Jeffrey Rudie
The Brain Tumor Segmentation (BraTS-METS) Challenge 2023: Brain Metastasis Segmentation on Pre-treatment MRI
Jun 01, 2023



Abstract:Clinical monitoring of metastatic disease to the brain can be a laborious and time-consuming process, especially in cases involving multiple metastases when the assessment is performed manually. The Response Assessment in Neuro-Oncology Brain Metastases (RANO-BM) guideline, which utilizes the unidimensional longest diameter, is commonly used in clinical and research settings to evaluate response to therapy in patients with brain metastases. However, accurate volumetric assessment of the lesion and surrounding peri-lesional edema holds significant importance in clinical decision-making and can greatly enhance outcome prediction. The unique challenge in performing segmentations of brain metastases lies in their common occurrence as small lesions. Detection and segmentation of lesions that are smaller than 10 mm in size has not demonstrated high accuracy in prior publications. The brain metastases challenge sets itself apart from previously conducted MICCAI challenges on glioma segmentation due to the significant variability in lesion size. Unlike gliomas, which tend to be larger on presentation scans, brain metastases exhibit a wide range of sizes and tend to include small lesions. We hope that the BraTS-METS dataset and challenge will advance the field of automated brain metastasis detection and segmentation.
Federated Learning Enables Big Data for Rare Cancer Boundary Detection
Apr 25, 2022Abstract:Although machine learning (ML) has shown promise in numerous domains, there are concerns about generalizability to out-of-sample data. This is currently addressed by centrally sharing ample, and importantly diverse, data from multiple sites. However, such centralization is challenging to scale (or even not feasible) due to various limitations. Federated ML (FL) provides an alternative to train accurate and generalizable ML models, by only sharing numerical model updates. Here we present findings from the largest FL study to-date, involving data from 71 healthcare institutions across 6 continents, to generate an automatic tumor boundary detector for the rare disease of glioblastoma, utilizing the largest dataset of such patients ever used in the literature (25,256 MRI scans from 6,314 patients). We demonstrate a 33% improvement over a publicly trained model to delineate the surgically targetable tumor, and 23% improvement over the tumor's entire extent. We anticipate our study to: 1) enable more studies in healthcare informed by large and diverse data, ensuring meaningful results for rare diseases and underrepresented populations, 2) facilitate further quantitative analyses for glioblastoma via performance optimization of our consensus model for eventual public release, and 3) demonstrate the effectiveness of FL at such scale and task complexity as a paradigm shift for multi-site collaborations, alleviating the need for data sharing.
The Brain Tumor Sequence Registration Challenge: Establishing Correspondence between Pre-Operative and Follow-up MRI scans of diffuse glioma patients
Dec 13, 2021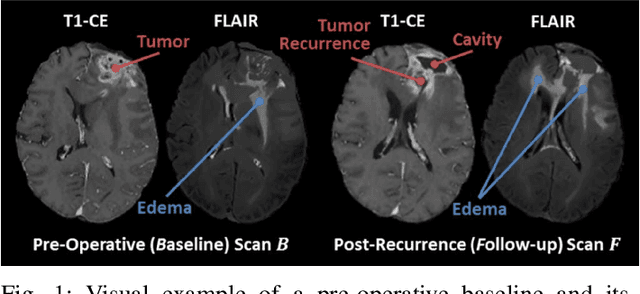
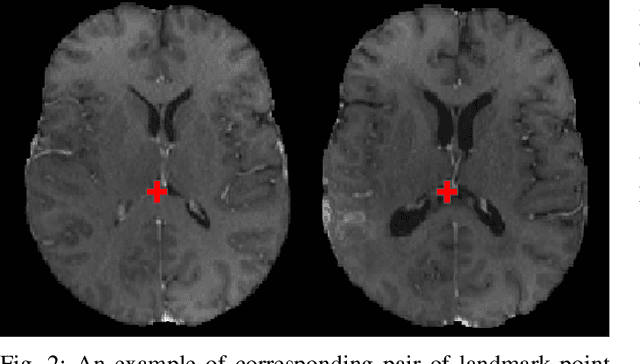
Abstract:Registration of longitudinal brain Magnetic Resonance Imaging (MRI) scans containing pathologies is challenging due to tissue appearance changes, and still an unsolved problem. This paper describes the first Brain Tumor Sequence Registration (BraTS-Reg) challenge, focusing on estimating correspondences between pre-operative and follow-up scans of the same patient diagnosed with a brain diffuse glioma. The BraTS-Reg challenge intends to establish a public benchmark environment for deformable registration algorithms. The associated dataset comprises de-identified multi-institutional multi-parametric MRI (mpMRI) data, curated for each scan's size and resolution, according to a common anatomical template. Clinical experts have generated extensive annotations of landmarks points within the scans, descriptive of distinct anatomical locations across the temporal domain. The training data along with these ground truth annotations will be released to participants to design and develop their registration algorithms, whereas the annotations for the validation and the testing data will be withheld by the organizers and used to evaluate the containerized algorithms of the participants. Each submitted algorithm will be quantitatively evaluated using several metrics, such as the Median Absolute Error (MAE), Robustness, and the Jacobian determinant.
The University of California San Francisco Preoperative Diffuse Glioma MRI Dataset
Aug 30, 2021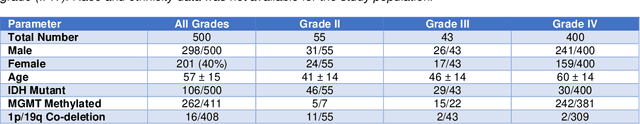
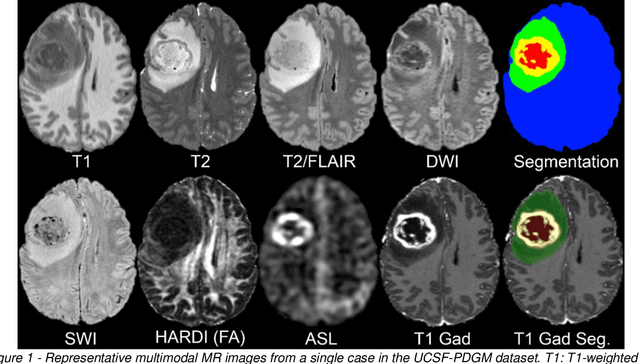
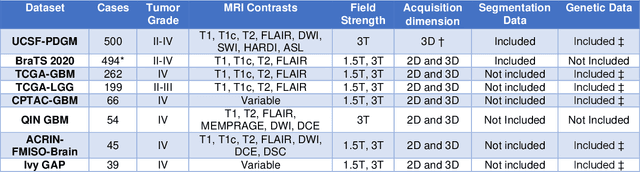
Abstract:Here we present the University of California San Francisco Preoperative Diffuse Glioma MRI (UCSF-PDGM) dataset. The UCSF-PDGM dataset includes 500 subjects with histopathologically-proven diffuse gliomas who were imaged with a standardized 3 Tesla preoperative brain tumor MRI protocol featuring predominantly 3D imaging, as well as advanced diffusion and perfusion imaging techniques. The dataset also includes isocitrate dehydrogenase (IDH) mutation status for all cases and O6-methylguanine-DNA methyltransferase (MGMT) promotor methylation status for World Health Organization (WHO) grade III and IV gliomas. The UCSF-PDGM has been made publicly available in the hopes that researchers around the world will use these data to continue to push the boundaries of AI applications for diffuse gliomas.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge