Mohammad Areeb Qazi
Quran-MD: A Fine-Grained Multilingual Multimodal Dataset of the Quran
Jan 25, 2026Abstract:We present Quran MD, a comprehensive multimodal dataset of the Quran that integrates textual, linguistic, and audio dimensions at the verse and word levels. For each verse (ayah), the dataset provides its original Arabic text, English translation, and phonetic transliteration. To capture the rich oral tradition of Quranic recitation, we include verse-level audio from 32 distinct reciters, reflecting diverse recitation styles and dialectical nuances. At the word level, each token is paired with its corresponding Arabic script, English translation, transliteration, and an aligned audio recording, allowing fine-grained analysis of pronunciation, phonology, and semantic context. This dataset supports various applications, including natural language processing, speech recognition, text-to-speech synthesis, linguistic analysis, and digital Islamic studies. Bridging text and audio modalities across multiple reciters, this dataset provides a unique resource to advance computational approaches to Quranic recitation and study. Beyond enabling tasks such as ASR, tajweed detection, and Quranic TTS, it lays the foundation for multimodal embeddings, semantic retrieval, style transfer, and personalized tutoring systems that can support both research and community applications. The dataset is available at https://huggingface.co/datasets/Buraaq/quran-audio-text-dataset
MAFM^3: Modular Adaptation of Foundation Models for Multi-Modal Medical AI
Nov 14, 2025Abstract:Foundational models are trained on extensive datasets to capture the general trends of a domain. However, in medical imaging, the scarcity of data makes pre-training for every domain, modality, or task challenging. Instead of building separate models, we propose MAFM^3 (Modular Adaptation of Foundation Models for Multi-Modal Medical AI), a framework that enables a single foundation model to expand into diverse domains, tasks, and modalities through lightweight modular components. These components serve as specialized skill sets that allow the system to flexibly activate the appropriate capability at the inference time, depending on the input type or clinical objective. Unlike conventional adaptation methods that treat each new task or modality in isolation, MAFM^3 provides a unified and expandable framework for efficient multitask and multimodality adaptation. Empirically, we validate our approach by adapting a chest CT foundation model initially trained for classification into prognosis and segmentation modules. Our results show improved performance on both tasks. Furthermore, by incorporating PET scans, MAFM^3 achieved an improvement in the Dice score 5% compared to the respective baselines. These findings establish that foundation models, when equipped with modular components, are not inherently constrained to their initial training scope but can evolve into multitask, multimodality systems for medical imaging. The code implementation of this work can be found at https://github.com/Areeb2735/CTscan_prognosis_VLM
PALM: Few-Shot Prompt Learning for Audio Language Models
Sep 29, 2024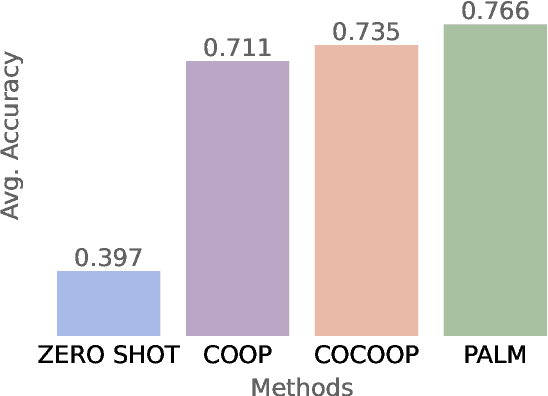
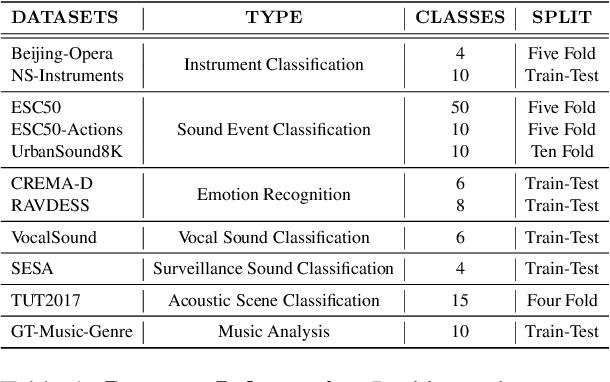
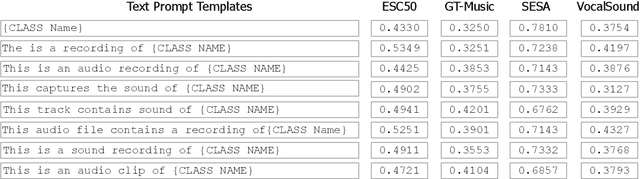
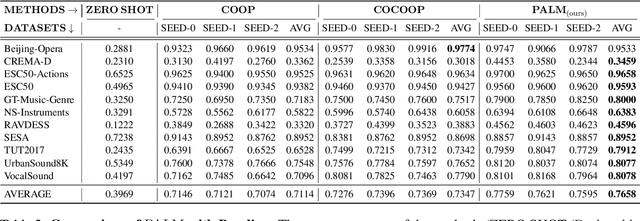
Abstract:Audio-Language Models (ALMs) have recently achieved remarkable success in zero-shot audio recognition tasks, which match features of audio waveforms with class-specific text prompt features, inspired by advancements in Vision-Language Models (VLMs). Given the sensitivity of zero-shot performance to the choice of hand-crafted text prompts, many prompt learning techniques have been developed for VLMs. We explore the efficacy of these approaches in ALMs and propose a novel method, Prompt Learning in Audio Language Models (PALM), which optimizes the feature space of the text encoder branch. Unlike existing methods that work in the input space, our approach results in greater training efficiency. We demonstrate the effectiveness of our approach on 11 audio recognition datasets, encompassing a variety of speech-processing tasks, and compare the results with three baselines in a few-shot learning setup. Our method is either on par with or outperforms other approaches while being computationally less demanding. Code is available at https://asif-hanif.github.io/palm/
Introducing SDICE: An Index for Assessing Diversity of Synthetic Medical Datasets
Sep 28, 2024



Abstract:Advancements in generative modeling are pushing the state-of-the-art in synthetic medical image generation. These synthetic images can serve as an effective data augmentation method to aid the development of more accurate machine learning models for medical image analysis. While the fidelity of these synthetic images has progressively increased, the diversity of these images is an understudied phenomenon. In this work, we propose the SDICE index, which is based on the characterization of similarity distributions induced by a contrastive encoder. Given a synthetic dataset and a reference dataset of real images, the SDICE index measures the distance between the similarity score distributions of original and synthetic images, where the similarity scores are estimated using a pre-trained contrastive encoder. This distance is then normalized using an exponential function to provide a consistent metric that can be easily compared across domains. Experiments conducted on the MIMIC-chest X-ray and ImageNet datasets demonstrate the effectiveness of SDICE index in assessing synthetic medical dataset diversity.
Continual Learning in Medical Imaging from Theory to Practice: A Survey and Practical Analysis
May 22, 2024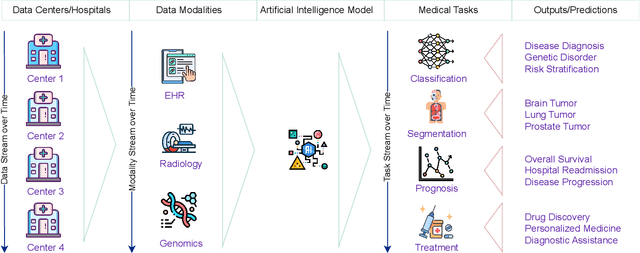
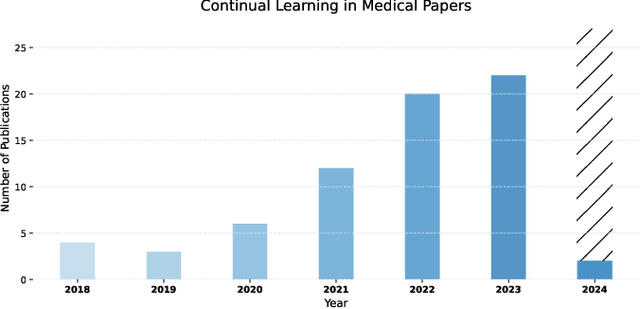

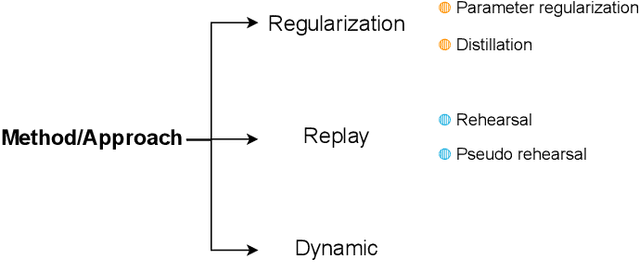
Abstract:Deep Learning has shown great success in reshaping medical imaging, yet it faces numerous challenges hindering widespread application. Issues like catastrophic forgetting and distribution shifts in the continuously evolving data stream increase the gap between research and applications. Continual Learning offers promise in addressing these hurdles by enabling the sequential acquisition of new knowledge without forgetting previous learnings in neural networks. In this survey, we comprehensively review the recent literature on continual learning in the medical domain, highlight recent trends, and point out the practical issues. Specifically, we survey the continual learning studies on classification, segmentation, detection, and other tasks in the medical domain. Furthermore, we develop a taxonomy for the reviewed studies, identify the challenges, and provide insights to overcome them. We also critically discuss the current state of continual learning in medical imaging, including identifying open problems and outlining promising future directions. We hope this survey will provide researchers with a useful overview of the developments in the field and will further increase interest in the community. To keep up with the fast-paced advancements in this field, we plan to routinely update the repository with the latest relevant papers at https://github.com/BioMedIA-MBZUAI/awesome-cl-in-medical .
DynaMMo: Dynamic Model Merging for Efficient Class Incremental Learning for Medical Images
Apr 22, 2024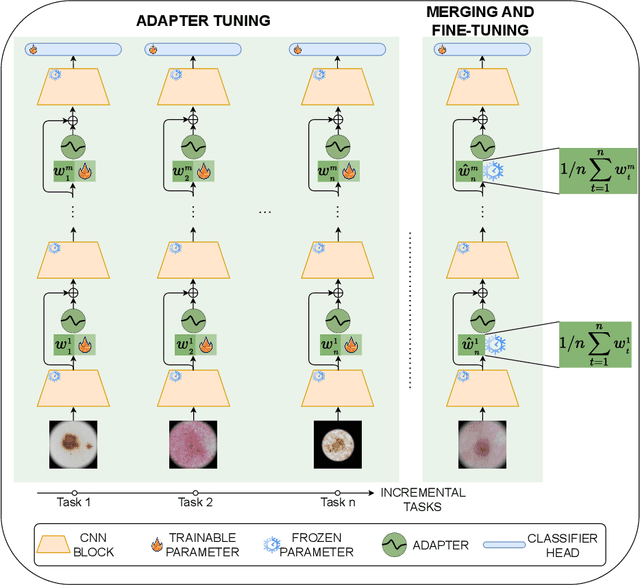
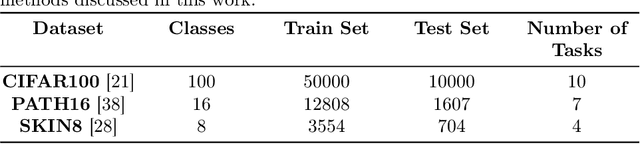
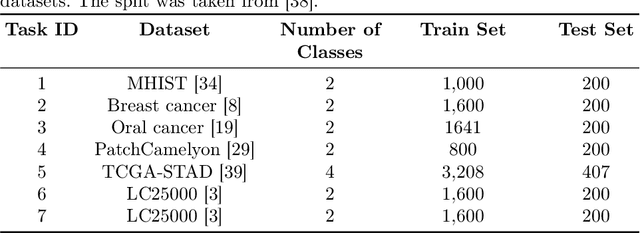
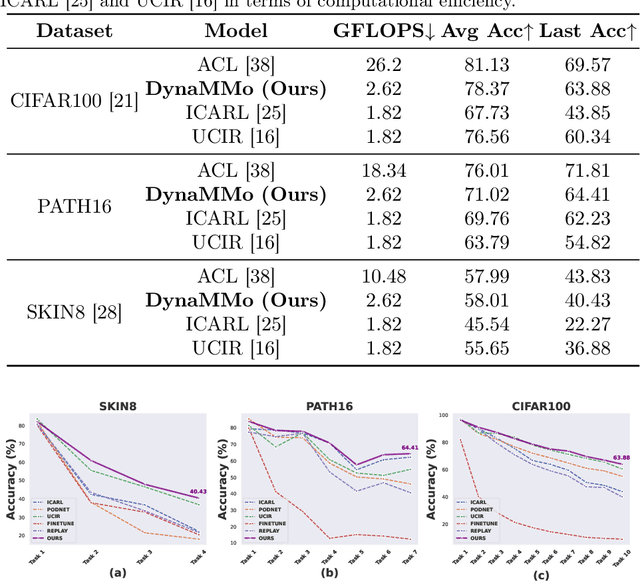
Abstract:Continual learning, the ability to acquire knowledge from new data while retaining previously learned information, is a fundamental challenge in machine learning. Various approaches, including memory replay, knowledge distillation, model regularization, and dynamic network expansion, have been proposed to address this issue. Thus far, dynamic network expansion methods have achieved state-of-the-art performance at the cost of incurring significant computational overhead. This is due to the need for additional model buffers, which makes it less feasible in resource-constrained settings, particularly in the medical domain. To overcome this challenge, we propose Dynamic Model Merging, DynaMMo, a method that merges multiple networks at different stages of model training to achieve better computational efficiency. Specifically, we employ lightweight learnable modules for each task and combine them into a unified model to minimize computational overhead. DynaMMo achieves this without compromising performance, offering a cost-effective solution for continual learning in medical applications. We evaluate DynaMMo on three publicly available datasets, demonstrating its effectiveness compared to existing approaches. DynaMMo offers around 10-fold reduction in GFLOPS with a small drop of 2.76 in average accuracy when compared to state-of-the-art dynamic-based approaches. The code implementation of this work will be available upon the acceptance of this work at https://github.com/BioMedIA-MBZUAI/DynaMMo.
FissionFusion: Fast Geometric Generation and Hierarchical Souping for Medical Image Analysis
Mar 20, 2024



Abstract:The scarcity of well-annotated medical datasets requires leveraging transfer learning from broader datasets like ImageNet or pre-trained models like CLIP. Model soups averages multiple fine-tuned models aiming to improve performance on In-Domain (ID) tasks and enhance robustness against Out-of-Distribution (OOD) datasets. However, applying these methods to the medical imaging domain faces challenges and results in suboptimal performance. This is primarily due to differences in error surface characteristics that stem from data complexities such as heterogeneity, domain shift, class imbalance, and distributional shifts between training and testing phases. To address this issue, we propose a hierarchical merging approach that involves local and global aggregation of models at various levels based on models' hyperparameter configurations. Furthermore, to alleviate the need for training a large number of models in the hyperparameter search, we introduce a computationally efficient method using a cyclical learning rate scheduler to produce multiple models for aggregation in the weight space. Our method demonstrates significant improvements over the model souping approach across multiple datasets (around 6% gain in HAM10000 and CheXpert datasets) while maintaining low computational costs for model generation and selection. Moreover, we achieve better results on OOD datasets than model soups. The code is available at https://github.com/BioMedIA-MBZUAI/FissionFusion.
MedMerge: Merging Models for Effective Transfer Learning to Medical Imaging Tasks
Mar 18, 2024



Abstract:Transfer learning has become a powerful tool to initialize deep learning models to achieve faster convergence and higher performance. This is especially useful in the medical imaging analysis domain, where data scarcity limits possible performance gains for deep learning models. Some advancements have been made in boosting the transfer learning performance gain by merging models starting from the same initialization. However, in the medical imaging analysis domain, there is an opportunity in merging models starting from different initialisations, thus combining the features learnt from different tasks. In this work, we propose MedMerge, a method whereby the weights of different models can be merged, and their features can be effectively utilized to boost performance on a new task. With MedMerge, we learn kernel-level weights that can later be used to merge the models into a single model, even when starting from different initializations. Testing on various medical imaging analysis tasks, we show that our merged model can achieve significant performance gains, with up to 3% improvement on the F1 score. The code implementation of this work will be available at www.github.com/BioMedIA-MBZUAI/MedMerge.
XReal: Realistic Anatomy and Pathology-Aware X-ray Generation via Controllable Diffusion Model
Mar 14, 2024



Abstract:Large-scale generative models have demonstrated impressive capacity in producing visually compelling images, with increasing applications in medical imaging. However, they continue to grapple with the challenge of image hallucination and the generation of anatomically inaccurate outputs. These limitations are mainly due to the sole reliance on textual inputs and lack of spatial control over the generated images, hindering the potential usefulness of such models in real-life settings. We present XReal, a novel controllable diffusion model for generating realistic chest X-ray images through precise anatomy and pathology location control. Our lightweight method can seamlessly integrate spatial control in a pre-trained text-to-image diffusion model without fine-tuning, retaining its existing knowledge while enhancing its generation capabilities. XReal outperforms state-of-the-art x-ray diffusion models in quantitative and qualitative metrics while showing 13% and 10% anatomy and pathology realism gain, respectively, based on the expert radiologist evaluation. Our model holds promise for advancing generative models in medical imaging, offering greater precision and adaptability while inviting further exploration in this evolving field. A large synthetically generated data with annotations and code is publicly available at https://github.com/BioMedIA-MBZUAI/XReal.
Multi-Task Learning Approach for Unified Biometric Estimation from Fetal Ultrasound Anomaly Scans
Nov 16, 2023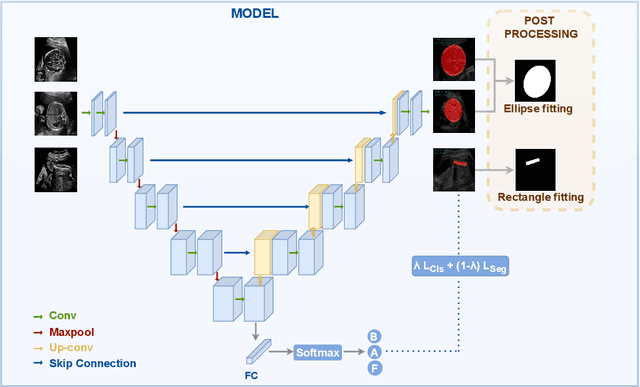
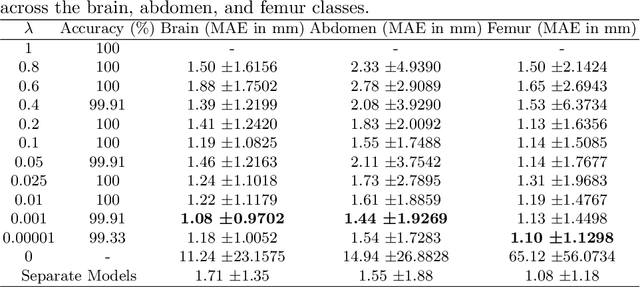

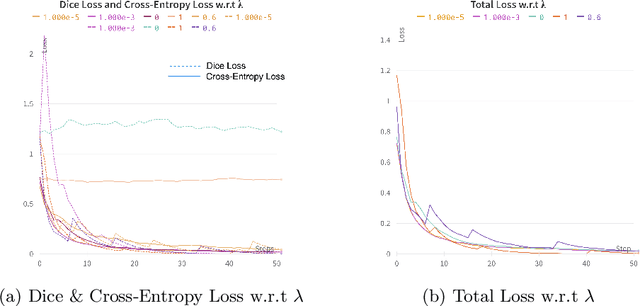
Abstract:Precise estimation of fetal biometry parameters from ultrasound images is vital for evaluating fetal growth, monitoring health, and identifying potential complications reliably. However, the automated computerized segmentation of the fetal head, abdomen, and femur from ultrasound images, along with the subsequent measurement of fetal biometrics, remains challenging. In this work, we propose a multi-task learning approach to classify the region into head, abdomen and femur as well as estimate the associated parameters. We were able to achieve a mean absolute error (MAE) of 1.08 mm on head circumference, 1.44 mm on abdomen circumference and 1.10 mm on femur length with a classification accuracy of 99.91\% on a dataset of fetal Ultrasound images. To achieve this, we leverage a weighted joint classification and segmentation loss function to train a U-Net architecture with an added classification head. The code can be accessed through \href{https://github.com/BioMedIA-MBZUAI/Multi-Task-Learning-Approach-for-Unified-Biometric-Estimation-from-Fetal-Ultrasound-Anomaly-Scans.git}{\texttt{Github}
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge