Kaiwen Huang
Bidirectional Channel-selective Semantic Interaction for Semi-Supervised Medical Segmentation
Jan 09, 2026Abstract:Semi-supervised medical image segmentation is an effective method for addressing scenarios with limited labeled data. Existing methods mainly rely on frameworks such as mean teacher and dual-stream consistency learning. These approaches often face issues like error accumulation and model structural complexity, while also neglecting the interaction between labeled and unlabeled data streams. To overcome these challenges, we propose a Bidirectional Channel-selective Semantic Interaction~(BCSI) framework for semi-supervised medical image segmentation. First, we propose a Semantic-Spatial Perturbation~(SSP) mechanism, which disturbs the data using two strong augmentation operations and leverages unsupervised learning with pseudo-labels from weak augmentations. Additionally, we employ consistency on the predictions from the two strong augmentations to further improve model stability and robustness. Second, to reduce noise during the interaction between labeled and unlabeled data, we propose a Channel-selective Router~(CR) component, which dynamically selects the most relevant channels for information exchange. This mechanism ensures that only highly relevant features are activated, minimizing unnecessary interference. Finally, the Bidirectional Channel-wise Interaction~(BCI) strategy is employed to supplement additional semantic information and enhance the representation of important channels. Experimental results on multiple benchmarking 3D medical datasets demonstrate that the proposed method outperforms existing semi-supervised approaches.
Uncertainty-aware Cross-training for Semi-supervised Medical Image Segmentation
Aug 12, 2025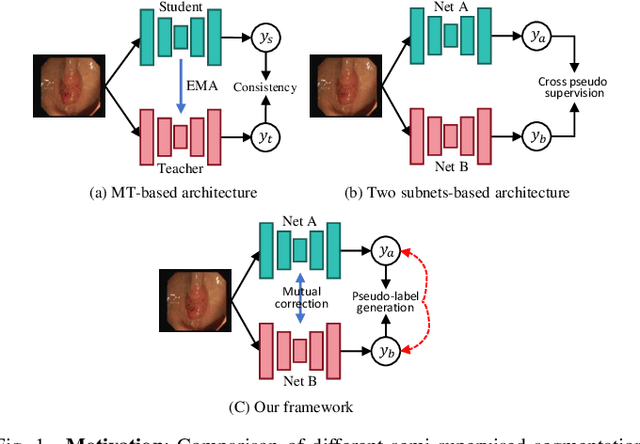
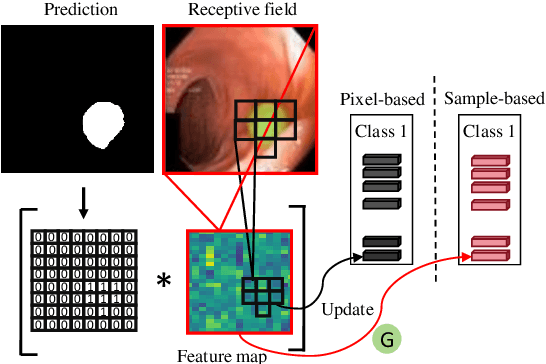
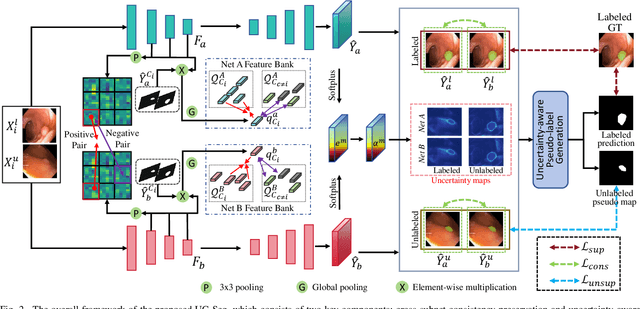
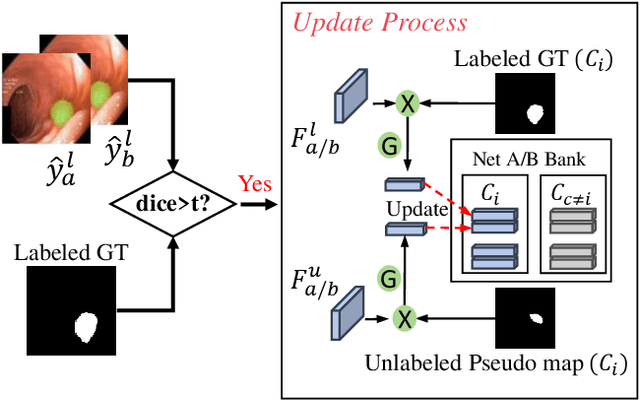
Abstract:Semi-supervised learning has gained considerable popularity in medical image segmentation tasks due to its capability to reduce reliance on expert-examined annotations. Several mean-teacher (MT) based semi-supervised methods utilize consistency regularization to effectively leverage valuable information from unlabeled data. However, these methods often heavily rely on the student model and overlook the potential impact of cognitive biases within the model. Furthermore, some methods employ co-training using pseudo-labels derived from different inputs, yet generating high-confidence pseudo-labels from perturbed inputs during training remains a significant challenge. In this paper, we propose an Uncertainty-aware Cross-training framework for semi-supervised medical image Segmentation (UC-Seg). Our UC-Seg framework incorporates two distinct subnets to effectively explore and leverage the correlation between them, thereby mitigating cognitive biases within the model. Specifically, we present a Cross-subnet Consistency Preservation (CCP) strategy to enhance feature representation capability and ensure feature consistency across the two subnets. This strategy enables each subnet to correct its own biases and learn shared semantics from both labeled and unlabeled data. Additionally, we propose an Uncertainty-aware Pseudo-label Generation (UPG) component that leverages segmentation results and corresponding uncertainty maps from both subnets to generate high-confidence pseudo-labels. We extensively evaluate the proposed UC-Seg on various medical image segmentation tasks involving different modality images, such as MRI, CT, ultrasound, colonoscopy, and so on. The results demonstrate that our method achieves superior segmentation accuracy and generalization performance compared to other state-of-the-art semi-supervised methods. Our code will be released at https://github.com/taozh2017/UCSeg.
Text-driven Multiplanar Visual Interaction for Semi-supervised Medical Image Segmentation
Jul 16, 2025Abstract:Semi-supervised medical image segmentation is a crucial technique for alleviating the high cost of data annotation. When labeled data is limited, textual information can provide additional context to enhance visual semantic understanding. However, research exploring the use of textual data to enhance visual semantic embeddings in 3D medical imaging tasks remains scarce. In this paper, we propose a novel text-driven multiplanar visual interaction framework for semi-supervised medical image segmentation (termed Text-SemiSeg), which consists of three main modules: Text-enhanced Multiplanar Representation (TMR), Category-aware Semantic Alignment (CSA), and Dynamic Cognitive Augmentation (DCA). Specifically, TMR facilitates text-visual interaction through planar mapping, thereby enhancing the category awareness of visual features. CSA performs cross-modal semantic alignment between the text features with introduced learnable variables and the intermediate layer of visual features. DCA reduces the distribution discrepancy between labeled and unlabeled data through their interaction, thus improving the model's robustness. Finally, experiments on three public datasets demonstrate that our model effectively enhances visual features with textual information and outperforms other methods. Our code is available at https://github.com/taozh2017/Text-SemiSeg.
Learnable Prompting SAM-induced Knowledge Distillation for Semi-supervised Medical Image Segmentation
Dec 18, 2024



Abstract:The limited availability of labeled data has driven advancements in semi-supervised learning for medical image segmentation. Modern large-scale models tailored for general segmentation, such as the Segment Anything Model (SAM), have revealed robust generalization capabilities. However, applying these models directly to medical image segmentation still exposes performance degradation. In this paper, we propose a learnable prompting SAM-induced Knowledge distillation framework (KnowSAM) for semi-supervised medical image segmentation. Firstly, we propose a Multi-view Co-training (MC) strategy that employs two distinct sub-networks to employ a co-teaching paradigm, resulting in more robust outcomes. Secondly, we present a Learnable Prompt Strategy (LPS) to dynamically produce dense prompts and integrate an adapter to fine-tune SAM specifically for medical image segmentation tasks. Moreover, we propose SAM-induced Knowledge Distillation (SKD) to transfer useful knowledge from SAM to two sub-networks, enabling them to learn from SAM's predictions and alleviate the effects of incorrect pseudo-labels during training. Notably, the predictions generated by our subnets are used to produce mask prompts for SAM, facilitating effective inter-module information exchange. Extensive experimental results on various medical segmentation tasks demonstrate that our model outperforms the state-of-the-art semi-supervised segmentation approaches. Crucially, our SAM distillation framework can be seamlessly integrated into other semi-supervised segmentation methods to enhance performance. The code will be released upon acceptance of this manuscript at: https://github.com/taozh2017/KnowSAM
SegRap2023: A Benchmark of Organs-at-Risk and Gross Tumor Volume Segmentation for Radiotherapy Planning of Nasopharyngeal Carcinoma
Dec 15, 2023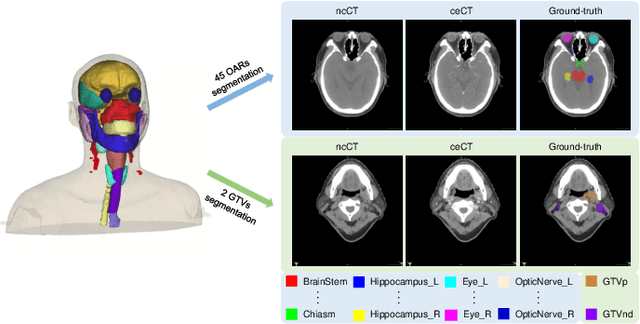
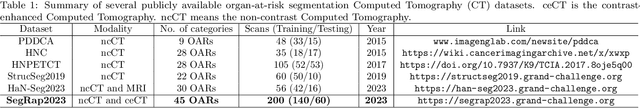
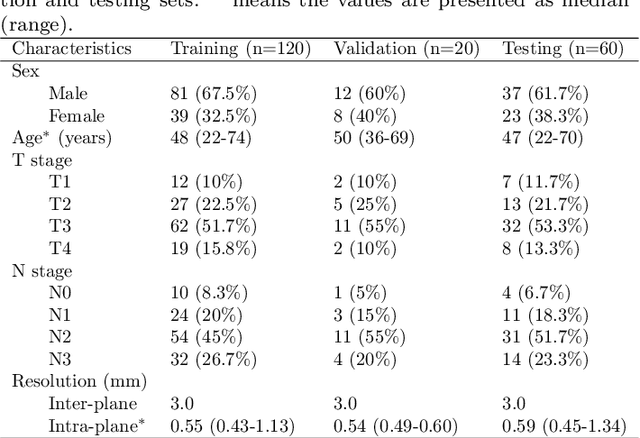
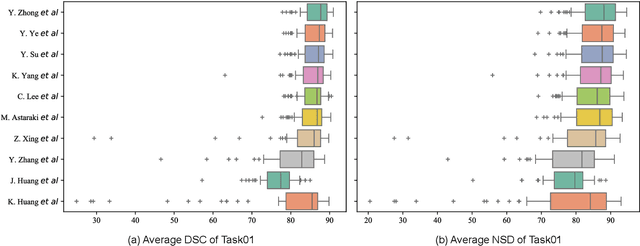
Abstract:Radiation therapy is a primary and effective NasoPharyngeal Carcinoma (NPC) treatment strategy. The precise delineation of Gross Tumor Volumes (GTVs) and Organs-At-Risk (OARs) is crucial in radiation treatment, directly impacting patient prognosis. Previously, the delineation of GTVs and OARs was performed by experienced radiation oncologists. Recently, deep learning has achieved promising results in many medical image segmentation tasks. However, for NPC OARs and GTVs segmentation, few public datasets are available for model development and evaluation. To alleviate this problem, the SegRap2023 challenge was organized in conjunction with MICCAI2023 and presented a large-scale benchmark for OAR and GTV segmentation with 400 Computed Tomography (CT) scans from 200 NPC patients, each with a pair of pre-aligned non-contrast and contrast-enhanced CT scans. The challenge's goal was to segment 45 OARs and 2 GTVs from the paired CT scans. In this paper, we detail the challenge and analyze the solutions of all participants. The average Dice similarity coefficient scores for all submissions ranged from 76.68\% to 86.70\%, and 70.42\% to 73.44\% for OARs and GTVs, respectively. We conclude that the segmentation of large-size OARs is well-addressed, and more efforts are needed for GTVs and small-size or thin-structure OARs. The benchmark will remain publicly available here: https://segrap2023.grand-challenge.org
A Survey on Deep Learning for Polyp Segmentation: Techniques, Challenges and Future Trends
Nov 30, 2023Abstract:Early detection and assessment of polyps play a crucial role in the prevention and treatment of colorectal cancer (CRC). Polyp segmentation provides an effective solution to assist clinicians in accurately locating and segmenting polyp regions. In the past, people often relied on manually extracted lower-level features such as color, texture, and shape, which often had issues capturing global context and lacked robustness to complex scenarios. With the advent of deep learning, more and more outstanding medical image segmentation algorithms based on deep learning networks have emerged, making significant progress in this field. This paper provides a comprehensive review of polyp segmentation algorithms. We first review some traditional algorithms based on manually extracted features and deep segmentation algorithms, then detail benchmark datasets related to the topic. Specifically, we carry out a comprehensive evaluation of recent deep learning models and results based on polyp sizes, considering the pain points of research topics and differences in network structures. Finally, we discuss the challenges of polyp segmentation and future trends in this field. The models, benchmark datasets, and source code links we collected are all published at https://github.com/taozh2017/Awesome-Polyp-Segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge