Matthew Welborn
NeuralPLexer3: Accurate Biomolecular Complex Structure Prediction with Flow Models
Dec 18, 2024
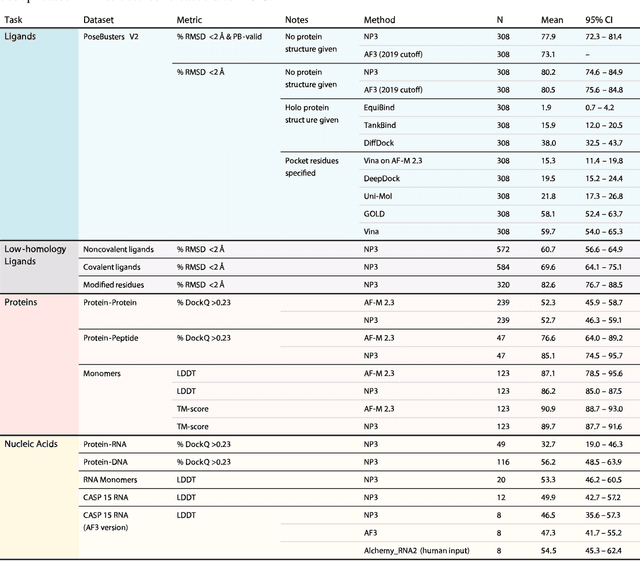
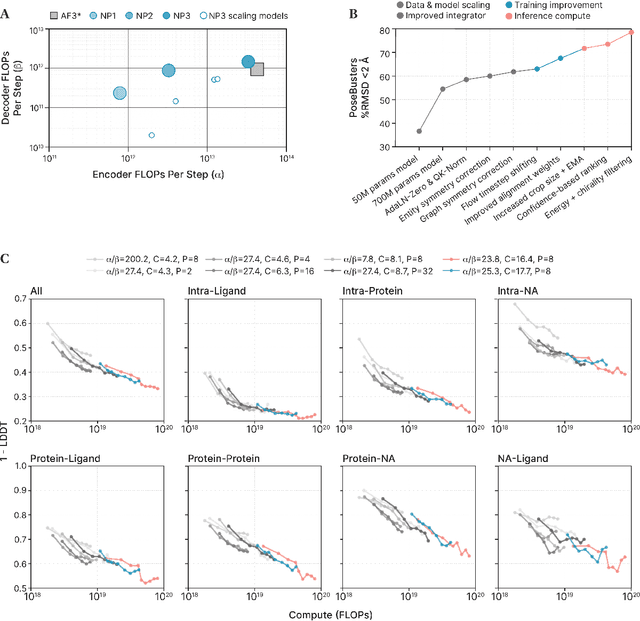
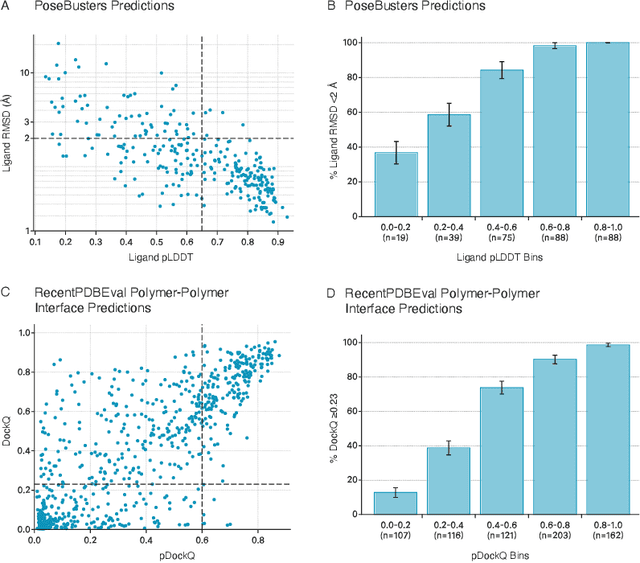
Abstract:Structure determination is essential to a mechanistic understanding of diseases and the development of novel therapeutics. Machine-learning-based structure prediction methods have made significant advancements by computationally predicting protein and bioassembly structures from sequences and molecular topology alone. Despite substantial progress in the field, challenges remain to deliver structure prediction models to real-world drug discovery. Here, we present NeuralPLexer3 -- a physics-inspired flow-based generative model that achieves state-of-the-art prediction accuracy on key biomolecular interaction types and improves training and sampling efficiency compared to its predecessors and alternative methodologies. Examined through newly developed benchmarking strategies, NeuralPLexer3 excels in vital areas that are crucial to structure-based drug design, such as physical validity and ligand-induced conformational changes.
UNiTE: Unitary N-body Tensor Equivariant Network with Applications to Quantum Chemistry
Jun 06, 2021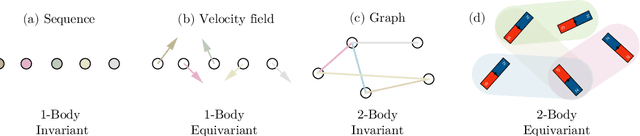
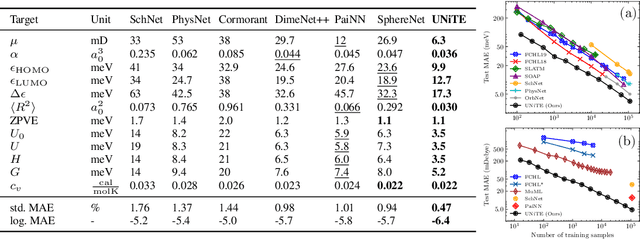
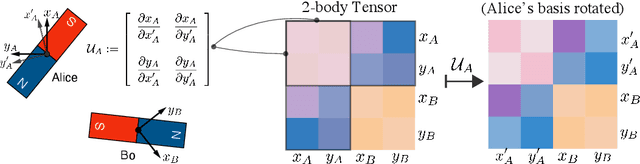
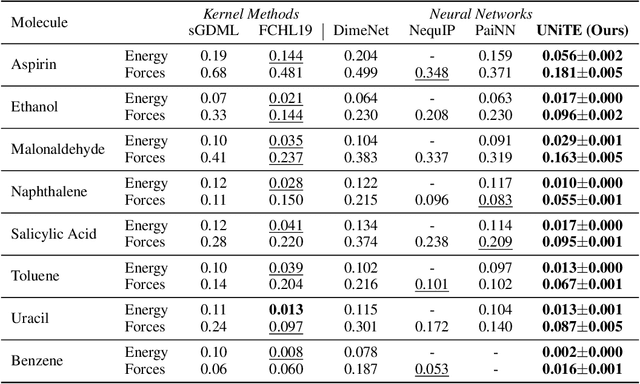
Abstract:Equivariant neural networks have been successful in incorporating various types of symmetries, but are mostly limited to vector representations of geometric objects. Despite the prevalence of higher-order tensors in various application domains, e.g. in quantum chemistry, equivariant neural networks for general tensors remain unexplored. Previous strategies for learning equivariant functions on tensors mostly rely on expensive tensor factorization which is not scalable when the dimensionality of the problem becomes large. In this work, we propose unitary $N$-body tensor equivariant neural network (UNiTE), an architecture for a general class of symmetric tensors called $N$-body tensors. The proposed neural network is equivariant with respect to the actions of a unitary group, such as the group of 3D rotations. Furthermore, it has a linear time complexity with respect to the number of non-zero elements in the tensor. We also introduce a normalization method, viz., Equivariant Normalization, to improve generalization of the neural network while preserving symmetry. When applied to quantum chemistry, UNiTE outperforms all state-of-the-art machine learning methods of that domain with over 110% average improvements on multiple benchmarks. Finally, we show that UNiTE achieves a robust zero-shot generalization performance on diverse down stream chemistry tasks, while being three orders of magnitude faster than conventional numerical methods with competitive accuracy.
Multi-task learning for electronic structure to predict and explore molecular potential energy surfaces
Nov 11, 2020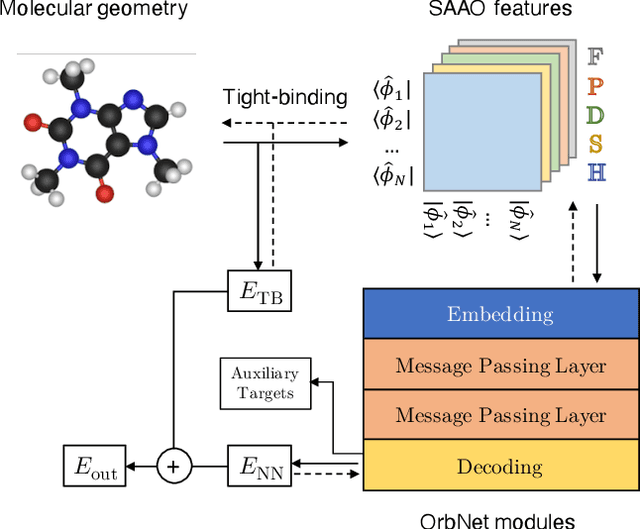

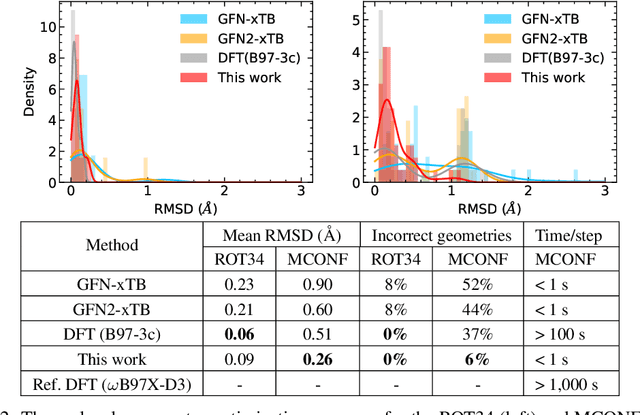
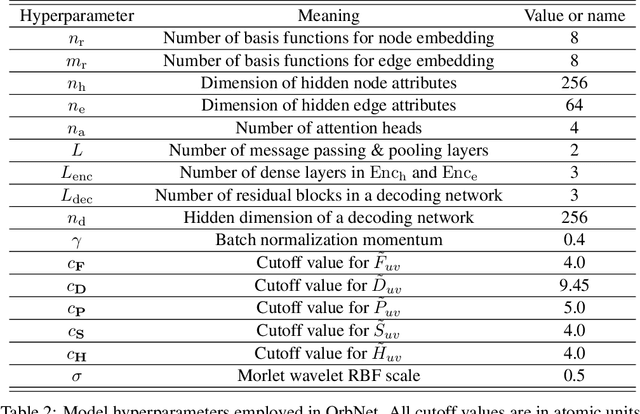
Abstract:We refine the OrbNet model to accurately predict energy, forces, and other response properties for molecules using a graph neural-network architecture based on features from low-cost approximated quantum operators in the symmetry-adapted atomic orbital basis. The model is end-to-end differentiable due to the derivation of analytic gradients for all electronic structure terms, and is shown to be transferable across chemical space due to the use of domain-specific features. The learning efficiency is improved by incorporating physically motivated constraints on the electronic structure through multi-task learning. The model outperforms existing methods on energy prediction tasks for the QM9 dataset and for molecular geometry optimizations on conformer datasets, at a computational cost that is thousand-fold or more reduced compared to conventional quantum-chemistry calculations (such as density functional theory) that offer similar accuracy.
OrbNet: Deep Learning for Quantum Chemistry Using Symmetry-Adapted Atomic-Orbital Features
Jul 15, 2020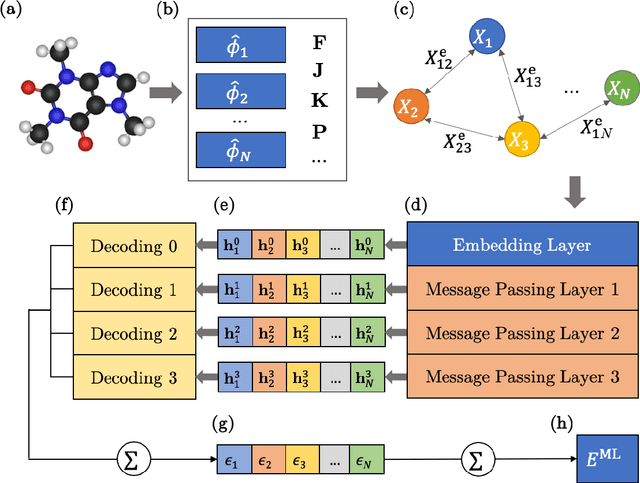
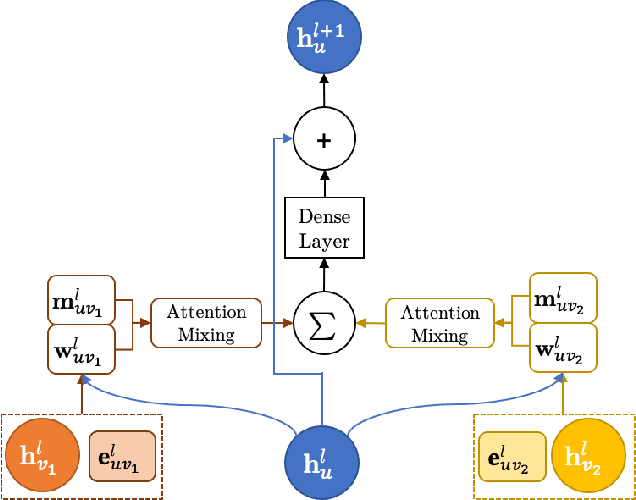
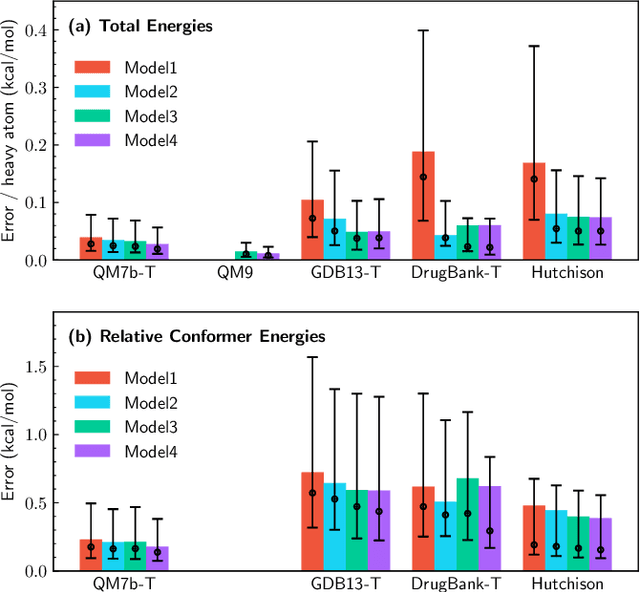
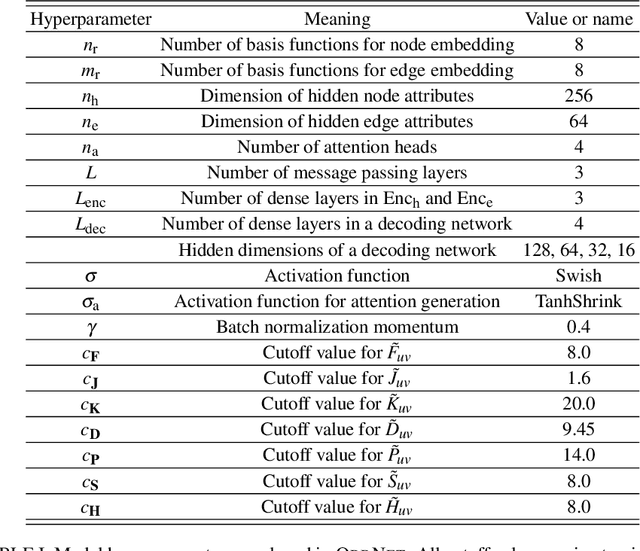
Abstract:We introduce a machine learning method in which energy solutions from the Schrodinger equation are predicted using symmetry adapted atomic orbitals features and a graph neural-network architecture. \textsc{OrbNet} is shown to outperform existing methods in terms of learning efficiency and transferability for the prediction of density functional theory results while employing low-cost features that are obtained from semi-empirical electronic structure calculations. For applications to datasets of drug-like molecules, including QM7b-T, QM9, GDB-13-T, DrugBank, and the conformer benchmark dataset of Folmsbee and Hutchison, \textsc{OrbNet} predicts energies within chemical accuracy of DFT at a computational cost that is thousand-fold or more reduced.
Regression-clustering for Improved Accuracy and Training Cost with Molecular-Orbital-Based Machine Learning
Sep 09, 2019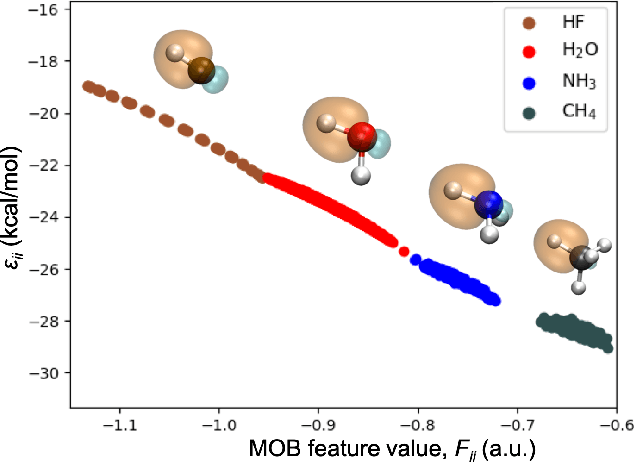
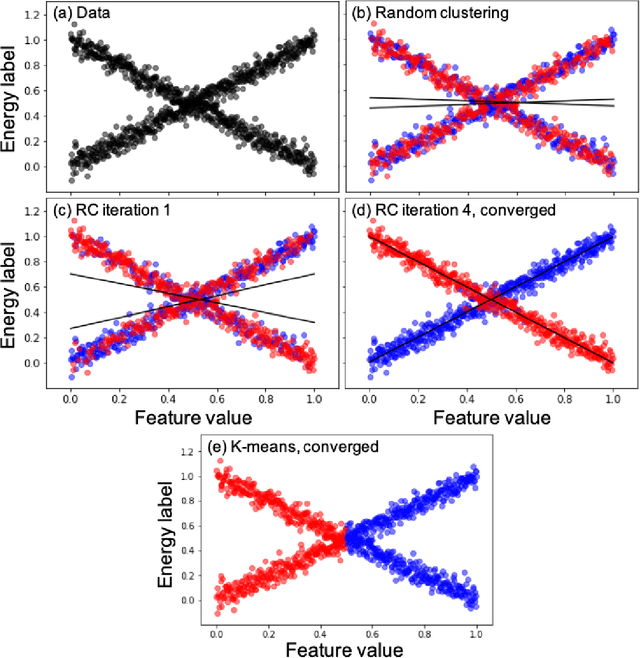
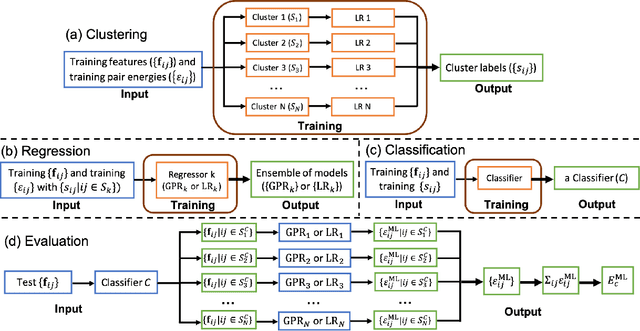
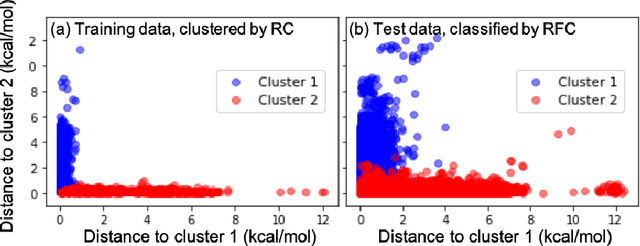
Abstract:Machine learning (ML) in the representation of molecular-orbital-based (MOB) features has been shown to be an accurate and transferable approach to the prediction of post-Hartree-Fock correlation energies. Previous applications of MOB-ML employed Gaussian Process Regression (GPR), which provides good prediction accuracy with small training sets; however, the cost of GPR training scales cubically with the amount of data and becomes a computational bottleneck for large training sets. In the current work, we address this problem by introducing a clustering/regression/classification implementation of MOB-ML. In a first step, regression clustering (RC) is used to partition the training data to best fit an ensemble of linear regression (LR) models; in a second step, each cluster is regressed independently, using either LR or GPR; and in a third step, a random forest classifier (RFC) is trained for the prediction of cluster assignments based on MOB feature values. Upon inspection, RC is found to recapitulate chemically intuitive groupings of the frontier molecular orbitals, and the combined RC/LR/RFC and RC/GPR/RFC implementations of MOB-ML are found to provide good prediction accuracy with greatly reduced wall-clock training times. For a dataset of thermalized geometries of 7211 organic molecules of up to seven heavy atoms, both implementations reach chemical accuracy (1 kcal/mol error) with only 300 training molecules, while providing 35000-fold and 4500-fold reductions in the wall-clock training time, respectively, compared to MOB-ML without clustering. The resulting models are also demonstrated to retain transferability for the prediction of large-molecule energies with only small-molecule training data. Finally, it is shown that capping the number of training datapoints per cluster leads to further improvements in prediction accuracy with negligible increases in wall-clock training time.
A Universal Density Matrix Functional from Molecular Orbital-Based Machine Learning: Transferability across Organic Molecules
Jan 10, 2019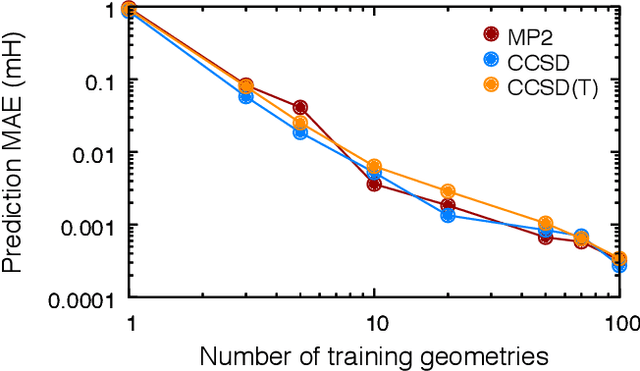
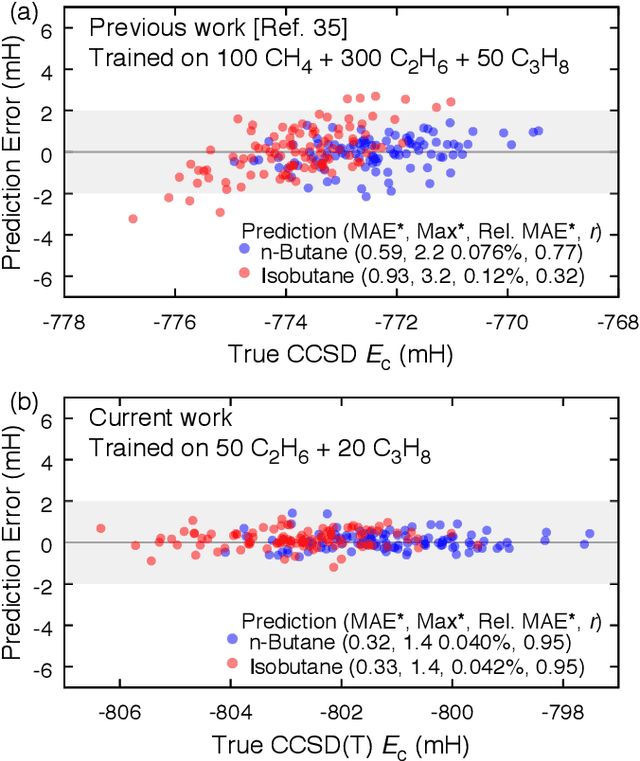
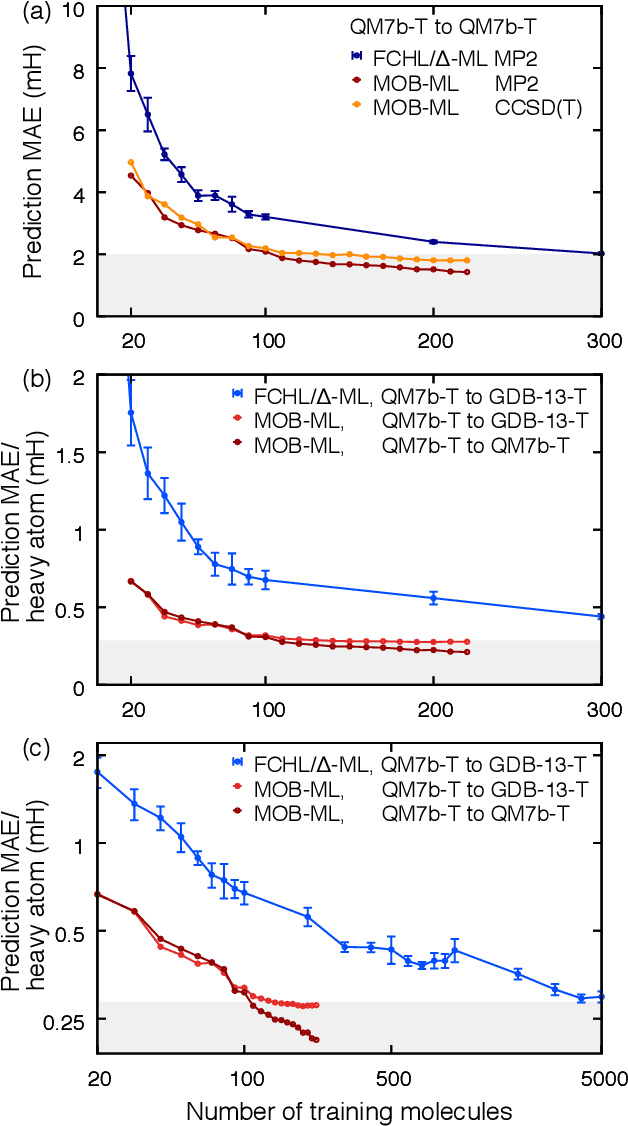
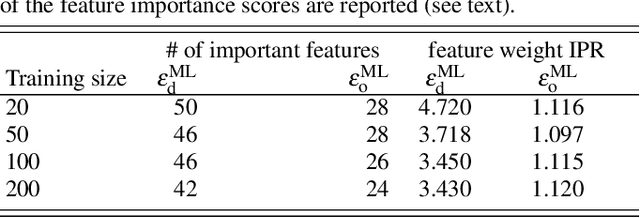
Abstract:We address the degree to which machine learning can be used to accurately and transferably predict post-Hartree-Fock correlation energies. After presenting refined strategies for feature design and selection, the molecular-orbital-based machine learning (MOB-ML) method is first applied to benchmark test systems. It is shown that the total electronic energy for a set of 1000 randomized geometries of water can be described to within 1 millihartree using a model that is trained at the level of MP2, CCSD, or CCSD(T) using only a single reference calculation at a randomized geometry. To explore the breadth of chemical diversity that can be described, the MOB-ML method is then applied a set of 7211 organic models with up to seven heavy atoms. It is shown that MP2 calculations on only 90 molecules are needed to train a model that predicts MP2 energies to within 2 millihartree accuracy for the remaining 7121 molecules; likewise, CCSD(T) calculations on only 150 molecules are needed to train a model that predicts CCSD(T) energies for the remaining molecules to within 2 millihartree accuracy. The MP2 model, trained with only 90 reference calculations on seven-heavy-atom molecules, is then applied to a diverse set of 1000 thirteen-heavy-atom organic molecules, demonstrating transferable preservation of chemical accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge