Julian Schwarting
Framing image registration as a landmark detection problem for better representation of clinical relevance
Jul 31, 2023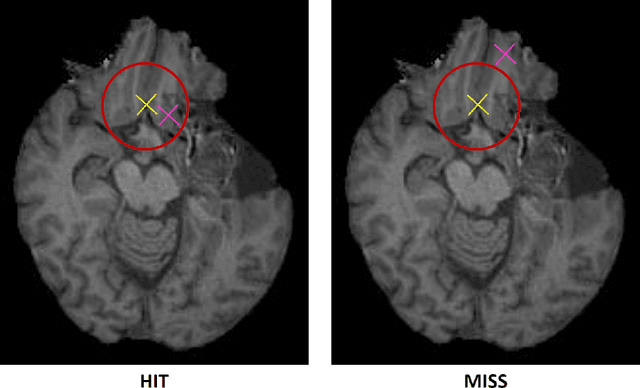
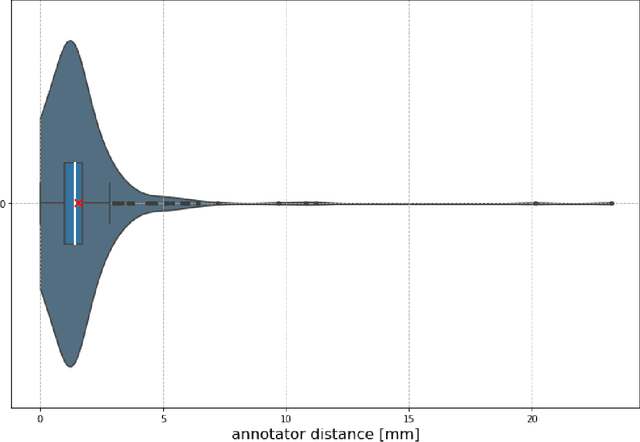
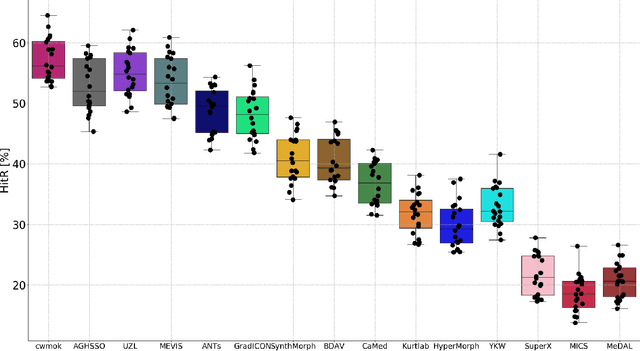
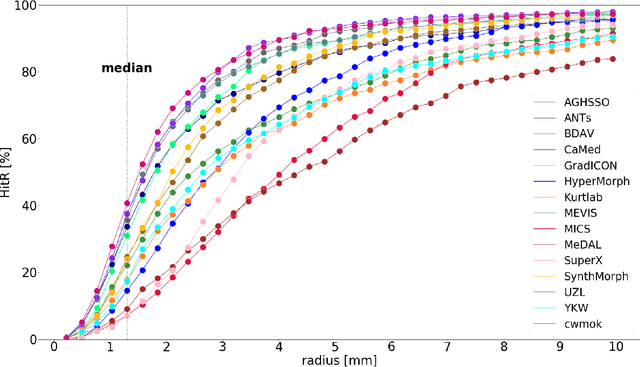
Abstract:Nowadays, registration methods are typically evaluated based on sub-resolution tracking error differences. In an effort to reinfuse this evaluation process with clinical relevance, we propose to reframe image registration as a landmark detection problem. Ideally, landmark-specific detection thresholds are derived from an inter-rater analysis. To approximate this costly process, we propose to compute hit rate curves based on the distribution of errors of a sub-sample inter-rater analysis. Therefore, we suggest deriving thresholds from the error distribution using the formula: median + delta * median absolute deviation. The method promises differentiation of previously indistinguishable registration algorithms and further enables assessing the clinical significance in algorithm development.
Primitive Simultaneous Optimization of Similarity Metrics for Image Registration
Apr 04, 2023



Abstract:Even though simultaneous optimization of similarity metrics represents a standard procedure in the field of semantic segmentation, surprisingly, this does not hold true for image registration. To close this unexpected gap in the literature, we investigate in a complex multi-modal 3D setting whether simultaneous optimization of registration metrics, here implemented by means of primitive summation, can benefit image registration. We evaluate two challenging datasets containing collections of pre- to post-operative and pre- to intra-operative Magnetic Resonance Imaging (MRI) of glioma. Employing the proposed optimization we demonstrate improved registration accuracy in terms of Target Registration Error (TRE) on expert neuroradiologists' landmark annotations.
The Brain Tumor Sequence Registration Challenge: Establishing Correspondence between Pre-Operative and Follow-up MRI scans of diffuse glioma patients
Dec 13, 2021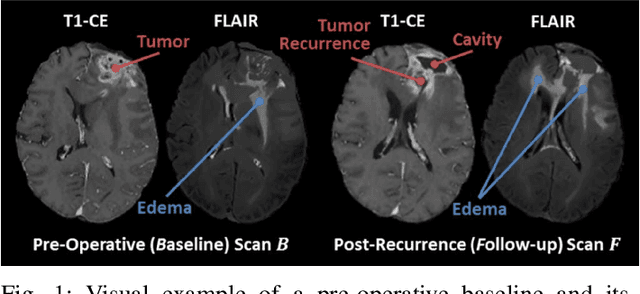
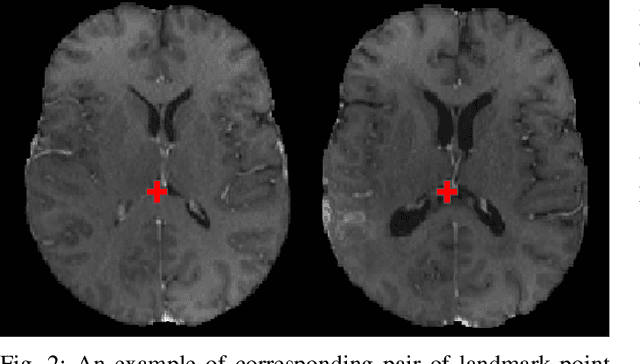
Abstract:Registration of longitudinal brain Magnetic Resonance Imaging (MRI) scans containing pathologies is challenging due to tissue appearance changes, and still an unsolved problem. This paper describes the first Brain Tumor Sequence Registration (BraTS-Reg) challenge, focusing on estimating correspondences between pre-operative and follow-up scans of the same patient diagnosed with a brain diffuse glioma. The BraTS-Reg challenge intends to establish a public benchmark environment for deformable registration algorithms. The associated dataset comprises de-identified multi-institutional multi-parametric MRI (mpMRI) data, curated for each scan's size and resolution, according to a common anatomical template. Clinical experts have generated extensive annotations of landmarks points within the scans, descriptive of distinct anatomical locations across the temporal domain. The training data along with these ground truth annotations will be released to participants to design and develop their registration algorithms, whereas the annotations for the validation and the testing data will be withheld by the organizers and used to evaluate the containerized algorithms of the participants. Each submitted algorithm will be quantitatively evaluated using several metrics, such as the Median Absolute Error (MAE), Robustness, and the Jacobian determinant.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge