Fabian Bongratz
Cortex-Grounded Diffusion Models for Brain Image Generation
Jan 27, 2026Abstract:Synthetic neuroimaging data can mitigate critical limitations of real-world datasets, including the scarcity of rare phenotypes, domain shifts across scanners, and insufficient longitudinal coverage. However, existing generative models largely rely on weak conditioning signals, such as labels or text, which lack anatomical grounding and often produce biologically implausible outputs. To this end, we introduce Cor2Vox, a cortex-grounded generative framework for brain magnetic resonance image (MRI) synthesis that ties image generation to continuous structural priors of the cerebral cortex. It leverages high-resolution cortical surfaces to guide a 3D shape-to-image Brownian bridge diffusion process, enabling topologically faithful synthesis and precise control over underlying anatomies. To support the generation of new, realistic brain shapes, we developed a large-scale statistical shape model of cortical morphology derived from over 33,000 UK Biobank scans. We validated the fidelity of Cor2Vox based on traditional image quality metrics, advanced cortical surface reconstruction, and whole-brain segmentation quality, outperforming many baseline methods. Across three applications, namely (i) anatomically consistent synthesis, (ii) simulation of progressive gray matter atrophy, and (iii) harmonization of in-house frontotemporal dementia scans with public datasets, Cor2Vox preserved fine-grained cortical morphology at the sub-voxel level, exhibiting remarkable robustness to variations in cortical geometry and disease phenotype without retraining.
Template-Based Cortical Surface Reconstruction with Minimal Energy Deformation
Sep 18, 2025Abstract:Cortical surface reconstruction (CSR) from magnetic resonance imaging (MRI) is fundamental to neuroimage analysis, enabling morphological studies of the cerebral cortex and functional brain mapping. Recent advances in learning-based CSR have dramatically accelerated processing, allowing for reconstructions through the deformation of anatomical templates within seconds. However, ensuring the learned deformations are optimal in terms of deformation energy and consistent across training runs remains a particular challenge. In this work, we design a Minimal Energy Deformation (MED) loss, acting as a regularizer on the deformation trajectories and complementing the widely used Chamfer distance in CSR. We incorporate it into the recent V2C-Flow model and demonstrate considerable improvements in previously neglected training consistency and reproducibility without harming reconstruction accuracy and topological correctness.
Spherical Brownian Bridge Diffusion Models for Conditional Cortical Thickness Forecasting
Sep 10, 2025Abstract:Accurate forecasting of individualized, high-resolution cortical thickness (CTh) trajectories is essential for detecting subtle cortical changes, providing invaluable insights into neurodegenerative processes and facilitating earlier and more precise intervention strategies. However, CTh forecasting is a challenging task due to the intricate non-Euclidean geometry of the cerebral cortex and the need to integrate multi-modal data for subject-specific predictions. To address these challenges, we introduce the Spherical Brownian Bridge Diffusion Model (SBDM). Specifically, we propose a bidirectional conditional Brownian bridge diffusion process to forecast CTh trajectories at the vertex level of registered cortical surfaces. Our technical contribution includes a new denoising model, the conditional spherical U-Net (CoS-UNet), which combines spherical convolutions and dense cross-attention to integrate cortical surfaces and tabular conditions seamlessly. Compared to previous approaches, SBDM achieves significantly reduced prediction errors, as demonstrated by our experiments based on longitudinal datasets from the ADNI and OASIS. Additionally, we demonstrate SBDM's ability to generate individual factual and counterfactual CTh trajectories, offering a novel framework for exploring hypothetical scenarios of cortical development.
3D Shape-to-Image Brownian Bridge Diffusion for Brain MRI Synthesis from Cortical Surfaces
Feb 18, 2025



Abstract:Despite recent advances in medical image generation, existing methods struggle to produce anatomically plausible 3D structures. In synthetic brain magnetic resonance images (MRIs), characteristic fissures are often missing, and reconstructed cortical surfaces appear scattered rather than densely convoluted. To address this issue, we introduce Cor2Vox, the first diffusion model-based method that translates continuous cortical shape priors to synthetic brain MRIs. To achieve this, we leverage a Brownian bridge process which allows for direct structured mapping between shape contours and medical images. Specifically, we adapt the concept of the Brownian bridge diffusion model to 3D and extend it to embrace various complementary shape representations. Our experiments demonstrate significant improvements in the geometric accuracy of reconstructed structures compared to previous voxel-based approaches. Moreover, Cor2Vox excels in image quality and diversity, yielding high variation in non-target structures like the skull. Finally, we highlight the capability of our approach to simulate cortical atrophy at the sub-voxel level. Our code is available at https://github.com/ai-med/Cor2Vox.
TimeFlow: Longitudinal Brain Image Registration and Aging Progression Analysis
Jan 15, 2025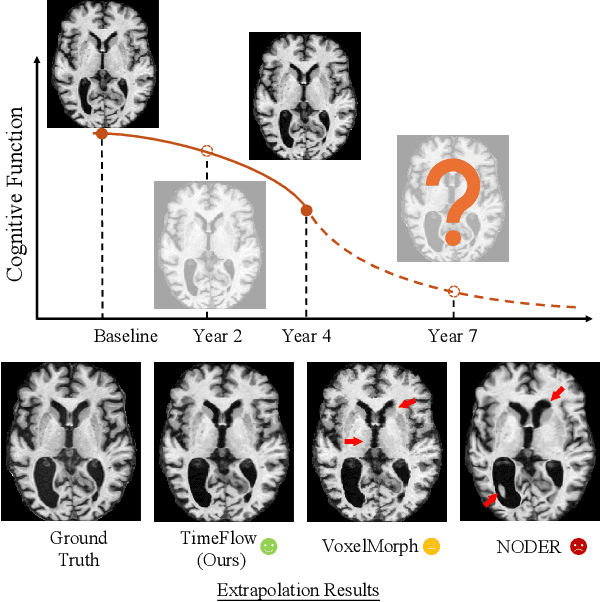
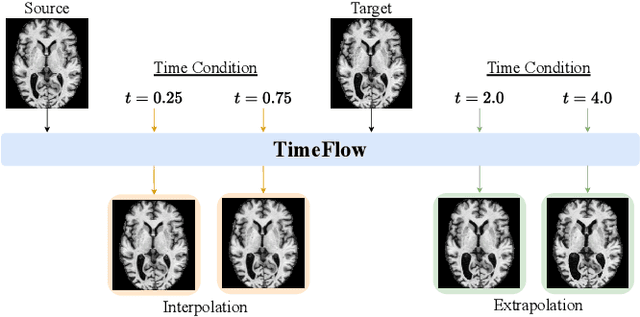
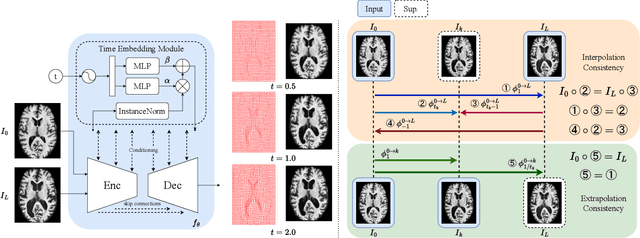
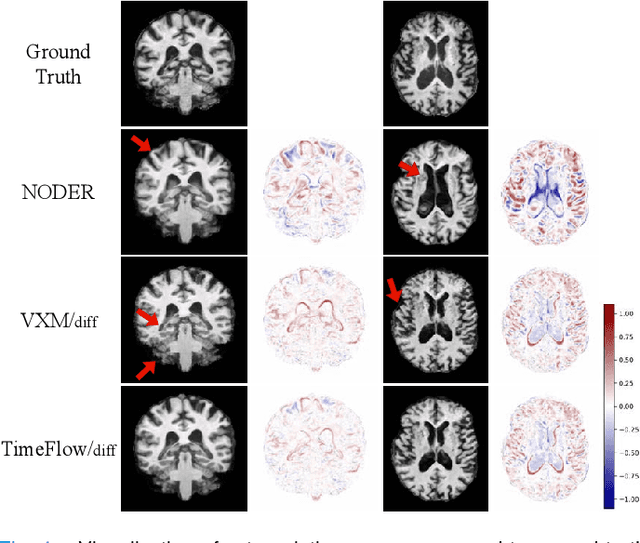
Abstract:Predicting future brain states is crucial for understanding healthy aging and neurodegenerative diseases. Longitudinal brain MRI registration, a cornerstone for such analyses, has long been limited by its inability to forecast future developments, reliance on extensive, dense longitudinal data, and the need to balance registration accuracy with temporal smoothness. In this work, we present \emph{TimeFlow}, a novel framework for longitudinal brain MRI registration that overcomes all these challenges. Leveraging a U-Net architecture with temporal conditioning inspired by diffusion models, TimeFlow enables accurate longitudinal registration and facilitates prospective analyses through future image prediction. Unlike traditional methods that depend on explicit smoothness regularizers and dense sequential data, TimeFlow achieves temporal consistency and continuity without these constraints. Experimental results highlight its superior performance in both future timepoint prediction and registration accuracy compared to state-of-the-art methods. Additionally, TimeFlow supports novel biological brain aging analyses, effectively differentiating neurodegenerative conditions from healthy aging. It eliminates the need for segmentation, thereby avoiding the challenges of non-trivial annotation and inconsistent segmentation errors. TimeFlow paves the way for accurate, data-efficient, and annotation-free prospective analyses of brain aging and chronic diseases.
MLV$^2$-Net: Rater-Based Majority-Label Voting for Consistent Meningeal Lymphatic Vessel Segmentation
Nov 13, 2024



Abstract:Meningeal lymphatic vessels (MLVs) are responsible for the drainage of waste products from the human brain. An impairment in their functionality has been associated with aging as well as brain disorders like multiple sclerosis and Alzheimer's disease. However, MLVs have only recently been described for the first time in magnetic resonance imaging (MRI), and their ramified structure renders manual segmentation particularly difficult. Further, as there is no consistent notion of their appearance, human-annotated MLV structures contain a high inter-rater variability that most automatic segmentation methods cannot take into account. In this work, we propose a new rater-aware training scheme for the popular nnU-Net model, and we explore rater-based ensembling strategies for accurate and consistent segmentation of MLVs. This enables us to boost nnU-Net's performance while obtaining explicit predictions in different annotation styles and a rater-based uncertainty estimation. Our final model, MLV$^2$-Net, achieves a Dice similarity coefficient of 0.806 with respect to the human reference standard. The model further matches the human inter-rater reliability and replicates age-related associations with MLV volume.
How to Choose a Reinforcement-Learning Algorithm
Jul 30, 2024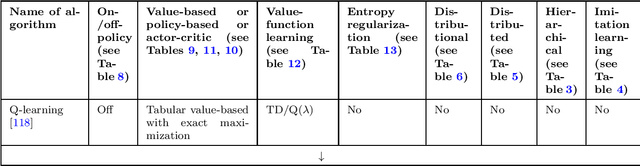
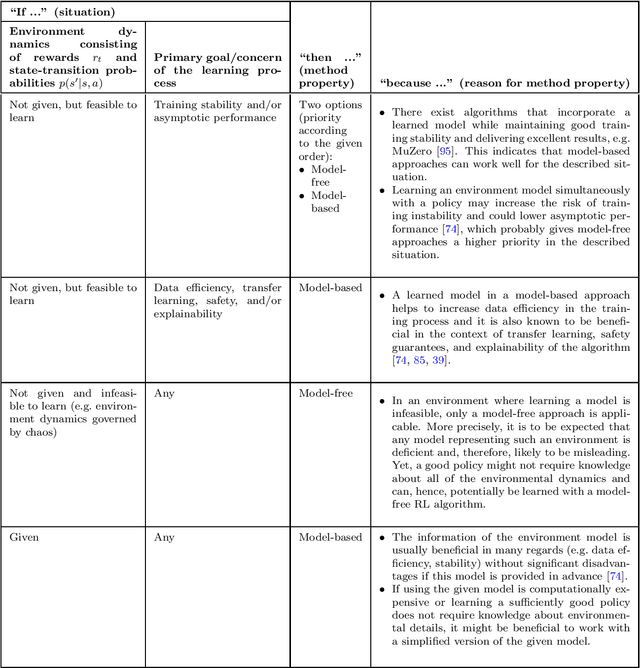
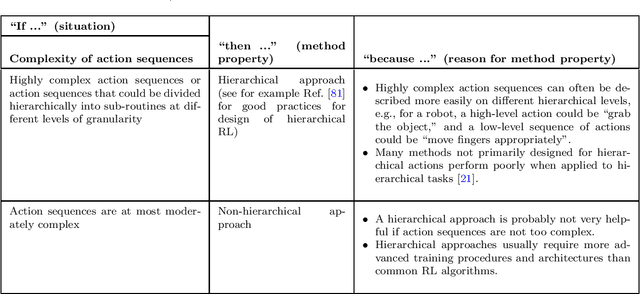

Abstract:The field of reinforcement learning offers a large variety of concepts and methods to tackle sequential decision-making problems. This variety has become so large that choosing an algorithm for a task at hand can be challenging. In this work, we streamline the process of choosing reinforcement-learning algorithms and action-distribution families. We provide a structured overview of existing methods and their properties, as well as guidelines for when to choose which methods. An interactive version of these guidelines is available online at https://rl-picker.github.io/.
Stochastic Cortical Self-Reconstruction
Mar 11, 2024Abstract:Magnetic resonance imaging (MRI) is critical for diagnosing neurodegenerative diseases, yet accurately assessing mild cortical atrophy remains a challenge due to its subtlety. Automated cortex reconstruction, paired with healthy reference ranges, aids in pinpointing pathological atrophy, yet their generalization is limited by biases from image acquisition and processing. We introduce the concept of stochastic cortical self-reconstruction (SCSR) that creates a subject-specific healthy reference by taking MRI-derived thicknesses as input and, therefore, implicitly accounting for potential confounders. SCSR randomly corrupts parts of the cortex and self-reconstructs them from the remaining information. Trained exclusively on healthy individuals, repeated self-reconstruction generates a stochastic reference cortex for assessing deviations from the norm. We present three implementations of this concept: XGBoost applied on parcels, and two autoencoders on vertex level -- one based on a multilayer perceptron and the other using a spherical U-Net. These models were trained on healthy subjects from the UK Biobank and subsequently evaluated across four public Alzheimer's datasets. Finally, we deploy the model on clinical in-house data, where deviation maps' high spatial resolution aids in discriminating between four types of dementia.
V2C-Long: Longitudinal Cortex Reconstruction with Spatiotemporal Correspondence
Feb 27, 2024



Abstract:Reconstructing the cortex from longitudinal MRI is indispensable for analyzing morphological changes in the human brain. Despite the recent disruption of cortical surface reconstruction with deep learning, challenges arising from longitudinal data are still persistent. Especially the lack of strong spatiotemporal point correspondence hinders downstream analyses due to the introduced noise. To address this issue, we present V2C-Long, the first dedicated deep learning-based cortex reconstruction method for longitudinal MRI. In contrast to existing methods, V2C-Long surfaces are directly comparable in a cross-sectional and longitudinal manner. We establish strong inherent spatiotemporal correspondences via a novel composition of two deep mesh deformation networks and fast aggregation of feature-enhanced within-subject templates. The results on internal and external test data demonstrate that V2C-Long yields cortical surfaces with improved accuracy and consistency compared to previous methods. Finally, this improvement manifests in higher sensitivity to regional cortical atrophy in Alzheimer's disease.
Neural deformation fields for template-based reconstruction of cortical surfaces from MRI
Jan 23, 2024



Abstract:The reconstruction of cortical surfaces is a prerequisite for quantitative analyses of the cerebral cortex in magnetic resonance imaging (MRI). Existing segmentation-based methods separate the surface registration from the surface extraction, which is computationally inefficient and prone to distortions. We introduce Vox2Cortex-Flow (V2C-Flow), a deep mesh-deformation technique that learns a deformation field from a brain template to the cortical surfaces of an MRI scan. To this end, we present a geometric neural network that models the deformation-describing ordinary differential equation in a continuous manner. The network architecture comprises convolutional and graph-convolutional layers, which allows it to work with images and meshes at the same time. V2C-Flow is not only very fast, requiring less than two seconds to infer all four cortical surfaces, but also establishes vertex-wise correspondences to the template during reconstruction. In addition, V2C-Flow is the first approach for cortex reconstruction that models white matter and pial surfaces jointly, therefore avoiding intersections between them. Our comprehensive experiments on internal and external test data demonstrate that V2C-Flow results in cortical surfaces that are state-of-the-art in terms of accuracy. Moreover, we show that the established correspondences are more consistent than in FreeSurfer and that they can directly be utilized for cortex parcellation and group analyses of cortical thickness.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge