DINeuro: Distilling Knowledge from 2D Natural Images via Deformable Tubular Transferring Strategy for 3D Neuron Reconstruction
Paper and Code
Oct 29, 2024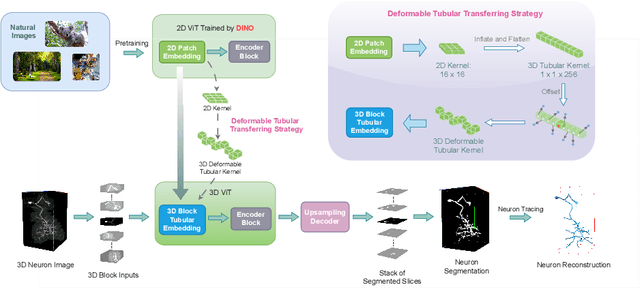
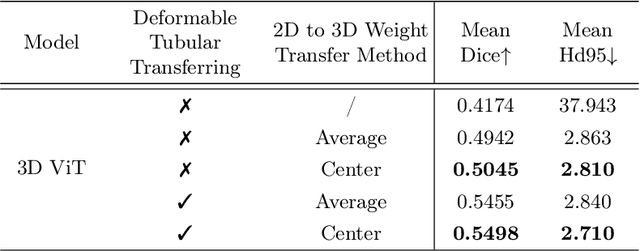
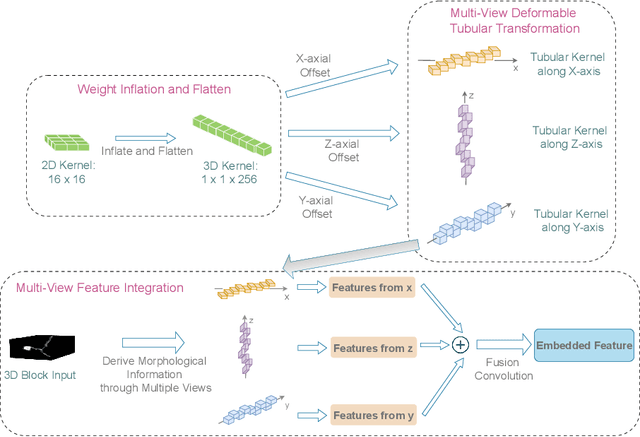
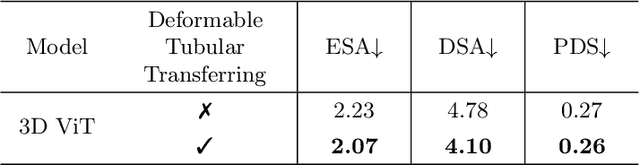
Reconstructing neuron morphology from 3D light microscope imaging data is critical to aid neuroscientists in analyzing brain networks and neuroanatomy. With the boost from deep learning techniques, a variety of learning-based segmentation models have been developed to enhance the signal-to-noise ratio of raw neuron images as a pre-processing step in the reconstruction workflow. However, most existing models directly encode the latent representative features of volumetric neuron data but neglect their intrinsic morphological knowledge. To address this limitation, we design a novel framework that distills the prior knowledge from a 2D Vision Transformer pre-trained on extensive 2D natural images to facilitate neuronal morphological learning of our 3D Vision Transformer. To bridge the knowledge gap between the 2D natural image and 3D microscopic morphologic domains, we propose a deformable tubular transferring strategy that adapts the pre-trained 2D natural knowledge to the inherent tubular characteristics of neuronal structure in the latent embedding space. The experimental results on the Janelia dataset of the BigNeuron project demonstrate that our method achieves a segmentation performance improvement of 4.53% in mean Dice and 3.56% in mean 95% Hausdorff distance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge