Yui Lo
DeepMultiConnectome: Deep Multi-Task Prediction of Structural Connectomes Directly from Diffusion MRI Tractography
May 27, 2025



Abstract:Diffusion MRI (dMRI) tractography enables in vivo mapping of brain structural connections, but traditional connectome generation is time-consuming and requires gray matter parcellation, posing challenges for large-scale studies. We introduce DeepMultiConnectome, a deep-learning model that predicts structural connectomes directly from tractography, bypassing the need for gray matter parcellation while supporting multiple parcellation schemes. Using a point-cloud-based neural network with multi-task learning, the model classifies streamlines according to their connected regions across two parcellation schemes, sharing a learned representation. We train and validate DeepMultiConnectome on tractography from the Human Connectome Project Young Adult dataset ($n = 1000$), labeled with an 84 and 164 region gray matter parcellation scheme. DeepMultiConnectome predicts multiple structural connectomes from a whole-brain tractogram containing 3 million streamlines in approximately 40 seconds. DeepMultiConnectome is evaluated by comparing predicted connectomes with traditional connectomes generated using the conventional method of labeling streamlines using a gray matter parcellation. The predicted connectomes are highly correlated with traditionally generated connectomes ($r = 0.992$ for an 84-region scheme; $r = 0.986$ for a 164-region scheme) and largely preserve network properties. A test-retest analysis of DeepMultiConnectome demonstrates reproducibility comparable to traditionally generated connectomes. The predicted connectomes perform similarly to traditionally generated connectomes in predicting age and cognitive function. Overall, DeepMultiConnectome provides a scalable, fast model for generating subject-specific connectomes across multiple parcellation schemes.
A Multimodal Deep Learning Approach for White Matter Shape Prediction in Diffusion MRI Tractography
Apr 25, 2025Abstract:Shape measures have emerged as promising descriptors of white matter tractography, offering complementary insights into anatomical variability and associations with cognitive and clinical phenotypes. However, conventional methods for computing shape measures are computationally expensive and time-consuming for large-scale datasets due to reliance on voxel-based representations. We propose Tract2Shape, a novel multimodal deep learning framework that leverages geometric (point cloud) and scalar (tabular) features to predict ten white matter tractography shape measures. To enhance model efficiency, we utilize a dimensionality reduction algorithm for the model to predict five primary shape components. The model is trained and evaluated on two independently acquired datasets, the HCP-YA dataset, and the PPMI dataset. We evaluate the performance of Tract2Shape by training and testing it on the HCP-YA dataset and comparing the results with state-of-the-art models. To further assess its robustness and generalization ability, we also test Tract2Shape on the unseen PPMI dataset. Tract2Shape outperforms SOTA deep learning models across all ten shape measures, achieving the highest average Pearson's r and the lowest nMSE on the HCP-YA dataset. The ablation study shows that both multimodal input and PCA contribute to performance gains. On the unseen testing PPMI dataset, Tract2Shape maintains a high Pearson's r and low nMSE, demonstrating strong generalizability in cross-dataset evaluation. Tract2Shape enables fast, accurate, and generalizable prediction of white matter shape measures from tractography data, supporting scalable analysis across datasets. This framework lays a promising foundation for future large-scale white matter shape analysis.
TractCloud-FOV: Deep Learning-based Robust Tractography Parcellation in Diffusion MRI with Incomplete Field of View
Feb 28, 2025Abstract:Tractography parcellation classifies streamlines reconstructed from diffusion MRI into anatomically defined fiber tracts for clinical and research applications. However, clinical scans often have incomplete fields of view (FOV) where brain regions are partially imaged, leading to partial or truncated fiber tracts. To address this challenge, we introduce TractCloud-FOV, a deep learning framework that robustly parcellates tractography under conditions of incomplete FOV. We propose a novel training strategy, FOV-Cut Augmentation (FOV-CA), in which we synthetically cut tractograms to simulate a spectrum of real-world inferior FOV cutoff scenarios. This data augmentation approach enriches the training set with realistic truncated streamlines, enabling the model to achieve superior generalization. We evaluate the proposed TractCloud-FOV on both synthetically cut tractography and two real-life datasets with incomplete FOV. TractCloud-FOV significantly outperforms several state-of-the-art methods on all testing datasets in terms of streamline classification accuracy, generalization ability, tract anatomical depiction, and computational efficiency. Overall, TractCloud-FOV achieves efficient and consistent tractography parcellation in diffusion MRI with incomplete FOV.
DINeuro: Distilling Knowledge from 2D Natural Images via Deformable Tubular Transferring Strategy for 3D Neuron Reconstruction
Oct 29, 2024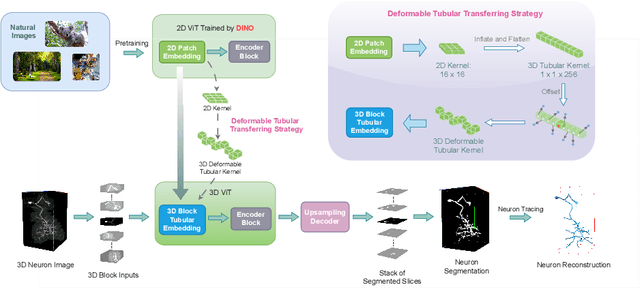
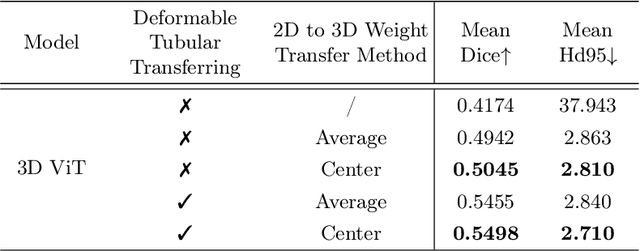
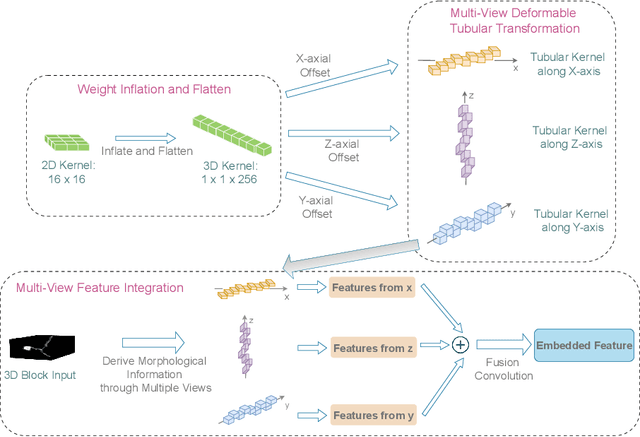
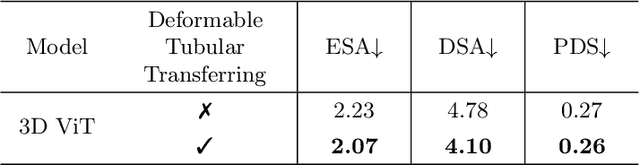
Abstract:Reconstructing neuron morphology from 3D light microscope imaging data is critical to aid neuroscientists in analyzing brain networks and neuroanatomy. With the boost from deep learning techniques, a variety of learning-based segmentation models have been developed to enhance the signal-to-noise ratio of raw neuron images as a pre-processing step in the reconstruction workflow. However, most existing models directly encode the latent representative features of volumetric neuron data but neglect their intrinsic morphological knowledge. To address this limitation, we design a novel framework that distills the prior knowledge from a 2D Vision Transformer pre-trained on extensive 2D natural images to facilitate neuronal morphological learning of our 3D Vision Transformer. To bridge the knowledge gap between the 2D natural image and 3D microscopic morphologic domains, we propose a deformable tubular transferring strategy that adapts the pre-trained 2D natural knowledge to the inherent tubular characteristics of neuronal structure in the latent embedding space. The experimental results on the Janelia dataset of the BigNeuron project demonstrate that our method achieves a segmentation performance improvement of 4.53% in mean Dice and 3.56% in mean 95% Hausdorff distance.
TractShapeNet: Efficient Multi-Shape Learning with 3D Tractography Point Clouds
Oct 29, 2024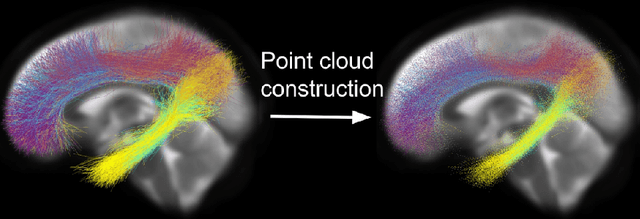
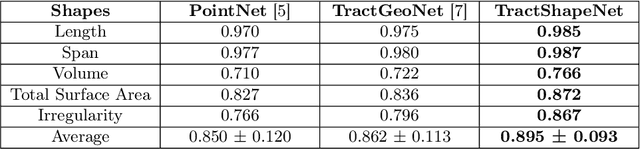

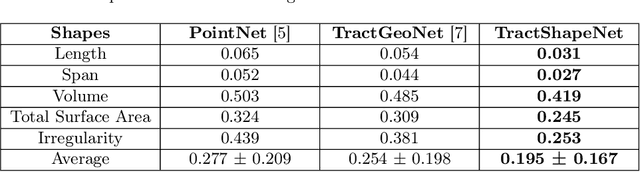
Abstract:Brain imaging studies have demonstrated that diffusion MRI tractography geometric shape descriptors can inform the study of the brain's white matter pathways and their relationship to brain function. In this work, we investigate the possibility of utilizing a deep learning model to compute shape measures of the brain's white matter connections. We introduce a novel framework, TractShapeNet, that leverages a point cloud representation of tractography to compute five shape measures: length, span, volume, total surface area, and irregularity. We assess the performance of the method on a large dataset including 1065 healthy young adults. Experiments for shape measure computation demonstrate that our proposed TractShapeNet outperforms other point cloud-based neural network models in both the Pearson correlation coefficient and normalized error metrics. We compare the inference runtime results with the conventional shape computation tool DSI-Studio. Our results demonstrate that a deep learning approach enables faster and more efficient shape measure computation. We also conduct experiments on two downstream language cognition prediction tasks, showing that shape measures from TractShapeNet perform similarly to those computed by DSI-Studio. Our code will be available at: https://github.com/SlicerDMRI/TractShapeNet.
The shape of the brain's connections is predictive of cognitive performance: an explainable machine learning study
Oct 19, 2024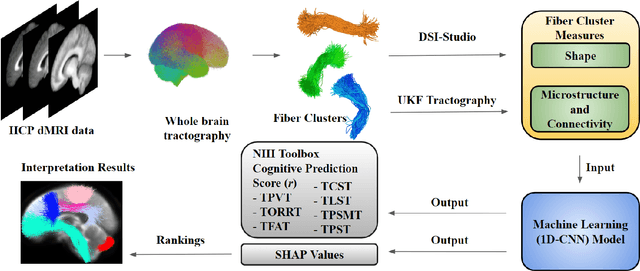
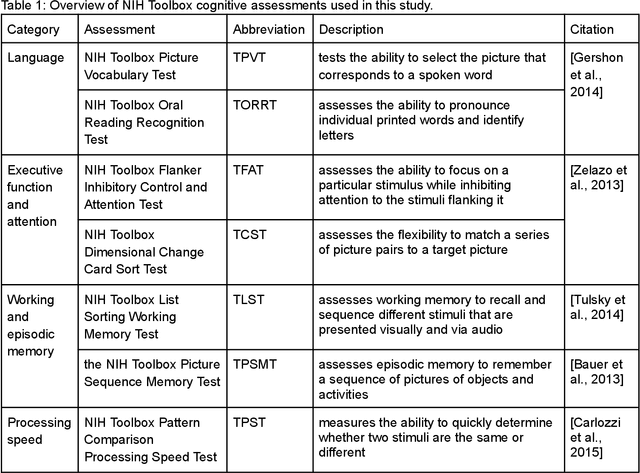

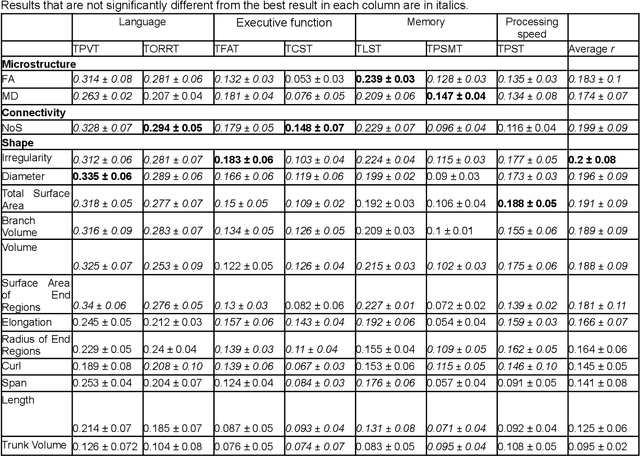
Abstract:The shape of the brain's white matter connections is relatively unexplored in diffusion MRI tractography analysis. While it is known that tract shape varies in populations and across the human lifespan, it is unknown if the variability in dMRI tractography-derived shape may relate to the brain's functional variability across individuals. This work explores the potential of leveraging tractography fiber cluster shape measures to predict subject-specific cognitive performance. We implement machine learning models to predict individual cognitive performance scores. We study a large-scale database from the HCP-YA study. We apply an atlas-based fiber cluster parcellation to the dMRI tractography of each individual. We compute 15 shape, microstructure, and connectivity features for each fiber cluster. Using these features as input, we train a total of 210 models to predict 7 different NIH Toolbox cognitive performance assessments. We apply an explainable AI technique, SHAP, to assess the importance of each fiber cluster for prediction. Our results demonstrate that shape measures are predictive of individual cognitive performance. The studied shape measures, such as irregularity, diameter, total surface area, volume, and branch volume, are as effective for prediction as microstructure and connectivity measures. The overall best-performing feature is a shape feature, irregularity, which describes how different a cluster's shape is from an idealized cylinder. Further interpretation using SHAP values suggest that fiber clusters with features highly predictive of cognitive ability are widespread throughout the brain, including fiber clusters from the superficial association, deep association, cerebellar, striatal, and projection pathways. This study demonstrates the strong potential of shape descriptors to enhance the study of the brain's white matter and its relationship to cognitive function.
White Matter Geometry-Guided Score-Based Diffusion Model for Tissue Microstructure Imputation in Tractography Imaging
Jul 28, 2024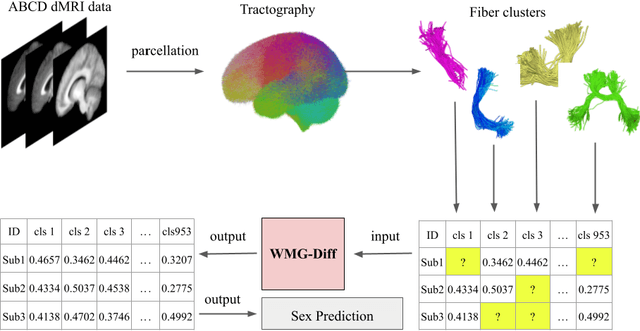
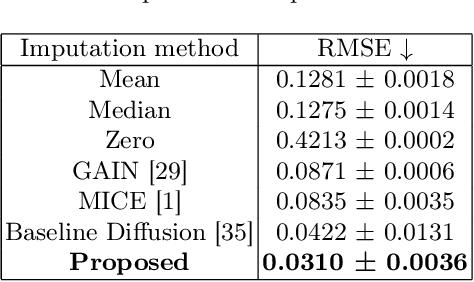
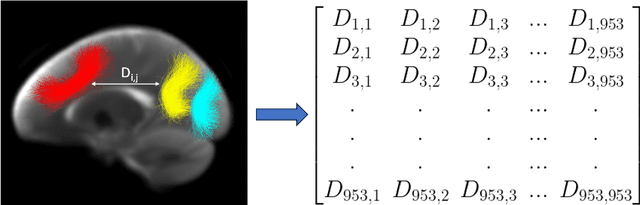
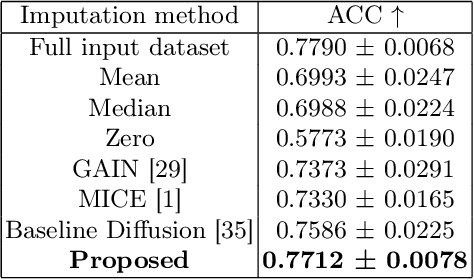
Abstract:Parcellation of white matter tractography provides anatomical features for disease prediction, anatomical tract segmentation, surgical brain mapping, and non-imaging phenotype classifications. However, parcellation does not always reach 100% accuracy due to various factors, including inter-individual anatomical variability and the quality of neuroimaging scan data. The failure to identify parcels causes a problem of missing microstructure data values, which is especially challenging for downstream tasks that analyze large brain datasets. In this work, we propose a novel deep-learning model to impute tissue microstructure: the White Matter Geometry-guided Diffusion (WMG-Diff) model. Specifically, we first propose a deep score-based guided diffusion model to impute tissue microstructure for diffusion magnetic resonance imaging (dMRI) tractography fiber clusters. Second, we propose a white matter atlas geometric relationship-guided denoising function to guide the reverse denoising process at the subject-specific level. Third, we train and evaluate our model on a large dataset with 9342 subjects. Comprehensive experiments for tissue microstructure imputation and a downstream non-imaging phenotype prediction task demonstrate that our proposed WMG-Diff outperforms state-of-the-art methods.
Cross-domain Fiber Cluster Shape Analysis for Language Performance Cognitive Score Prediction
Mar 30, 2024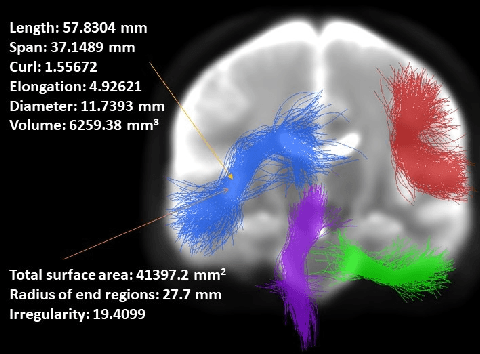
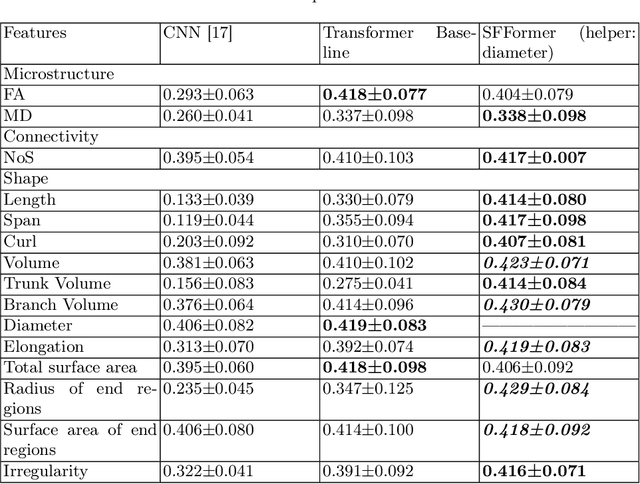
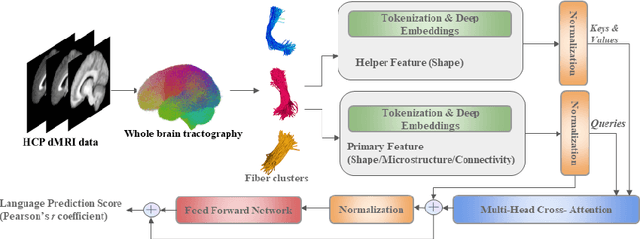
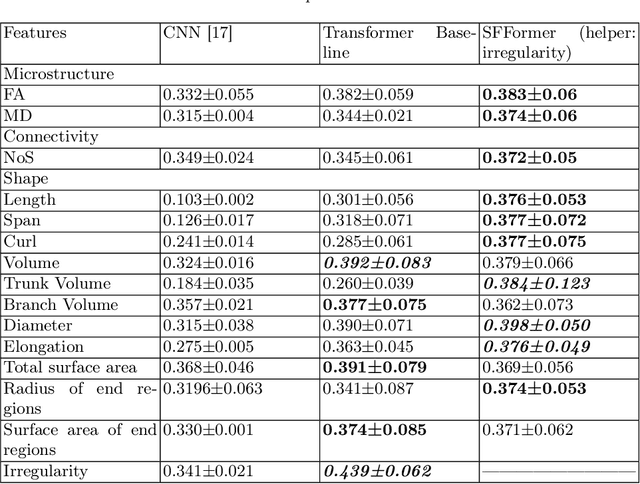
Abstract:Shape plays an important role in computer graphics, offering informative features to convey an object's morphology and functionality. Shape analysis in brain imaging can help interpret structural and functionality correlations of the human brain. In this work, we investigate the shape of the brain's 3D white matter connections and its potential predictive relationship to human cognitive function. We reconstruct brain connections as sequences of 3D points using diffusion magnetic resonance imaging (dMRI) tractography. To describe each connection, we extract 12 shape descriptors in addition to traditional dMRI connectivity and tissue microstructure features. We introduce a novel framework, Shape--fused Fiber Cluster Transformer (SFFormer), that leverages a multi-head cross-attention feature fusion module to predict subject-specific language performance based on dMRI tractography. We assess the performance of the method on a large dataset including 1065 healthy young adults. The results demonstrate that both the transformer-based SFFormer model and its inter/intra feature fusion with shape, microstructure, and connectivity are informative, and together, they improve the prediction of subject-specific language performance scores. Overall, our results indicate that the shape of the brain's connections is predictive of human language function.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge