Seyed Sadegh Mohseni Salehi
Dual-Domain Self-Supervised Learning for Accelerated Non-Cartesian MRI Reconstruction
Feb 18, 2023



Abstract:While enabling accelerated acquisition and improved reconstruction accuracy, current deep MRI reconstruction networks are typically supervised, require fully sampled data, and are limited to Cartesian sampling patterns. These factors limit their practical adoption as fully-sampled MRI is prohibitively time-consuming to acquire clinically. Further, non-Cartesian sampling patterns are particularly desirable as they are more amenable to acceleration and show improved motion robustness. To this end, we present a fully self-supervised approach for accelerated non-Cartesian MRI reconstruction which leverages self-supervision in both k-space and image domains. In training, the undersampled data are split into disjoint k-space domain partitions. For the k-space self-supervision, we train a network to reconstruct the input undersampled data from both the disjoint partitions and from itself. For the image-level self-supervision, we enforce appearance consistency obtained from the original undersampled data and the two partitions. Experimental results on our simulated multi-coil non-Cartesian MRI dataset demonstrate that DDSS can generate high-quality reconstruction that approaches the accuracy of the fully supervised reconstruction, outperforming previous baseline methods. Finally, DDSS is shown to scale to highly challenging real-world clinical MRI reconstruction acquired on a portable low-field (0.064 T) MRI scanner with no data available for supervised training while demonstrating improved image quality as compared to traditional reconstruction, as determined by a radiologist study.
ContraReg: Contrastive Learning of Multi-modality Unsupervised Deformable Image Registration
Jun 27, 2022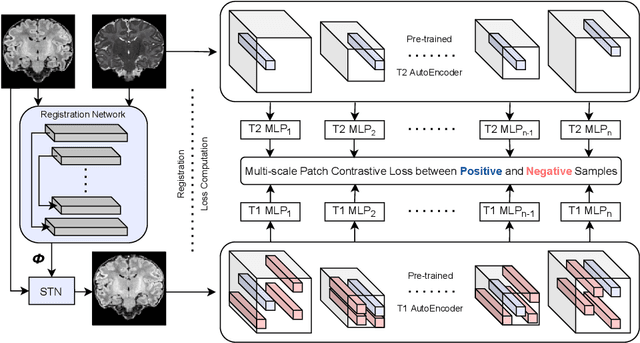
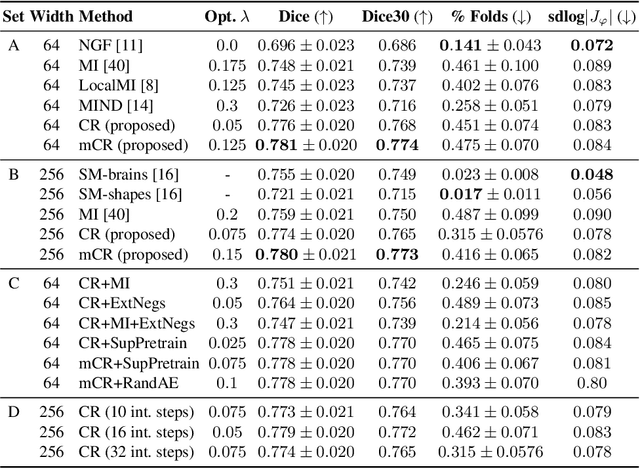
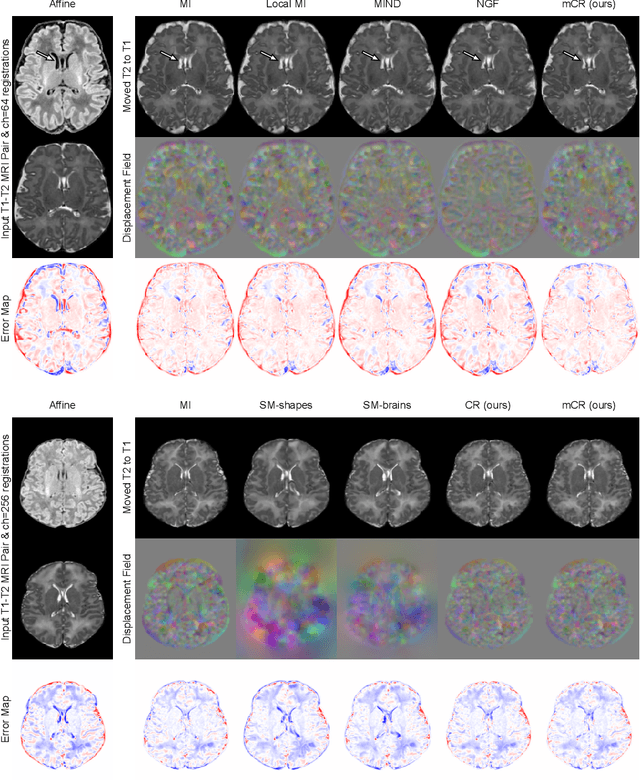
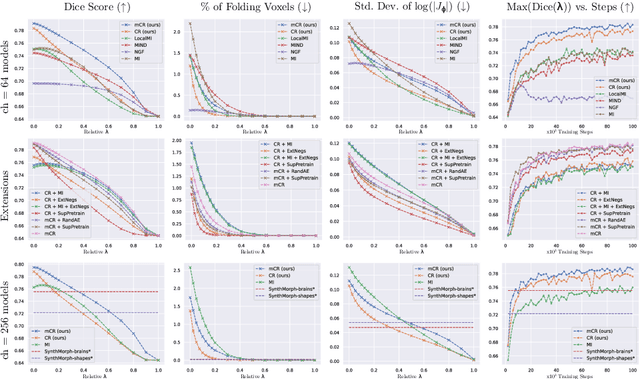
Abstract:Establishing voxelwise semantic correspondence across distinct imaging modalities is a foundational yet formidable computer vision task. Current multi-modality registration techniques maximize hand-crafted inter-domain similarity functions, are limited in modeling nonlinear intensity-relationships and deformations, and may require significant re-engineering or underperform on new tasks, datasets, and domain pairs. This work presents ContraReg, an unsupervised contrastive representation learning approach to multi-modality deformable registration. By projecting learned multi-scale local patch features onto a jointly learned inter-domain embedding space, ContraReg obtains representations useful for non-rigid multi-modality alignment. Experimentally, ContraReg achieves accurate and robust results with smooth and invertible deformations across a series of baselines and ablations on a neonatal T1-T2 brain MRI registration task with all methods validated over a wide range of deformation regularization strengths.
DSFormer: A Dual-domain Self-supervised Transformer for Accelerated Multi-contrast MRI Reconstruction
Jan 26, 2022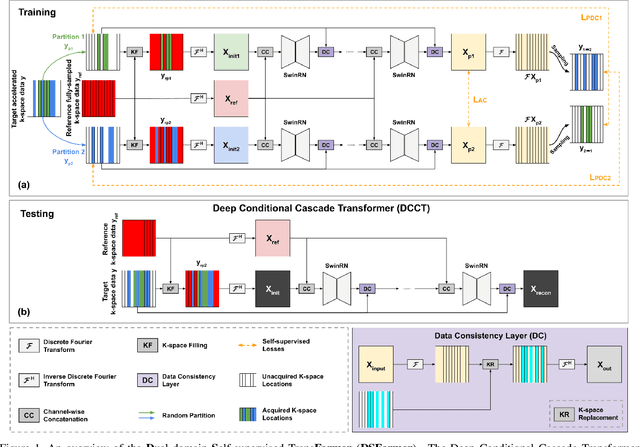
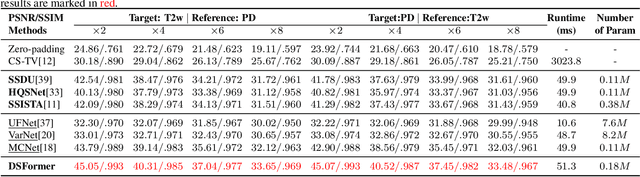
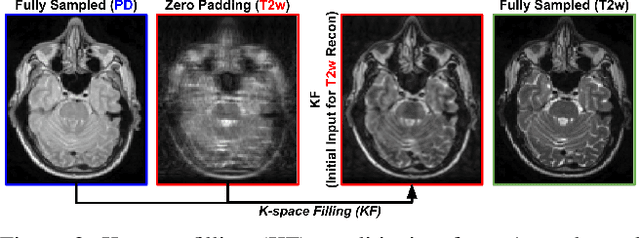
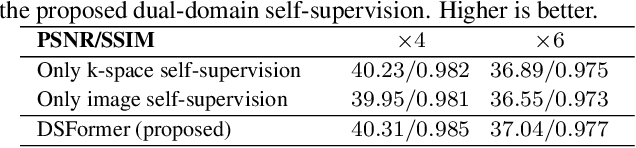
Abstract:Multi-contrast MRI (MC-MRI) captures multiple complementary imaging modalities to aid in radiological decision-making. Given the need for lowering the time cost of multiple acquisitions, current deep accelerated MRI reconstruction networks focus on exploiting the redundancy between multiple contrasts. However, existing works are largely supervised with paired data and/or prohibitively expensive fully-sampled MRI sequences. Further, reconstruction networks typically rely on convolutional architectures which are limited in their capacity to model long-range interactions and may lead to suboptimal recovery of fine anatomical detail. To these ends, we present a dual-domain self-supervised transformer (DSFormer) for accelerated MC-MRI reconstruction. DSFormer develops a deep conditional cascade transformer (DCCT) consisting of several cascaded Swin transformer reconstruction networks (SwinRN) trained under two deep conditioning strategies to enable MC-MRI information sharing. We further present a dual-domain (image and k-space) self-supervised learning strategy for DCCT to alleviate the costs of acquiring fully sampled training data. DSFormer generates high-fidelity reconstructions which experimentally outperform current fully-supervised baselines. Moreover, we find that DSFormer achieves nearly the same performance when trained either with full supervision or with our proposed dual-domain self-supervision.
Deep Predictive Motion Tracking in Magnetic Resonance Imaging: Application to Fetal Imaging
Sep 25, 2019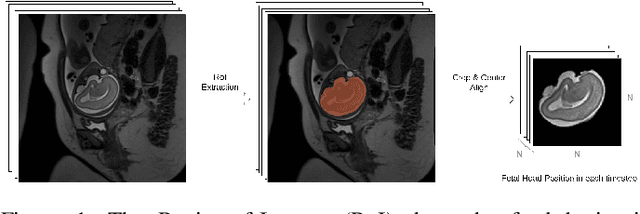
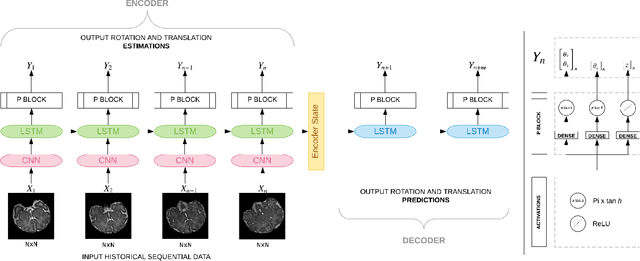
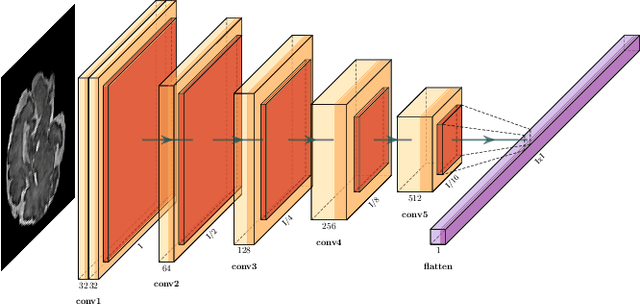
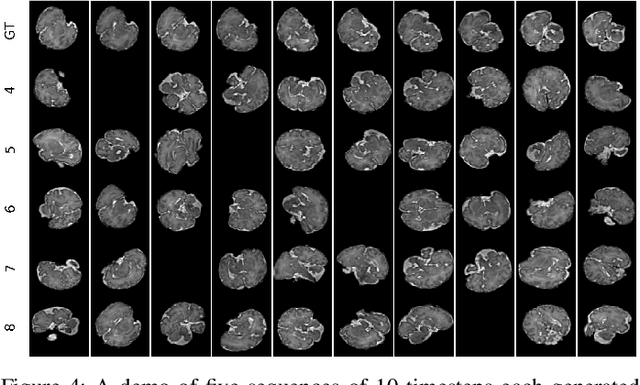
Abstract:Fetal magnetic resonance imaging (MRI) is challenged by uncontrollable, large, and irregular fetal movements. Fetal MRI is performed in a fully interactive manner in which a technologist monitors motion to prescribe slices in right angles with respect to the anatomy of interest. Current practice involves repeated acquisitions to ensure diagnostic-quality images are acquired; and the scans are retrospectively registered slice-by-slice to reconstruct 3D images. Nonetheless, manual monitoring of 3D fetal motion based on displayed 2D slices and navigation at the level of stacks-of-slices (instead of slices) is sub-optimal and inefficient. The current process is highly operator-dependent, requires extensive training, and significantly increases the length of fetal MRI scans which makes them difficult for pregnant women, and costly. With that motivation, we presented a new real-time image-based motion tracking technique in MRI using deep learning that can significantly improve state of the art. Through a combination of spatial and temporal encoder-decoder networks, our system learns to predict 3D pose of the fetal head based on dynamics of motion inferred directly from sequences of acquired slices. Compared to recent works that estimate static 3D pose of the subject from slices, our method learns to predict dynamics of 3D motion. We compared our trained network on held-out test sets (including data with different characteristics, e.g. different age ranges, and motion trajectories recorded from volunteer subjects) with networks designed for estimation as well as methods adopted to make predictions. The results of all estimation and prediction tasks show that we achieved reliable motion tracking in fetal MRI. This technique can be augmented with deep learning based fast anatomy detection, segmentation, and image registration techniques to build real-time motion tracking and navigation systems.
Real-time Deep Pose Estimation with Geodesic Loss for Image-to-Template Rigid Registration
Aug 18, 2018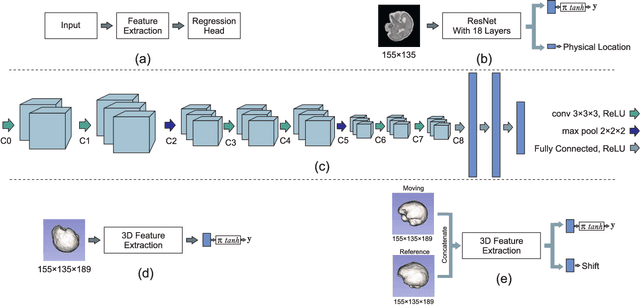
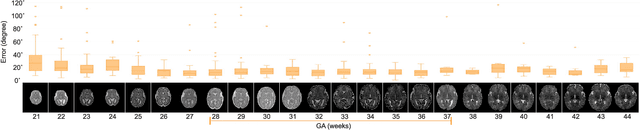
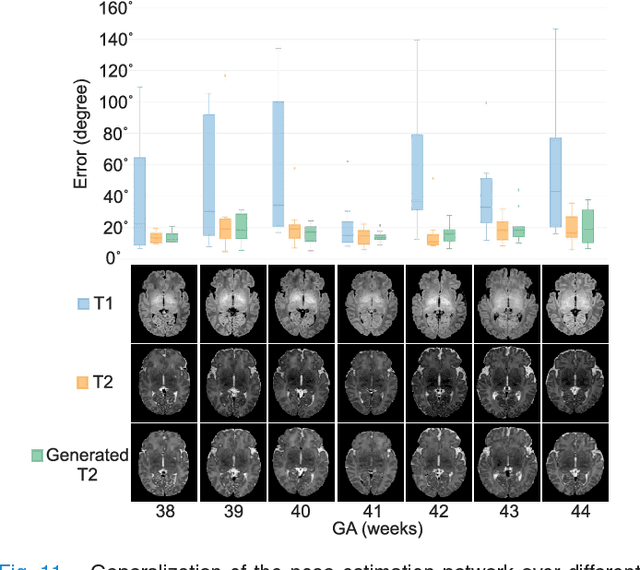
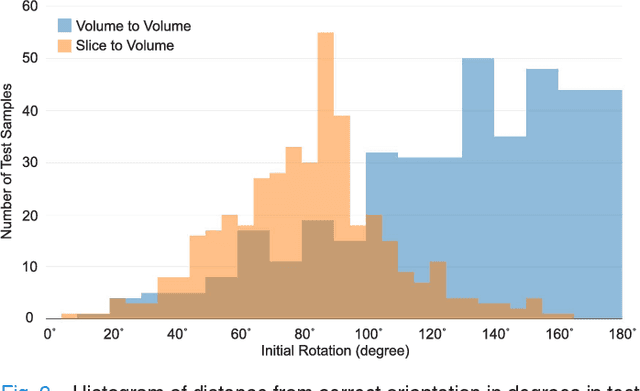
Abstract:With an aim to increase the capture range and accelerate the performance of state-of-the-art inter-subject and subject-to-template 3D registration, we propose deep learning-based methods that are trained to find the 3D position of arbitrarily oriented subjects or anatomy based on slices or volumes of medical images. For this, we propose regression CNNs that learn to predict the angle-axis representation of 3D rotations and translations using image features. We use and compare mean square error and geodesic loss to train regression CNNs for 3D pose estimation used in two different scenarios: slice-to-volume registration and volume-to-volume registration. Our results show that in such registration applications that are amendable to learning, the proposed deep learning methods with geodesic loss minimization can achieve accurate results with a wide capture range in real-time (<100ms). We also tested the generalization capability of the trained CNNs on an expanded age range and on images of newborn subjects with similar and different MR image contrasts. We trained our models on T2-weighted fetal brain MRI scans and used them to predict the 3D pose of newborn brains based on T1-weighted MRI scans. We showed that the trained models generalized well for the new domain when we performed image contrast transfer through a conditional generative adversarial network. This indicates that the domain of application of the trained deep regression CNNs can be further expanded to image modalities and contrasts other than those used in training. A combination of our proposed methods with accelerated optimization-based registration algorithms can dramatically enhance the performance of automatic imaging devices and image processing methods of the future.
Asymmetric Similarity Loss Function to Balance Precision and Recall in Highly Unbalanced Deep Medical Image Segmentation
Jun 29, 2018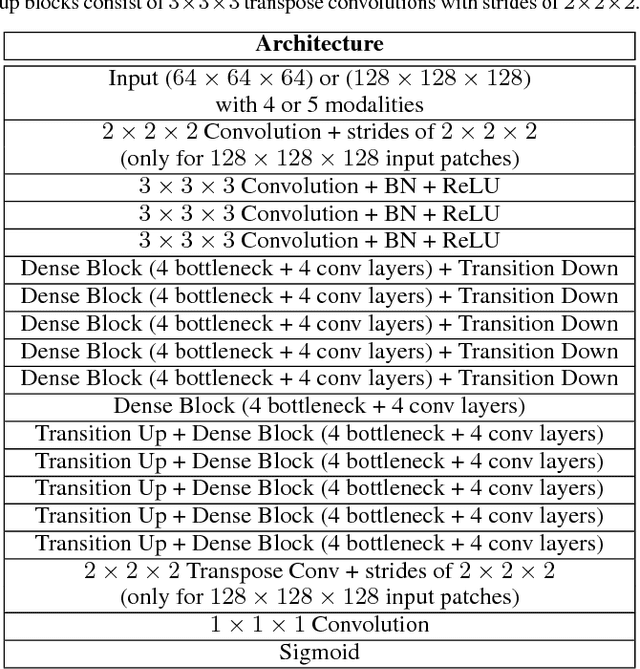
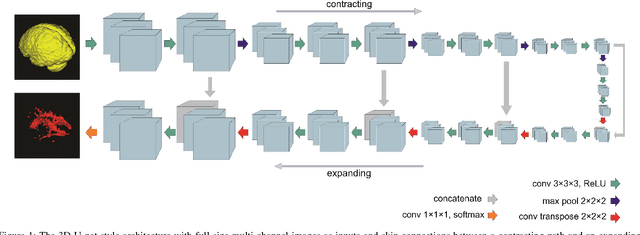
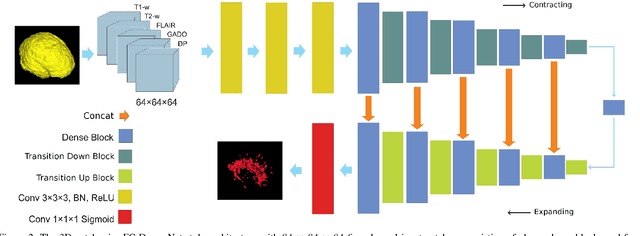
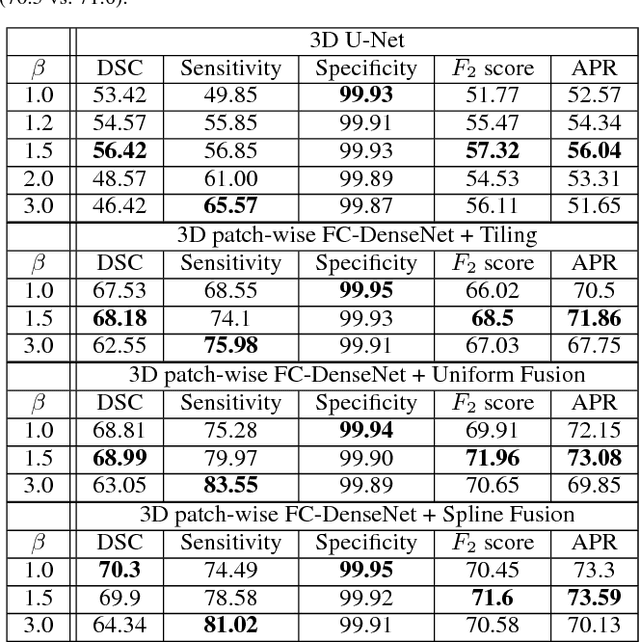
Abstract:Fully convolutional deep neural networks have been asserted to be fast and precise frameworks with great potential in image segmentation. One of the major challenges in utilizing such networks raises when data is unbalanced, which is common in many medical imaging applications such as lesion segmentation where lesion class voxels are often much lower in numbers than non-lesion voxels. A trained network with unbalanced data may make predictions with high precision and low recall, being severely biased towards the non-lesion class which is particularly undesired in medical applications where false negatives are actually more important than false positives. Various methods have been proposed to address this problem including two step training, sample re-weighting, balanced sampling, and similarity loss functions. In this paper we developed a patch-wise 3D densely connected network with an asymmetric loss function, where we used large overlapping image patches for intrinsic and extrinsic data augmentation, a patch selection algorithm, and a patch prediction fusion strategy based on B-spline weighted soft voting to take into account the uncertainty of prediction in patch borders. We applied this method to lesion segmentation based on the MSSEG 2016 and ISBI 2015 challenges, where we achieved average Dice similarity coefficient of 69.9% and 65.74%, respectively. In addition to the proposed loss, we trained our network with focal and generalized Dice loss functions. Significant improvement in $F_1$ and $F_2$ scores and the APR curve was achieved in test using the asymmetric similarity loss layer and our 3D patch prediction fusion. The asymmetric similarity loss based on $F_\beta$ scores generalizes the Dice similarity coefficient and can be effectively used with the patch-wise strategy developed here to train fully convolutional deep neural networks for highly unbalanced image segmentation.
Real-Time Automatic Fetal Brain Extraction in Fetal MRI by Deep Learning
Oct 25, 2017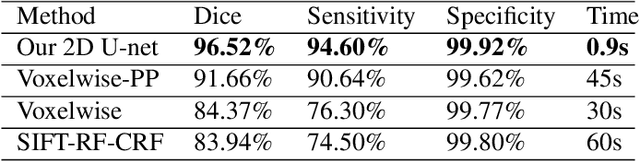
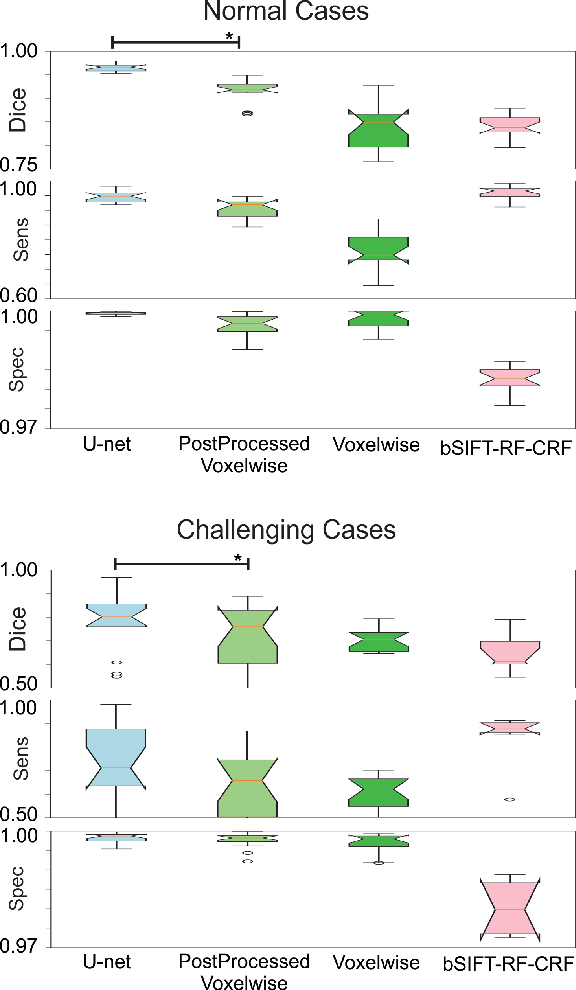
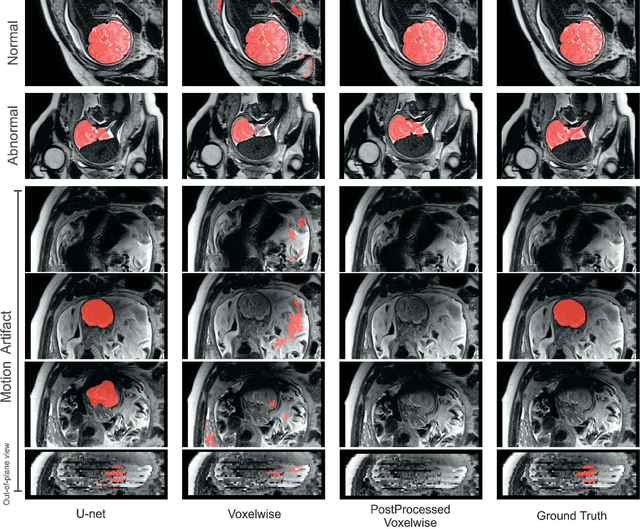
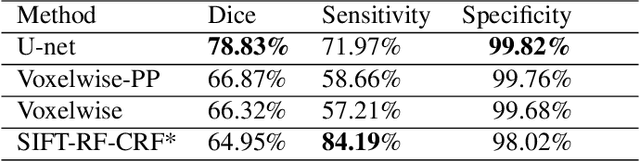
Abstract:Brain segmentation is a fundamental first step in neuroimage analysis. In the case of fetal MRI, it is particularly challenging and important due to the arbitrary orientation of the fetus, organs that surround the fetal head, and intermittent fetal motion. Several promising methods have been proposed but are limited in their performance in challenging cases and in real-time segmentation. We aimed to develop a fully automatic segmentation method that independently segments sections of the fetal brain in 2D fetal MRI slices in real-time. To this end, we developed and evaluated a deep fully convolutional neural network based on 2D U-net and autocontext, and compared it to two alternative fast methods based on 1) a voxelwise fully convolutional network and 2) a method based on SIFT features, random forest and conditional random field. We trained the networks with manual brain masks on 250 stacks of training images, and tested on 17 stacks of normal fetal brain images as well as 18 stacks of extremely challenging cases based on extreme motion, noise, and severely abnormal brain shape. Experimental results show that our U-net approach outperformed the other methods and achieved average Dice metrics of 96.52% and 78.83% in the normal and challenging test sets, respectively. With an unprecedented performance and a test run time of about 1 second, our network can be used to segment the fetal brain in real-time while fetal MRI slices are being acquired. This can enable real-time motion tracking, motion detection, and 3D reconstruction of fetal brain MRI.
Auto-context Convolutional Neural Network (Auto-Net) for Brain Extraction in Magnetic Resonance Imaging
Jun 19, 2017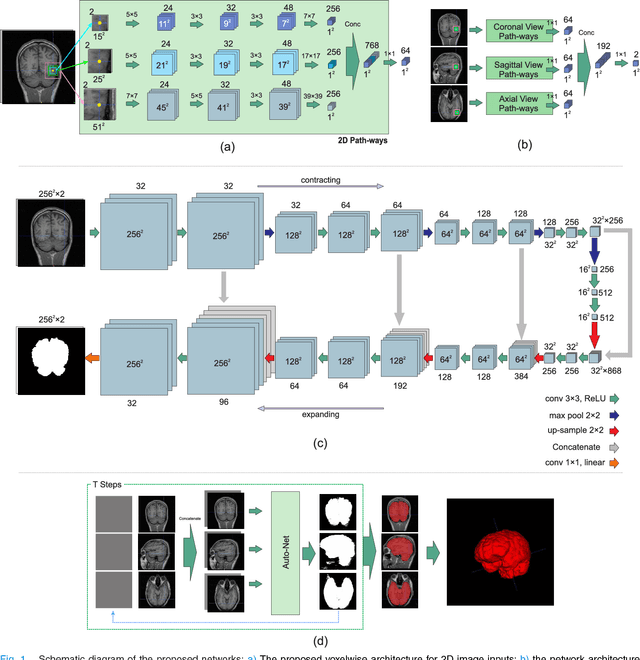
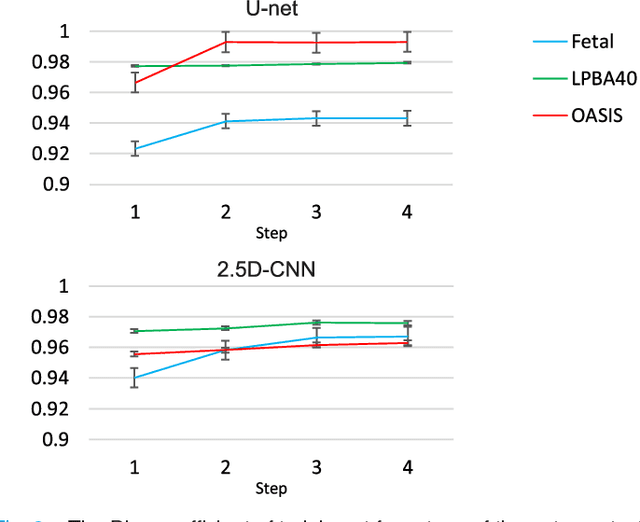
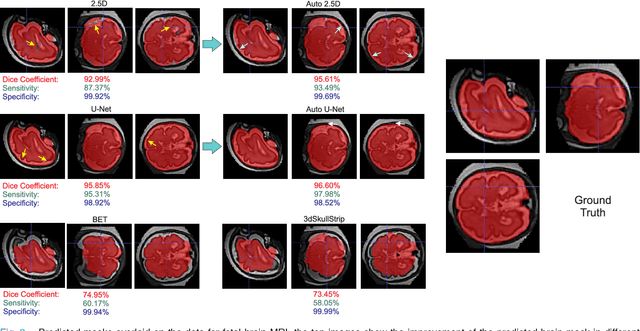

Abstract:Brain extraction or whole brain segmentation is an important first step in many of the neuroimage analysis pipelines. The accuracy and robustness of brain extraction, therefore, is crucial for the accuracy of the entire brain analysis process. With the aim of designing a learning-based, geometry-independent and registration-free brain extraction tool in this study, we present a technique based on an auto-context convolutional neural network (CNN), in which intrinsic local and global image features are learned through 2D patches of different window sizes. In this architecture three parallel 2D convolutional pathways for three different directions (axial, coronal, and sagittal) implicitly learn 3D image information without the need for computationally expensive 3D convolutions. Posterior probability maps generated by the network are used iteratively as context information along with the original image patches to learn the local shape and connectedness of the brain, to extract it from non-brain tissue. The brain extraction results we have obtained from our algorithm are superior to the recently reported results in the literature on two publicly available benchmark datasets, namely LPBA40 and OASIS, in which we obtained Dice overlap coefficients of 97.42% and 95.40%, respectively. Furthermore, we evaluated the performance of our algorithm in the challenging problem of extracting arbitrarily-oriented fetal brains in reconstructed fetal brain magnetic resonance imaging (MRI) datasets. In this application our algorithm performed much better than the other methods (Dice coefficient: 95.98%), where the other methods performed poorly due to the non-standard orientation and geometry of the fetal brain in MRI. Our CNN-based method can provide accurate, geometry-independent brain extraction in challenging applications.
Tversky loss function for image segmentation using 3D fully convolutional deep networks
Jun 18, 2017Abstract:Fully convolutional deep neural networks carry out excellent potential for fast and accurate image segmentation. One of the main challenges in training these networks is data imbalance, which is particularly problematic in medical imaging applications such as lesion segmentation where the number of lesion voxels is often much lower than the number of non-lesion voxels. Training with unbalanced data can lead to predictions that are severely biased towards high precision but low recall (sensitivity), which is undesired especially in medical applications where false negatives are much less tolerable than false positives. Several methods have been proposed to deal with this problem including balanced sampling, two step training, sample re-weighting, and similarity loss functions. In this paper, we propose a generalized loss function based on the Tversky index to address the issue of data imbalance and achieve much better trade-off between precision and recall in training 3D fully convolutional deep neural networks. Experimental results in multiple sclerosis lesion segmentation on magnetic resonance images show improved F2 score, Dice coefficient, and the area under the precision-recall curve in test data. Based on these results we suggest Tversky loss function as a generalized framework to effectively train deep neural networks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge