Sanjay P. Prabhu
BraTS-PEDs: Results of the Multi-Consortium International Pediatric Brain Tumor Segmentation Challenge 2023
Jul 11, 2024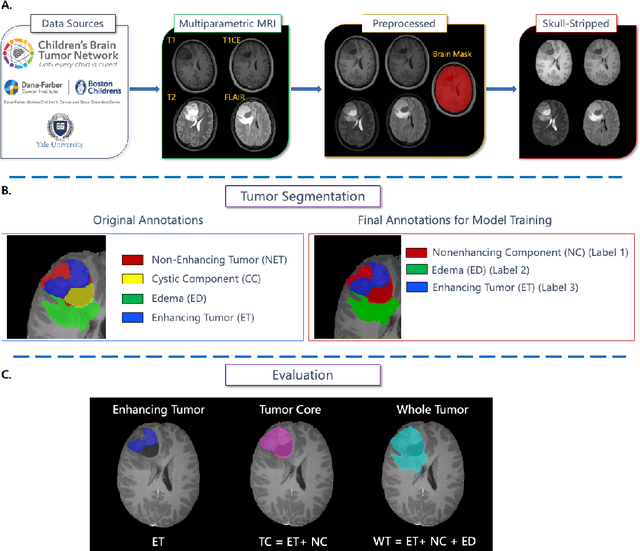

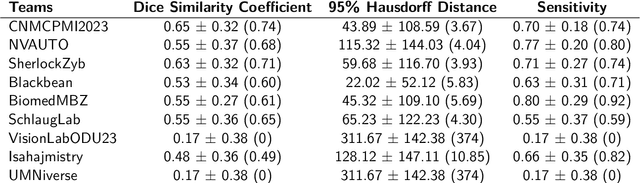
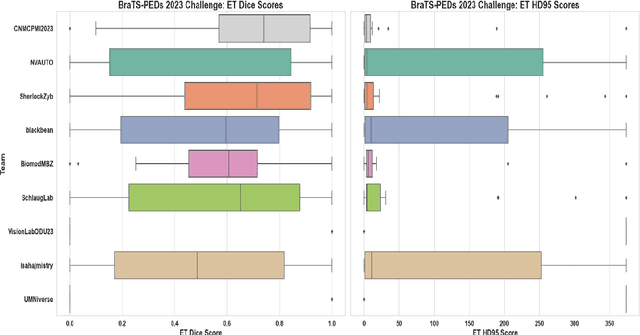
Abstract:Pediatric central nervous system tumors are the leading cause of cancer-related deaths in children. The five-year survival rate for high-grade glioma in children is less than 20%. The development of new treatments is dependent upon multi-institutional collaborative clinical trials requiring reproducible and accurate centralized response assessment. We present the results of the BraTS-PEDs 2023 challenge, the first Brain Tumor Segmentation (BraTS) challenge focused on pediatric brain tumors. This challenge utilized data acquired from multiple international consortia dedicated to pediatric neuro-oncology and clinical trials. BraTS-PEDs 2023 aimed to evaluate volumetric segmentation algorithms for pediatric brain gliomas from magnetic resonance imaging using standardized quantitative performance evaluation metrics employed across the BraTS 2023 challenges. The top-performing AI approaches for pediatric tumor analysis included ensembles of nnU-Net and Swin UNETR, Auto3DSeg, or nnU-Net with a self-supervised framework. The BraTSPEDs 2023 challenge fostered collaboration between clinicians (neuro-oncologists, neuroradiologists) and AI/imaging scientists, promoting faster data sharing and the development of automated volumetric analysis techniques. These advancements could significantly benefit clinical trials and improve the care of children with brain tumors.
Exclusive Independent Probability Estimation using Deep 3D Fully Convolutional DenseNets for IsoIntense Infant Brain MRI Segmentation
Sep 27, 2018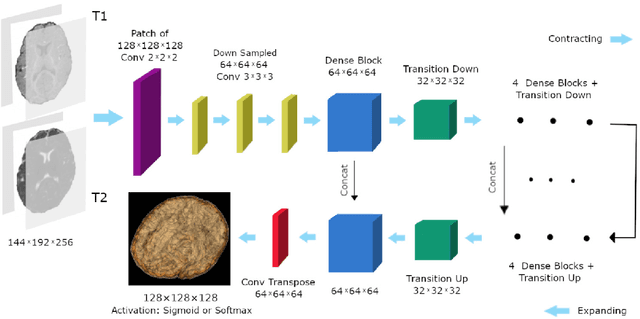

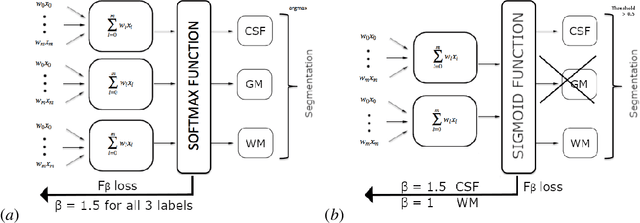
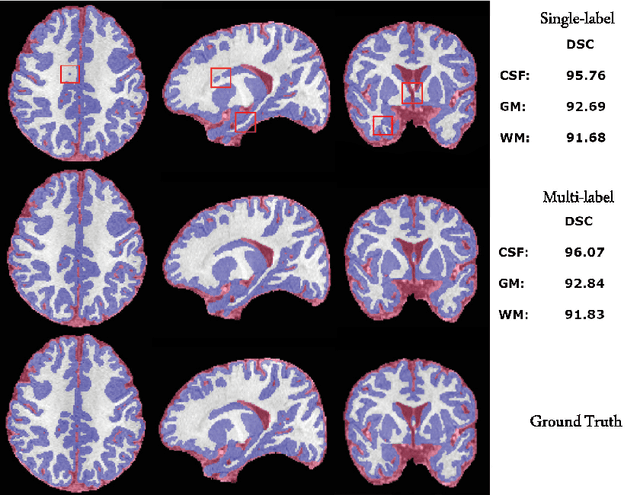
Abstract:The most recent fast and accurate image segmentation methods are built upon fully convolutional deep neural networks. Infant brain MRI tissue segmentation is a complex deep learning task, where the white matter and gray matter of the developing brain at about 6 months of age show similar T1 and T2 relaxation times, having similar intensity values on both T1 and T2-weighted MRIs. In this paper, we propose deep learning strategies to overcome the challenges of isointense infant brain MRI tissue segmentation. We introduce an exclusive multi-label training method to independently segment the mutually exclusive brain tissues with similarity loss function parameters that are balanced based on class prevalence. Using our training technique based on similarity loss functions and patch prediction fusion we decrease the number of parameters in the network, reduce the complexity of the training process focusing the attention on less number of tasks, while mitigating the effects of data imbalance between labels and inaccuracies near patch borders. By taking advantage of these strategies we were able to perform fast image segmentation, using a network with less parameters than many state-of-the-art networks, being image size independent overcoming issues such as 3D vs 2D training and large vs small patch size selection, while achieving the top performance in segmenting brain tissue among all methods in the 2017 iSeg challenge. We present a 3D FC-DenseNet architecture, an exclusive multilabel patchwise training technique with balanced similarity loss functions and a patch prediction fusion strategy that can be used on new classification and segmentation applications with two or more very similar classes. This strategy improves the training process by reducing its complexity while providing a trained model that works for any size input and is fast and more accurate than many state-of-the-art methods.
Asymmetric Similarity Loss Function to Balance Precision and Recall in Highly Unbalanced Deep Medical Image Segmentation
Jun 29, 2018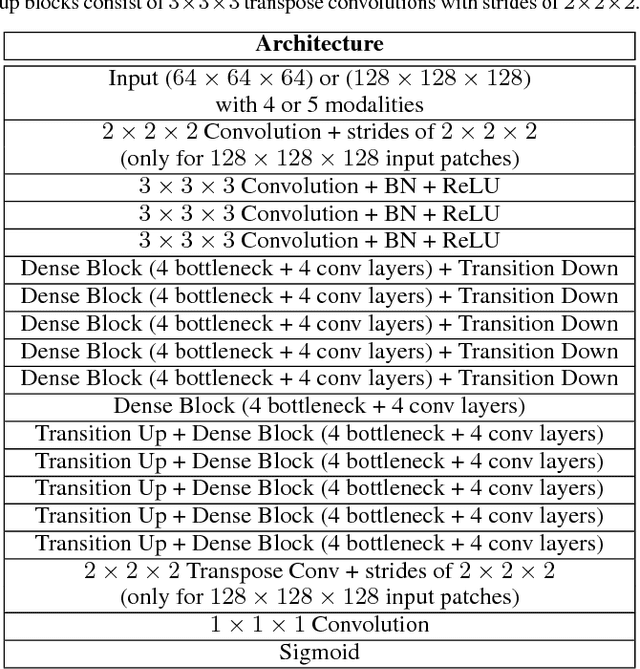
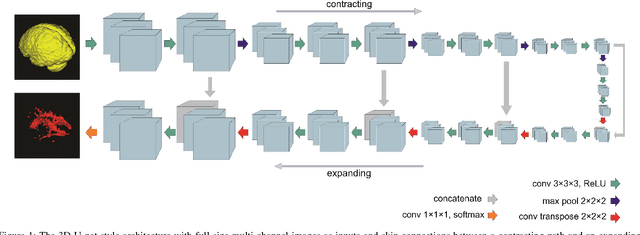
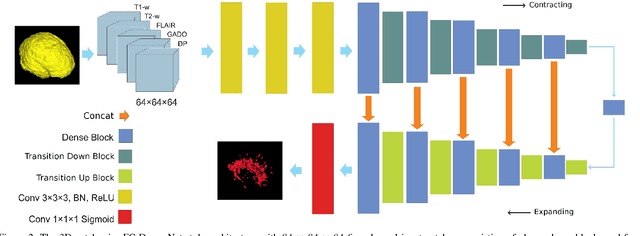
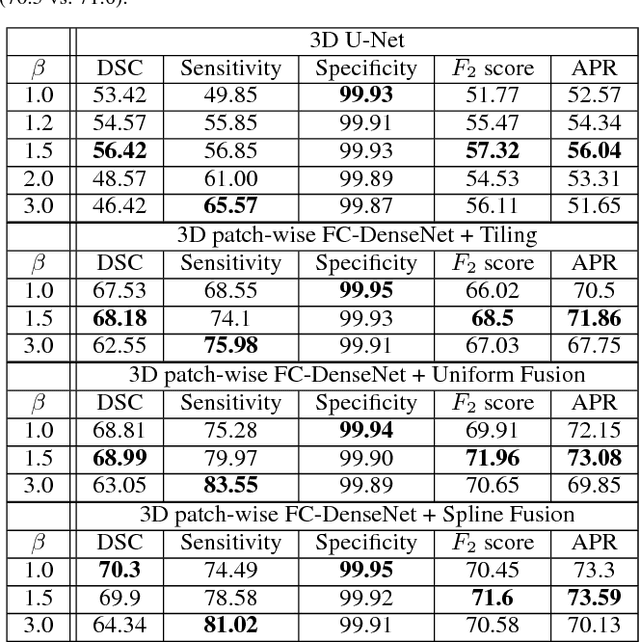
Abstract:Fully convolutional deep neural networks have been asserted to be fast and precise frameworks with great potential in image segmentation. One of the major challenges in utilizing such networks raises when data is unbalanced, which is common in many medical imaging applications such as lesion segmentation where lesion class voxels are often much lower in numbers than non-lesion voxels. A trained network with unbalanced data may make predictions with high precision and low recall, being severely biased towards the non-lesion class which is particularly undesired in medical applications where false negatives are actually more important than false positives. Various methods have been proposed to address this problem including two step training, sample re-weighting, balanced sampling, and similarity loss functions. In this paper we developed a patch-wise 3D densely connected network with an asymmetric loss function, where we used large overlapping image patches for intrinsic and extrinsic data augmentation, a patch selection algorithm, and a patch prediction fusion strategy based on B-spline weighted soft voting to take into account the uncertainty of prediction in patch borders. We applied this method to lesion segmentation based on the MSSEG 2016 and ISBI 2015 challenges, where we achieved average Dice similarity coefficient of 69.9% and 65.74%, respectively. In addition to the proposed loss, we trained our network with focal and generalized Dice loss functions. Significant improvement in $F_1$ and $F_2$ scores and the APR curve was achieved in test using the asymmetric similarity loss layer and our 3D patch prediction fusion. The asymmetric similarity loss based on $F_\beta$ scores generalizes the Dice similarity coefficient and can be effectively used with the patch-wise strategy developed here to train fully convolutional deep neural networks for highly unbalanced image segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge