerosis) were employed to develop and evaluate the proposed methods. From five-fold cross-validation, the proposed CircleMix augmentation achieved superior performance (Balanced Accuracy=73.0%) compared with the EfficientNet-B0 baseline (Balanced Accuracy=69.4%)
EasierPath: An Open-source Tool for Human-in-the-loop Deep Learning of Renal Pathology
Jul 28, 2020
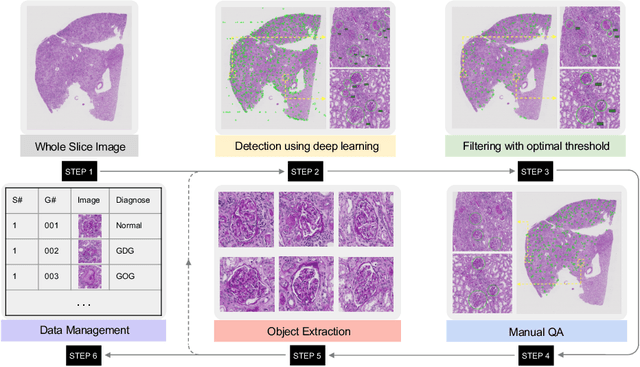


Abstract:Considerable morphological phenotyping studies in nephrology have emerged in the past few years, aiming to discover hidden regularities between clinical and imaging phenotypes. Such studies have been largely enabled by deep learning based image analysis to extract sparsely located targeting objects (e.g., glomeruli) on high-resolution whole slide images (WSI). However, such methods need to be trained using labor-intensive high-quality annotations, ideally labeled by pathologists. Inspired by the recent "human-in-the-loop" strategy, we developed EasierPath, an open-source tool to integrate human physicians and deep learning algorithms for efficient large-scale pathological image quantification as a loop. Using EasierPath, physicians are able to (1) optimize the recall and precision of deep learning object detection outcomes adaptively, (2) seamlessly support deep learning outcomes refining using either our EasierPath or prevalent ImageScope software without changing physician's user habit, and (3) manage and phenotype each object with user-defined classes. As a user case of EasierPath, we present the procedure of curating large-scale glomeruli in an efficient human-in-the-loop fashion (with two loops). From the experiments, the EasierPath saved 57 % of the annotation efforts to curate 8,833 glomeruli during the second loop. Meanwhile, the average precision of glomerular detection was leveraged from 0.504 to 0.620. The EasierPath software has been released as open-source to enable the large-scale glomerular prototyping. The code can be found in https://github.com/yuankaihuo/EasierPath
CircleNet: Anchor-free Detection with Circle Representation
Jun 03, 2020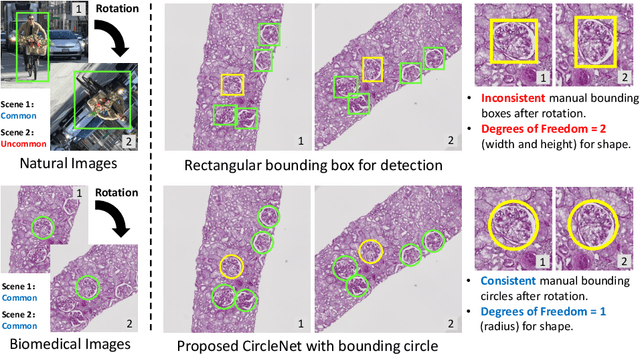

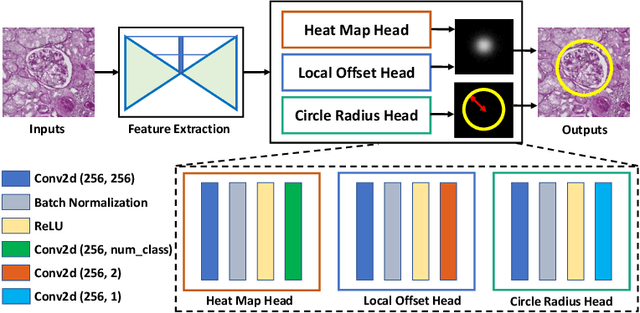
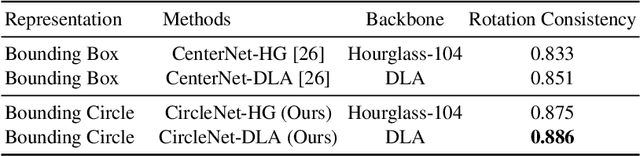
Abstract:Object detection networks are powerful in computer vision, but not necessarily optimized for biomedical object detection. In this work, we propose CircleNet, a simple anchor-free detection method with circle representation for detection of the ball-shaped glomerulus. Different from the traditional bounding box based detection method, the bounding circle (1) reduces the degrees of freedom of detection representation, (2) is naturally rotation invariant, (3) and optimized for ball-shaped objects. The key innovation to enable this representation is the anchor-free framework with the circle detection head. We evaluate CircleNet in the context of detection of glomerulus. CircleNet increases average precision of the glomerulus detection from 0.598 to 0.647. Another key advantage is that CircleNet achieves better rotation consistency compared with bounding box representations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge