George Yiasemis
Towards Universal Learning-based Model for Cardiac Image Reconstruction: Summary of the CMRxRecon2024 Challenge
Mar 05, 2025Abstract:Cardiovascular magnetic resonance (CMR) offers diverse imaging contrasts for assessment of cardiac function and tissue characterization. However, acquiring each single CMR modality is often time-consuming, and comprehensive clinical protocols require multiple modalities with various sampling patterns, further extending the overall acquisition time and increasing susceptibility to motion artifacts. Existing deep learning-based reconstruction methods are often designed for specific acquisition parameters, which limits their ability to generalize across a variety of scan scenarios. As part of the CMRxRecon Series, the CMRxRecon2024 challenge provides diverse datasets encompassing multi-modality multi-view imaging with various sampling patterns, and a platform for the international community to develop and benchmark reconstruction solutions in two well-crafted tasks. Task 1 is a modality-universal setting, evaluating the out-of-distribution generalization of the reconstructed model, while Task 2 follows sampling-universal setting assessing the one-for-all adaptability of the universal model. Main contributions include providing the first and largest publicly available multi-modality, multi-view cardiac k-space dataset; developing a benchmarking platform that simulates clinical acceleration protocols, with a shared code library and tutorial for various k-t undersampling patterns and data processing; giving technical insights of enhanced data consistency based on physic-informed networks and adaptive prompt-learning embedding to be versatile to different clinical settings; additional finding on evaluation metrics to address the limitations of conventional ground-truth references in universal reconstruction tasks.
Deep End-to-end Adaptive k-Space Sampling, Reconstruction, and Registration for Dynamic MRI
Nov 27, 2024Abstract:Dynamic MRI enables a range of clinical applications, including cardiac function assessment, organ motion tracking, and radiotherapy guidance. However, fully sampling the dynamic k-space data is often infeasible due to time constraints and physiological motion such as respiratory and cardiac motion. This necessitates undersampling, which degrades the quality of reconstructed images. Poor image quality not only hinders visualization but also impairs the estimation of deformation fields, crucial for registering dynamic (moving) images to a static reference image. This registration enables tasks such as motion correction, treatment planning, and quantitative analysis in applications like cardiac imaging and MR-guided radiotherapy. To overcome the challenges posed by undersampling and motion, we introduce an end-to-end deep learning (DL) framework that integrates adaptive dynamic k-space sampling, reconstruction, and registration. Our approach begins with a DL-based adaptive sampling strategy, optimizing dynamic k-space acquisition to capture the most relevant data for each specific case. This is followed by a DL-based reconstruction module that produces images optimized for accurate deformation field estimation from the undersampled moving data. Finally, a registration module estimates the deformation fields aligning the reconstructed dynamic images with a static reference. The proposed framework is independent of specific reconstruction and registration modules allowing for plug-and-play integration of these components. The entire framework is jointly trained using a combination of supervised and unsupervised loss functions, enabling end-to-end optimization for improved performance across all components. Through controlled experiments and ablation studies, we validate each component, demonstrating that each choice contributes to robust motion estimation from undersampled dynamic data.
Deep Multi-contrast Cardiac MRI Reconstruction via vSHARP with Auxiliary Refinement Network
Nov 02, 2024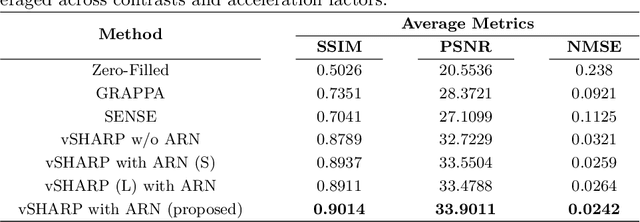

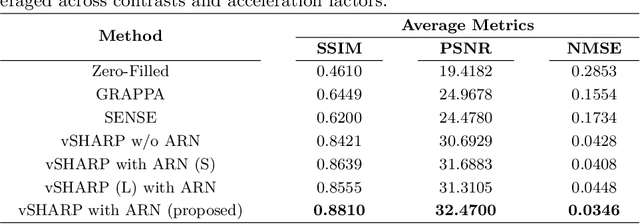

Abstract:Cardiac MRI (CMRI) is a cornerstone imaging modality that provides in-depth insights into cardiac structure and function. Multi-contrast CMRI (MCCMRI), which acquires sequences with varying contrast weightings, significantly enhances diagnostic capabilities by capturing a wide range of cardiac tissue characteristics. However, MCCMRI is often constrained by lengthy acquisition times and susceptibility to motion artifacts. To mitigate these challenges, accelerated imaging techniques that use k-space undersampling via different sampling schemes at acceleration factors have been developed to shorten scan durations. In this context, we propose a deep learning-based reconstruction method for 2D dynamic multi-contrast, multi-scheme, and multi-acceleration MRI. Our approach integrates the state-of-the-art vSHARP model, which utilizes half-quadratic variable splitting and ADMM optimization, with a Variational Network serving as an Auxiliary Refinement Network (ARN) to better adapt to the diverse nature of MCCMRI data. Specifically, the subsampled k-space data is fed into the ARN, which produces an initial prediction for the denoising step used by vSHARP. This, along with the subsampled k-space, is then used by vSHARP to generate high-quality 2D sequence predictions. Our method outperforms traditional reconstruction techniques and other vSHARP-based models.
The state-of-the-art in Cardiac MRI Reconstruction: Results of the CMRxRecon Challenge in MICCAI 2023
Apr 01, 2024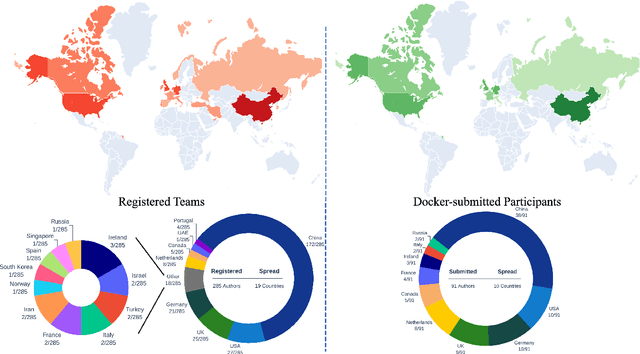
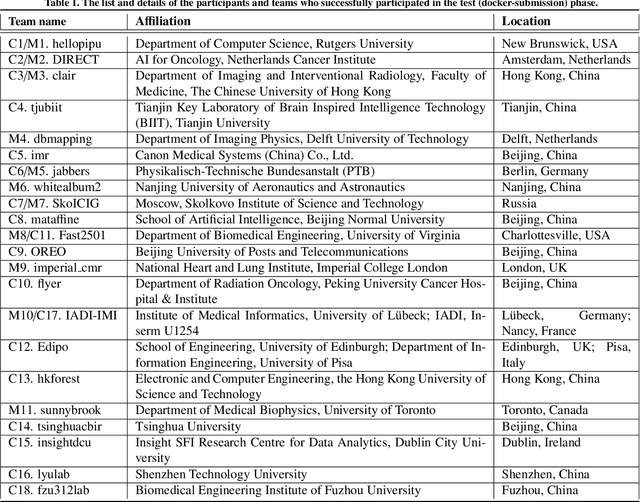
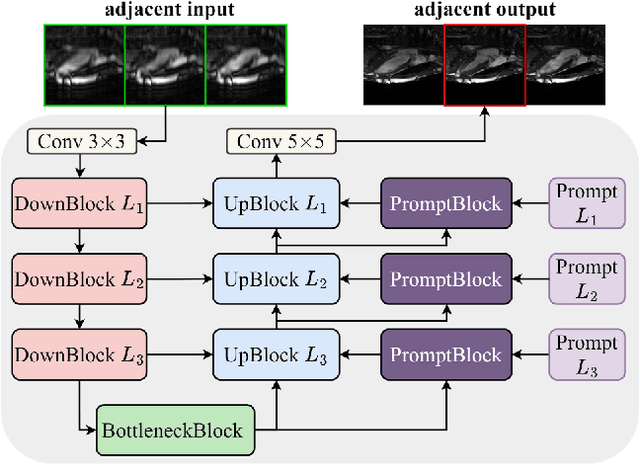
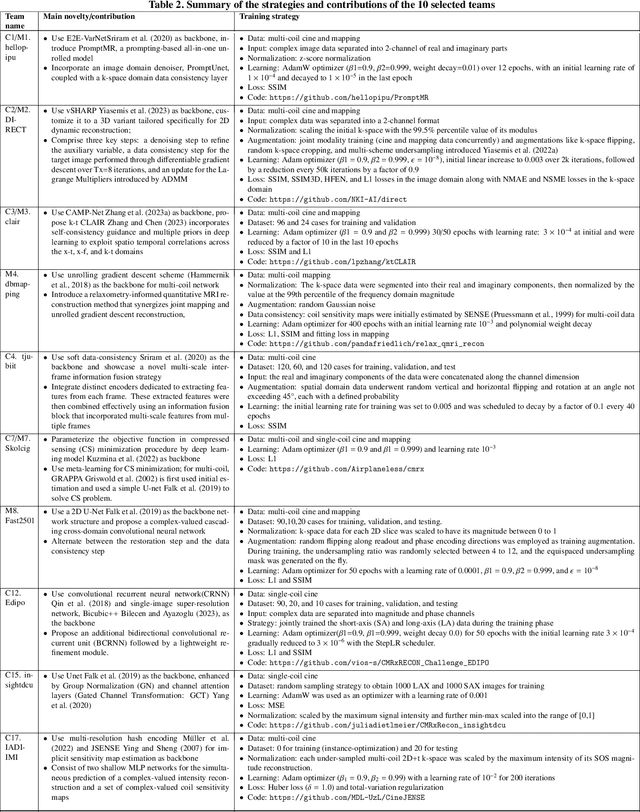
Abstract:Cardiac MRI, crucial for evaluating heart structure and function, faces limitations like slow imaging and motion artifacts. Undersampling reconstruction, especially data-driven algorithms, has emerged as a promising solution to accelerate scans and enhance imaging performance using highly under-sampled data. Nevertheless, the scarcity of publicly available cardiac k-space datasets and evaluation platform hinder the development of data-driven reconstruction algorithms. To address this issue, we organized the Cardiac MRI Reconstruction Challenge (CMRxRecon) in 2023, in collaboration with the 26th International Conference on MICCAI. CMRxRecon presented an extensive k-space dataset comprising cine and mapping raw data, accompanied by detailed annotations of cardiac anatomical structures. With overwhelming participation, the challenge attracted more than 285 teams and over 600 participants. Among them, 22 teams successfully submitted Docker containers for the testing phase, with 7 teams submitted for both cine and mapping tasks. All teams use deep learning based approaches, indicating that deep learning has predominately become a promising solution for the problem. The first-place winner of both tasks utilizes the E2E-VarNet architecture as backbones. In contrast, U-Net is still the most popular backbone for both multi-coil and single-coil reconstructions. This paper provides a comprehensive overview of the challenge design, presents a summary of the submitted results, reviews the employed methods, and offers an in-depth discussion that aims to inspire future advancements in cardiac MRI reconstruction models. The summary emphasizes the effective strategies observed in Cardiac MRI reconstruction, including backbone architecture, loss function, pre-processing techniques, physical modeling, and model complexity, thereby providing valuable insights for further developments in this field.
End-to-end Adaptive Dynamic Subsampling and Reconstruction for Cardiac MRI
Mar 15, 2024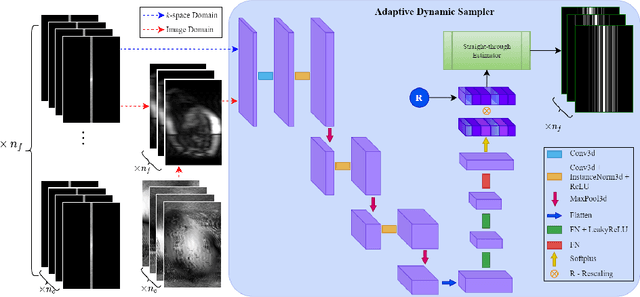
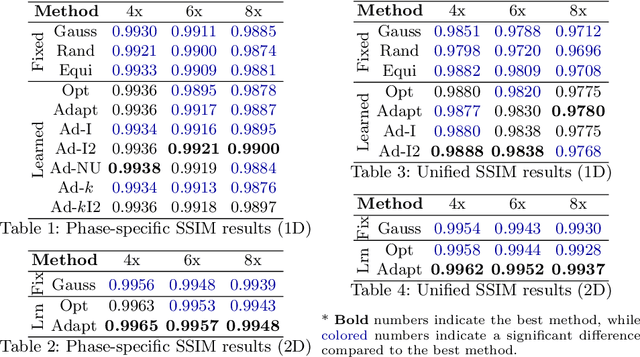
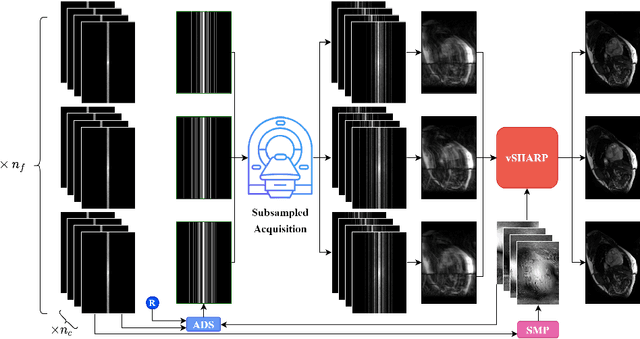
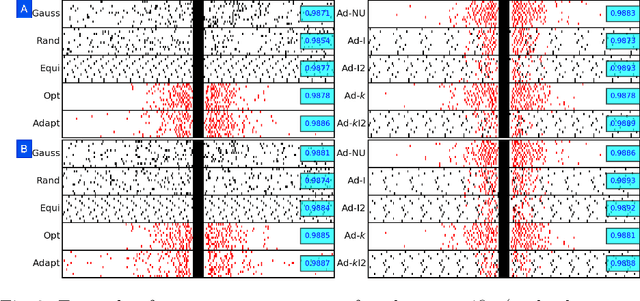
Abstract:Accelerating dynamic MRI is essential for enhancing clinical applications, such as adaptive radiotherapy, and improving patient comfort. Traditional deep learning (DL) approaches for accelerated dynamic MRI reconstruction typically rely on predefined or random subsampling patterns, applied uniformly across all temporal phases. This standard practice overlooks the potential benefits of leveraging temporal correlations and lacks the adaptability required for case-specific subsampling optimization, which holds the potential for maximizing reconstruction quality. Addressing this gap, we present a novel end-to-end framework for adaptive dynamic MRI subsampling and reconstruction. Our pipeline integrates a DL-based adaptive sampler, generating case-specific dynamic subsampling patterns, trained end-to-end with a state-of-the-art 2D dynamic reconstruction network, namely vSHARP, which effectively reconstructs the adaptive dynamic subsampled data into a moving image. Our method is assessed using dynamic cine cardiac MRI data, comparing its performance against vSHARP models that employ common subsampling trajectories, and pipelines trained to optimize dataset-specific sampling schemes alongside vSHARP reconstruction. Our results indicate superior reconstruction quality, particularly at high accelerations.
JSSL: Joint Supervised and Self-supervised Learning for MRI Reconstruction
Nov 27, 2023Abstract:Magnetic Resonance Imaging represents an important diagnostic modality; however, its inherently slow acquisition process poses challenges in obtaining fully sampled k-space data under motion in clinical scenarios such as abdominal, cardiac, and prostate imaging. In the absence of fully sampled acquisitions, which can serve as ground truth data, training deep learning algorithms in a supervised manner to predict the underlying ground truth image becomes an impossible task. To address this limitation, self-supervised methods have emerged as a viable alternative, leveraging available subsampled k-space data to train deep learning networks for MRI reconstruction. Nevertheless, these self-supervised approaches often fall short when compared to supervised methodologies. In this paper, we introduce JSSL (Joint Supervised and Self-supervised Learning), a novel training approach for deep learning-based MRI reconstruction algorithms aimed at enhancing reconstruction quality in scenarios where target dataset(s) containing fully sampled k-space measurements are unavailable. Our proposed method operates by simultaneously training a model in a self-supervised learning setting, using subsampled data from the target dataset(s), and in a supervised learning manner, utilizing data from other datasets, referred to as proxy datasets, where fully sampled k-space data is accessible. To demonstrate the efficacy of JSSL, we utilized subsampled prostate parallel MRI measurements as the target dataset, while employing fully sampled brain and knee k-space acquisitions as proxy datasets. Our results showcase a substantial improvement over conventional self-supervised training methods, thereby underscoring the effectiveness of our joint approach. We provide a theoretical motivation for JSSL and establish a practical "rule-of-thumb" for selecting the most appropriate training approach for deep MRI reconstruction.
Deep Cardiac MRI Reconstruction with ADMM
Oct 10, 2023
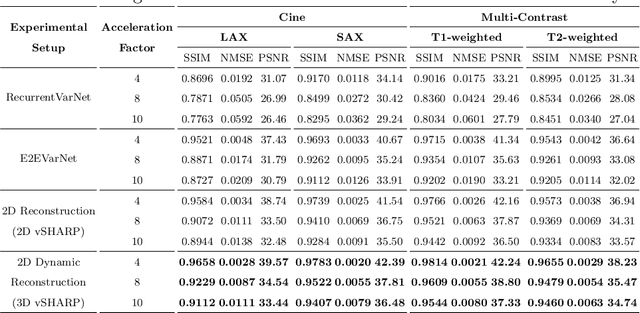


Abstract:Cardiac magnetic resonance imaging is a valuable non-invasive tool for identifying cardiovascular diseases. For instance, Cine MRI is the benchmark modality for assessing the cardiac function and anatomy. On the other hand, multi-contrast (T1 and T2) mapping has the potential to assess pathologies and abnormalities in the myocardium and interstitium. However, voluntary breath-holding and often arrhythmia, in combination with MRI's slow imaging speed, can lead to motion artifacts, hindering real-time acquisition image quality. Although performing accelerated acquisitions can facilitate dynamic imaging, it induces aliasing, causing low reconstructed image quality in Cine MRI and inaccurate T1 and T2 mapping estimation. In this work, inspired by related work in accelerated MRI reconstruction, we present a deep learning (DL)-based method for accelerated cine and multi-contrast reconstruction in the context of dynamic cardiac imaging. We formulate the reconstruction problem as a least squares regularized optimization task, and employ vSHARP, a state-of-the-art DL-based inverse problem solver, which incorporates half-quadratic variable splitting and the alternating direction method of multipliers with neural networks. We treat the problem in two setups; a 2D reconstruction and a 2D dynamic reconstruction task, and employ 2D and 3D deep learning networks, respectively. Our method optimizes in both the image and k-space domains, allowing for high reconstruction fidelity. Although the target data is undersampled with a Cartesian equispaced scheme, we train our model using both Cartesian and simulated non-Cartesian undersampling schemes to enhance generalization of the model to unseen data. Furthermore, our model adopts a deep neural network to learn and refine the sensitivity maps of multi-coil k-space data. Lastly, our method is jointly trained on both, undersampled cine and multi-contrast data.
vSHARP: variable Splitting Half-quadratic ADMM algorithm for Reconstruction of inverse-Problems
Sep 18, 2023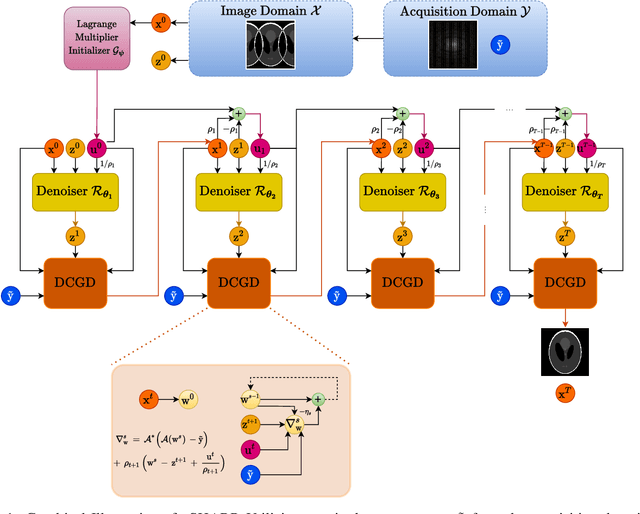
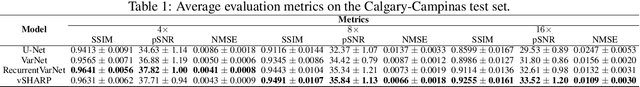
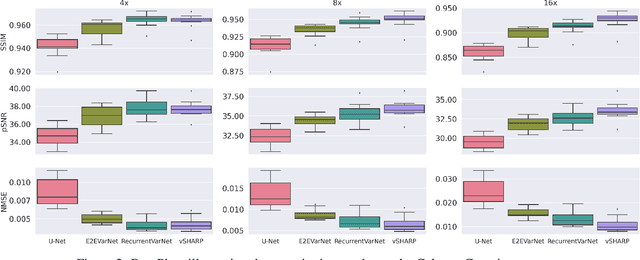

Abstract:Medical Imaging (MI) tasks, such as accelerated Parallel Magnetic Resonance Imaging (MRI), often involve reconstructing an image from noisy or incomplete measurements. This amounts to solving ill-posed inverse problems, where a satisfactory closed-form analytical solution is not available. Traditional methods such as Compressed Sensing (CS) in MRI reconstruction can be time-consuming or prone to obtaining low-fidelity images. Recently, a plethora of supervised and self-supervised Deep Learning (DL) approaches have demonstrated superior performance in inverse-problem solving, surpassing conventional methods. In this study, we propose vSHARP (variable Splitting Half-quadratic ADMM algorithm for Reconstruction of inverse Problems), a novel DL-based method for solving ill-posed inverse problems arising in MI. vSHARP utilizes the Half-Quadratic Variable Splitting method and employs the Alternating Direction Method of Multipliers (ADMM) to unroll the optimization process. For data consistency, vSHARP unrolls a differentiable gradient descent process in the image domain, while a DL-based denoiser, such as a U-Net architecture, is applied to enhance image quality. vSHARP also employs a dilated-convolution DL-based model to predict the Lagrange multipliers for the ADMM initialization. We evaluate the proposed model by applying it to the task of accelerated Parallel MRI Reconstruction on two distinct datasets. We present a comparative analysis of our experimental results with state-of-the-art approaches, highlighting the superior performance of vSHARP.
On Retrospective k-space Subsampling schemes For Deep MRI Reconstruction
Jan 23, 2023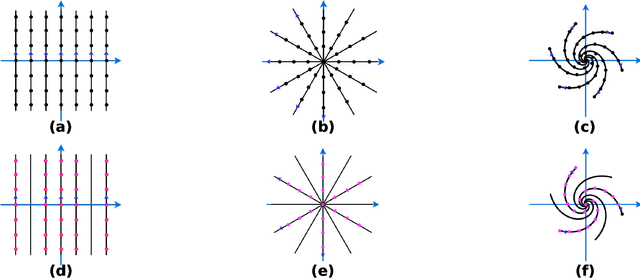
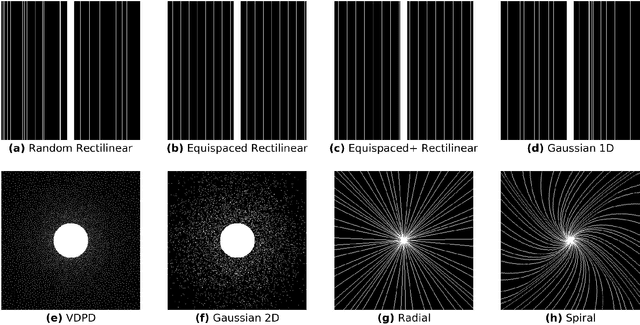
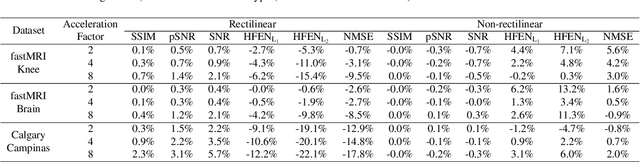

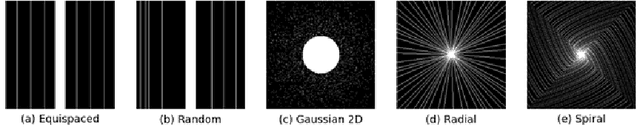
Abstract:$\textbf{Purpose:}$ The MRI $k$-space acquisition is time consuming. Traditional techniques aim to acquire accelerated data, which in conjunction with recent DL methods, aid in producing high-fidelity images in truncated times. Conventionally, subsampling the $k$-space is performed by utilizing Cartesian-rectilinear trajectories, which even with the use of DL, provide imprecise reconstructions, though, a plethora of non-rectilinear or non-Cartesian trajectories can be implemented in modern MRI scanners. This work investigates the effect of the $k$-space subsampling scheme on the quality of reconstructed accelerated MRI measurements produced by trained DL models. $\textbf{Methods:}$ The RecurrentVarNet was used as the DL-based MRI-reconstruction architecture. Cartesian fully-sampled multi-coil $k$-space measurements from three datasets with different accelerations were retrospectively subsampled using eight distinct subsampling schemes (four Cartesian-rectilinear, two Cartesian non-rectilinear, two non-Cartesian). Experiments were conducted in two frameworks: Scheme-specific, where a distinct model was trained and evaluated for each dataset-subsampling scheme pair, and multi-scheme, where for each dataset a single model was trained on data randomly subsampled by any of the eight schemes and evaluated on data subsampled by all schemes. $\textbf{Results:}$ In the scheme-specific setting RecurrentVarNets trained and evaluated on non-rectilinearly subsampled data demonstrated superior performance especially for high accelerations, whilst in the multi-scheme setting, reconstruction performance on rectilinearly subsampled data improved when compared to the scheme-specific experiments. $\textbf{Conclusion:}$ Training DL-based MRI reconstruction algorithms on non-rectilinearly subsampled measurements can produce more faithful reconstructions.
Recurrent Variational Network: A Deep Learning Inverse Problem Solver applied to the task of Accelerated MRI Reconstruction
Nov 18, 2021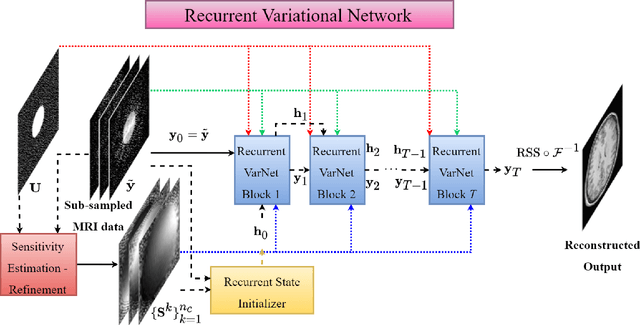
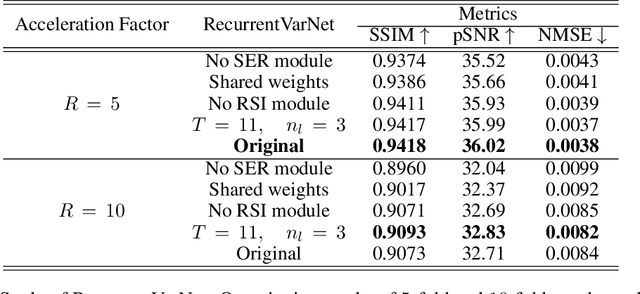
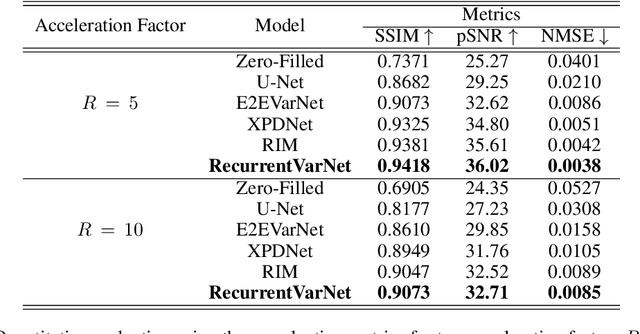
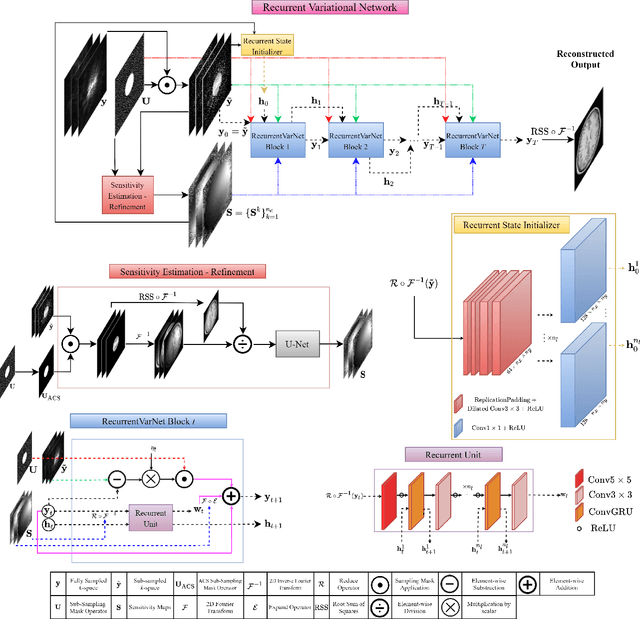

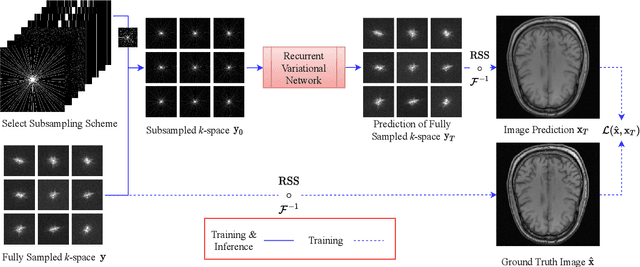
Abstract:Magnetic Resonance Imaging can produce detailed images of the anatomy and physiology of the human body that can assist doctors in diagnosing and treating pathologies such as tumours. However, MRI suffers from very long acquisition times that make it susceptible to patient motion artifacts and limit its potential to deliver dynamic treatments. Conventional approaches such as Parallel Imaging and Compressed Sensing allow for an increase in MRI acquisition speed by reconstructing MR images by acquiring less MRI data using multiple receiver coils. Recent advancements in Deep Learning combined with Parallel Imaging and Compressed Sensing techniques have the potential to produce high-fidelity reconstructions from highly accelerated MRI data. In this work we present a novel Deep Learning-based Inverse Problem solver applied to the task of accelerated MRI reconstruction, called Recurrent Variational Network (RecurrentVarNet) by exploiting the properties of Convolution Recurrent Networks and unrolled algorithms for solving Inverse Problems. The RecurrentVarNet consists of multiple blocks, each responsible for one unrolled iteration of the gradient descent optimization algorithm for solving inverse problems. Contrary to traditional approaches, the optimization steps are performed in the observation domain ($k$-space) instead of the image domain. Each recurrent block of RecurrentVarNet refines the observed $k$-space and is comprised of a data consistency term and a recurrent unit which takes as input a learned hidden state and the prediction of the previous block. Our proposed method achieves new state of the art qualitative and quantitative reconstruction results on 5-fold and 10-fold accelerated data from a public multi-channel brain dataset, outperforming previous conventional and deep learning-based approaches. We will release all models code and baselines on our public repository.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge