Natasha Thorley
Deep EM with Hierarchical Latent Label Modelling for Multi-Site Prostate Lesion Segmentation
Mar 15, 2026Abstract:Label variability is a major challenge for prostate lesion segmentation. In multi-site datasets, annotations often reflect centre-specific contouring protocols, causing segmentation networks to overfit to local styles and generalise poorly to unseen sites in inference. We treat each observed annotation as a noisy observation of an underlying latent 'clean' lesion mask, and propose a hierarchical expectation-maximisation (HierEM) framework that alternates between: (1) inferring a voxel-wise posterior distribution over the latent mask, and (2) training a CNN using this posterior as a soft target and estimate site-specific sensitivity and specificity under a hierarchical prior. This hierarchical prior decomposes label-quality into a global mean with site- and case-level deviations, reducing site-specific bias by penalising the likelihood term contributed only by site deviations. Experiments on three cohorts demonstrate that the proposed hierarchical EM framework enhances cross-site generalisation compared to state-of-the-art methods. For pooled-dataset evaluation, the per-site mean DSC ranges from 29.50% to 39.69%; for leave-one-site-out generalisation, it ranges from 27.91% to 32.67%, yielding statistically significant improvements over comparison methods (p<0.039). The method also produces interpretable per-site latent label-quality estimates (sensitivity alpha ranges from 31.5% to 47.3% at specificity beta approximates 0.99), supporting post-hoc analyses of cross-site annotation variability. These results indicate that explicitly modelling site-dependent annotation can improve cross-site generalisation.
ProFound: A moderate-sized vision foundation model for multi-task prostate imaging
Mar 04, 2026Abstract:Many diagnostic and therapeutic clinical tasks for prostate cancer increasingly rely on multi-parametric MRI. Automating these tasks is challenging because they necessitate expert interpretations, which are difficult to scale to capitalise on modern deep learning. Although modern automated systems achieve expert-level performance in isolated tasks, their general clinical utility remains limited by the requirement of large task-specific labelled datasets. In this paper, we present ProFound, a domain-specialised vision foundation model for volumetric prostate mpMRI. ProFound is pre-trained using several variants of self-supervised approaches on a diverse, multi-institutional collection of 5,000 patients, with a total of over 22,000 unique 3D MRI volumes (over 1,800,000 2D image slices). We conducted a systematic evaluation of ProFound across a broad spectrum of $11$ downstream clinical tasks on over 3,000 independent patients, including prostate cancer detection, Gleason grading, lesion localisation, gland volume estimation, zonal and surrounding structure segmentation. Experimental results demonstrate that finetuned ProFound consistently outperforms or remains competitive with state-of-the-art specialised models and existing medical vision foundation models trained/finetuned on the same data.
Understanding the Transfer Limits of Vision Foundation Models
Jan 22, 2026Abstract:Foundation models leverage large-scale pretraining to capture extensive knowledge, demonstrating generalization in a wide range of language tasks. By comparison, vision foundation models (VFMs) often exhibit uneven improvements across downstream tasks, despite substantial computational investment. We postulate that this limitation arises from a mismatch between pretraining objectives and the demands of downstream vision-and-imaging tasks. Pretraining strategies like masked image reconstruction or contrastive learning shape representations for tasks such as recovery of generic visual patterns or global semantic structures, which may not align with the task-specific requirements of downstream applications including segmentation, classification, or image synthesis. To investigate this in a concrete real-world clinical area, we assess two VFMs, a reconstruction-focused MAE-based model (ProFound) and a contrastive-learning-based model (ProViCNet), on five prostate multiparametric MR imaging tasks, examining how such task alignment influences transfer performance, i.e., from pretraining to fine-tuning. Our findings indicate that better alignment between pretraining and downstream tasks, measured by simple divergence metrics such as maximum-mean-discrepancy (MMD) between the same features before and after fine-tuning, correlates with greater performance improvements and faster convergence, emphasizing the importance of designing and analyzing pretraining objectives with downstream applicability in mind.
T2-Only Prostate Cancer Prediction by Meta-Learning from Bi-Parametric MR Imaging
Nov 11, 2024


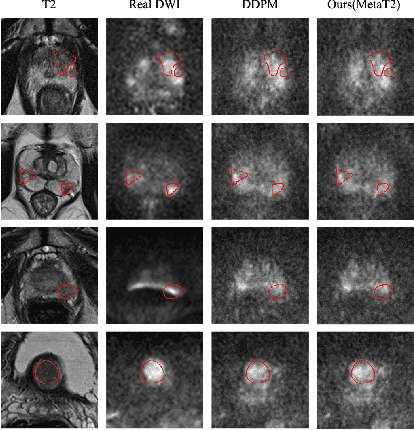
Abstract:Current imaging-based prostate cancer diagnosis requires both MR T2-weighted (T2w) and diffusion-weighted imaging (DWI) sequences, with additional sequences for potentially greater accuracy improvement. However, measuring diffusion patterns in DWI sequences can be time-consuming, prone to artifacts and sensitive to imaging parameters. While machine learning (ML) models have demonstrated radiologist-level accuracy in detecting prostate cancer from these two sequences, this study investigates the potential of ML-enabled methods using only the T2w sequence as input during inference time. We first discuss the technical feasibility of such a T2-only approach, and then propose a novel ML formulation, where DWI sequences - readily available for training purposes - are only used to train a meta-learning model, which subsequently only uses T2w sequences at inference. Using multiple datasets from more than 3,000 prostate cancer patients, we report superior or comparable performance in localising radiologist-identified prostate cancer using our proposed T2-only models, compared with alternative models using T2-only or both sequences as input. Real patient cases are presented and discussed to demonstrate, for the first time, the exclusively true-positive cases from models with different input sequences.
AI-assisted prostate cancer detection and localisation on biparametric MR by classifying radiologist-positives
Oct 30, 2024



Abstract:Prostate cancer diagnosis through MR imaging have currently relied on radiologists' interpretation, whilst modern AI-based methods have been developed to detect clinically significant cancers independent of radiologists. In this study, we propose to develop deep learning models that improve the overall cancer diagnostic accuracy, by classifying radiologist-identified patients or lesions (i.e. radiologist-positives), as opposed to the existing models that are trained to discriminate over all patients. We develop a single voxel-level classification model, with a simple percentage threshold to determine positive cases, at levels of lesions, Barzell-zones and patients. Based on the presented experiments from two clinical data sets, consisting of histopathology-labelled MR images from more than 800 and 500 patients in the respective UCLA and UCL PROMIS studies, we show that the proposed strategy can improve the diagnostic accuracy, by augmenting the radiologist reading of the MR imaging. Among varying definition of clinical significance, the proposed strategy, for example, achieved a specificity of 44.1% (with AI assistance) from 36.3% (by radiologists alone), at a controlled sensitivity of 80.0% on the publicly available UCLA data set. This provides measurable clinical values in a range of applications such as reducing unnecessary biopsies, lowering cost in cancer screening and quantifying risk in therapies.
Poisson Ordinal Network for Gleason Group Estimation Using Bi-Parametric MRI
Jul 08, 2024



Abstract:The Gleason groups serve as the primary histological grading system for prostate cancer, providing crucial insights into the cancer's potential for growth and metastasis. In clinical practice, pathologists determine the Gleason groups based on specimens obtained from ultrasound-guided biopsies. In this study, we investigate the feasibility of directly estimating the Gleason groups from MRI scans to reduce otherwise required biopsies. We identify two characteristics of this task, ordinality and the resulting dependent yet unknown variances between Gleason groups. In addition to the inter- / intra- observer variability in a multi-step Gleason scoring process based on the interpretation of Gleason patterns, our MR-based prediction is also subject to specimen sampling variance and, to a lesser degree, varying MR imaging protocols. To address this challenge, we propose a novel Poisson ordinal network (PON). PONs model the prediction using a Poisson distribution and leverages Poisson encoding and Poisson focal loss to capture a learnable dependency between ordinal classes (here, Gleason groups), rather than relying solely on the numerical ground-truth (e.g. Gleason Groups 1-5 or Gleason Scores 6-10). To improve this modelling efficacy, PONs also employ contrastive learning with a memory bank to regularise intra-class variance, decoupling the memory requirement of contrast learning from the batch size. Experimental results based on the images labelled by saturation biopsies from 265 prior-biopsy-blind patients, across two tasks demonstrate the superiority and effectiveness of our proposed method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge