Hanley Ong
A foundation model for human-AI collaboration in medical literature mining
Jan 27, 2025



Abstract:Systematic literature review is essential for evidence-based medicine, requiring comprehensive analysis of clinical trial publications. However, the application of artificial intelligence (AI) models for medical literature mining has been limited by insufficient training and evaluation across broad therapeutic areas and diverse tasks. Here, we present LEADS, an AI foundation model for study search, screening, and data extraction from medical literature. The model is trained on 633,759 instruction data points in LEADSInstruct, curated from 21,335 systematic reviews, 453,625 clinical trial publications, and 27,015 clinical trial registries. We showed that LEADS demonstrates consistent improvements over four cutting-edge generic large language models (LLMs) on six tasks. Furthermore, LEADS enhances expert workflows by providing supportive references following expert requests, streamlining processes while maintaining high-quality results. A study with 16 clinicians and medical researchers from 14 different institutions revealed that experts collaborating with LEADS achieved a recall of 0.81 compared to 0.77 experts working alone in study selection, with a time savings of 22.6%. In data extraction tasks, experts using LEADS achieved an accuracy of 0.85 versus 0.80 without using LEADS, alongside a 26.9% time savings. These findings highlight the potential of specialized medical literature foundation models to outperform generic models, delivering significant quality and efficiency benefits when integrated into expert workflows for medical literature mining.
Enhancing disease detection in radiology reports through fine-tuning lightweight LLM on weak labels
Sep 25, 2024Abstract:Despite significant progress in applying large language models (LLMs) to the medical domain, several limitations still prevent them from practical applications. Among these are the constraints on model size and the lack of cohort-specific labeled datasets. In this work, we investigated the potential of improving a lightweight LLM, such as Llama 3.1-8B, through fine-tuning with datasets using synthetic labels. Two tasks are jointly trained by combining their respective instruction datasets. When the quality of the task-specific synthetic labels is relatively high (e.g., generated by GPT4- o), Llama 3.1-8B achieves satisfactory performance on the open-ended disease detection task, with a micro F1 score of 0.91. Conversely, when the quality of the task-relevant synthetic labels is relatively low (e.g., from the MIMIC-CXR dataset), fine-tuned Llama 3.1-8B is able to surpass its noisy teacher labels (micro F1 score of 0.67 v.s. 0.63) when calibrated against curated labels, indicating the strong inherent underlying capability of the model. These findings demonstrate the potential of fine-tuning LLMs with synthetic labels, offering a promising direction for future research on LLM specialization in the medical domain.
Evaluating GPT-4 with Vision on Detection of Radiological Findings on Chest Radiographs
Apr 03, 2024Abstract:The study examines the application of GPT-4V, a multi-modal large language model equipped with visual recognition, in detecting radiological findings from a set of 100 chest radiographs and suggests that GPT-4V is currently not ready for real-world diagnostic usage in interpreting chest radiographs.
Ensembling Low Precision Models for Binary Biomedical Image Segmentation
Oct 16, 2020


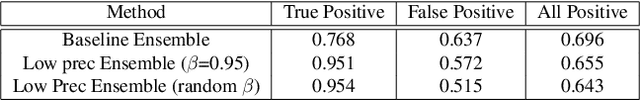
Abstract:Segmentation of anatomical regions of interest such as vessels or small lesions in medical images is still a difficult problem that is often tackled with manual input by an expert. One of the major challenges for this task is that the appearance of foreground (positive) regions can be similar to background (negative) regions. As a result, many automatic segmentation algorithms tend to exhibit asymmetric errors, typically producing more false positives than false negatives. In this paper, we aim to leverage this asymmetry and train a diverse ensemble of models with very high recall, while sacrificing their precision. Our core idea is straightforward: A diverse ensemble of low precision and high recall models are likely to make different false positive errors (classifying background as foreground in different parts of the image), but the true positives will tend to be consistent. Thus, in aggregate the false positive errors will cancel out, yielding high performance for the ensemble. Our strategy is general and can be applied with any segmentation model. In three different applications (carotid artery segmentation in a neck CT angiography, myocardium segmentation in a cardiovascular MRI and multiple sclerosis lesion segmentation in a brain MRI), we show how the proposed approach can significantly boost the performance of a baseline segmentation method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge