Valentin Comte
Retrieval Augmented Generation Evaluation for Health Documents
May 07, 2025Abstract:Safe and trustworthy use of Large Language Models (LLM) in the processing of healthcare documents and scientific papers could substantially help clinicians, scientists and policymakers in overcoming information overload and focusing on the most relevant information at a given moment. Retrieval Augmented Generation (RAG) is a promising method to leverage the potential of LLMs while enhancing the accuracy of their outcomes. This report assesses the potentials and shortcomings of such approaches in the automatic knowledge synthesis of different types of documents in the health domain. To this end, it describes: (1) an internally developed proof of concept pipeline that employs state-of-the-art practices to deliver safe and trustable analysis for healthcare documents and scientific papers called RAGEv (Retrieval Augmented Generation Evaluation); (2) a set of evaluation tools for LLM-based document retrieval and generation; (3) a benchmark dataset to verify the accuracy and veracity of the results called RAGEv-Bench. It concludes that careful implementations of RAG techniques could minimize most of the common problems in the use of LLMs for document processing in the health domain, obtaining very high scores both on short yes/no answers and long answers. There is a high potential for incorporating it into the day-to-day work of policy support tasks, but additional efforts are required to obtain a consistent and trustworthy tool.
Conformal Risk Control for Pulmonary Nodule Detection
Dec 28, 2024



Abstract:Quantitative tools are increasingly appealing for decision support in healthcare, driven by the growing capabilities of advanced AI systems. However, understanding the predictive uncertainties surrounding a tool's output is crucial for decision-makers to ensure reliable and transparent decisions. In this paper, we present a case study on pulmonary nodule detection for lung cancer screening, enhancing an advanced detection model with an uncertainty quantification technique called conformal risk control (CRC). We demonstrate that prediction sets with conformal guarantees are attractive measures of predictive uncertainty in the safety-critical healthcare domain, allowing end-users to achieve arbitrary validity by trading off false positives and providing formal statistical guarantees on model performance. Among ground-truth nodules annotated by at least three radiologists, our model achieves a sensitivity that is competitive with that generally achieved by individual radiologists, with a slight increase in false positives. Furthermore, we illustrate the risks of using off-the-shelve prediction models when faced with ontological uncertainty, such as when radiologists disagree on what constitutes the ground truth on pulmonary nodules.
Multi-Center Fetal Brain Tissue Annotation (FeTA) Challenge 2022 Results
Feb 08, 2024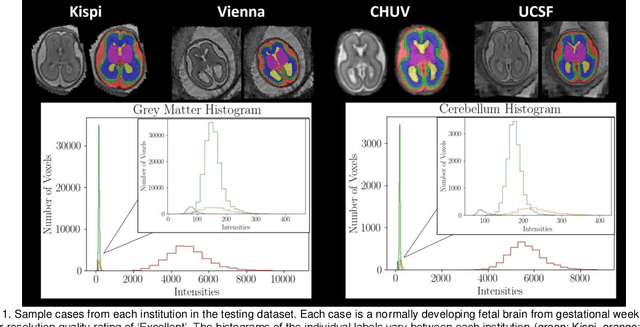
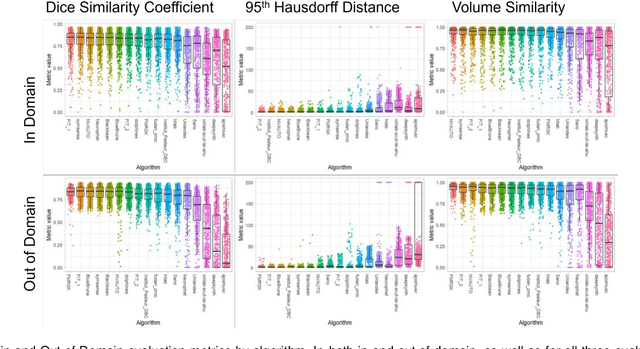
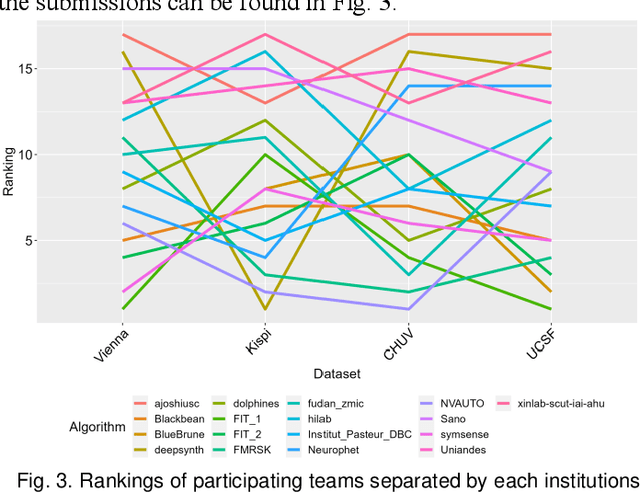
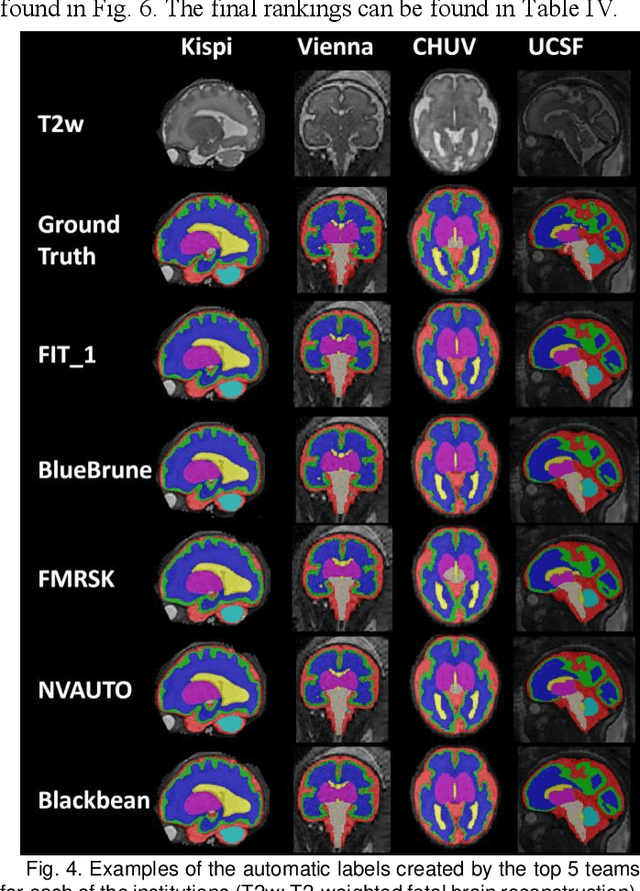
Abstract:Segmentation is a critical step in analyzing the developing human fetal brain. There have been vast improvements in automatic segmentation methods in the past several years, and the Fetal Brain Tissue Annotation (FeTA) Challenge 2021 helped to establish an excellent standard of fetal brain segmentation. However, FeTA 2021 was a single center study, and the generalizability of algorithms across different imaging centers remains unsolved, limiting real-world clinical applicability. The multi-center FeTA Challenge 2022 focuses on advancing the generalizability of fetal brain segmentation algorithms for magnetic resonance imaging (MRI). In FeTA 2022, the training dataset contained images and corresponding manually annotated multi-class labels from two imaging centers, and the testing data contained images from these two imaging centers as well as two additional unseen centers. The data from different centers varied in many aspects, including scanners used, imaging parameters, and fetal brain super-resolution algorithms applied. 16 teams participated in the challenge, and 17 algorithms were evaluated. Here, a detailed overview and analysis of the challenge results are provided, focusing on the generalizability of the submissions. Both in- and out of domain, the white matter and ventricles were segmented with the highest accuracy, while the most challenging structure remains the cerebral cortex due to anatomical complexity. The FeTA Challenge 2022 was able to successfully evaluate and advance generalizability of multi-class fetal brain tissue segmentation algorithms for MRI and it continues to benchmark new algorithms. The resulting new methods contribute to improving the analysis of brain development in utero.
Unsupervised Segmentation of Fetal Brain MRI using Deep Learning Cascaded Registration
Jul 07, 2023



Abstract:Accurate segmentation of fetal brain magnetic resonance images is crucial for analyzing fetal brain development and detecting potential neurodevelopmental abnormalities. Traditional deep learning-based automatic segmentation, although effective, requires extensive training data with ground-truth labels, typically produced by clinicians through a time-consuming annotation process. To overcome this challenge, we propose a novel unsupervised segmentation method based on multi-atlas segmentation, that accurately segments multiple tissues without relying on labeled data for training. Our method employs a cascaded deep learning network for 3D image registration, which computes small, incremental deformations to the moving image to align it precisely with the fixed image. This cascaded network can then be used to register multiple annotated images with the image to be segmented, and combine the propagated labels to form a refined segmentation. Our experiments demonstrate that the proposed cascaded architecture outperforms the state-of-the-art registration methods that were tested. Furthermore, the derived segmentation method achieves similar performance and inference time to nnU-Net while only using a small subset of annotated data for the multi-atlas segmentation task and none for training the network. Our pipeline for registration and multi-atlas segmentation is publicly available at https://github.com/ValBcn/CasReg.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge