Ellen Kuhl
Generative AI for material design: A mechanics perspective from burgers to matter
Apr 03, 2026Abstract:Generative artificial intelligence offers a new paradigm to design matter in high-dimensional spaces. However, its underlying mechanisms remain difficult to interpret and limit adoption in computational mechanics. This gap is striking because its core tools-diffusion, stochastic differential equations, and inverse problems-are fundamental to the mechanics of materials. Here we show that diffusion-based generative AI and computational mechanics are rooted in the same principles. We illustrate this connection using a three-ingredient burger as a minimal benchmark for material design in a low-dimensional space, where both forward and reverse diffusion admit analytical solutions: Markov chains with Bayesian inversion in the discrete case and the Ornstein-Uhlenbeck process with score-based reversal in the continuous case. We extend this framework to a high-dimensional design space with 146 ingredients and 8.9x10^43 possible configurations, where analytical solutions become intractable. We therefore learn the discrete and continuous reverse processes using neural network models that infer inverse dynamics from data. We train the models on only 2,260 recipes and generate one million samples that capture the statistical structure of the data, including ingredient prevalence and quantitative composition. We further generate five new burgers and validate them in a restaurant-based sensory study with 100 participants, where three of the AI-designed burgers outperform the classical Big Mac in overall liking, flavor, and texture. These results establish diffusion-based generative modeling as a physically grounded approach to design in high-dimensional spaces. They position generative AI as a natural extension of computational mechanics, with applications from burgers to matter, and establish a path toward data-driven, physics-informed generative design.
A Convex Route to Thermomechanics: Learning Internal Energy and Dissipation
Mar 31, 2026Abstract:We present a physics-based neural network framework for the discovery of constitutive models in fully coupled thermomechanics. In contrast to classical formulations based on the Helmholtz energy, we adopt the internal energy and a dissipation potential as primary constitutive functions, expressed in terms of deformation and entropy. This choice avoids the need to enforce mixed convexity--concavity conditions and facilitates a consistent incorporation of thermodynamic principles. In this contribution, we focus on materials without preferred directions or internal variables. While the formulation is posed in terms of entropy, the temperature is treated as the independent observable, and the entropy is inferred internally through the constitutive relation, enabling thermodynamically consistent modeling without requiring entropy data. Thermodynamic admissibility of the networks is guaranteed by construction. The internal energy and dissipation potential are represented by input convex neural networks, ensuring convexity and compliance with the second law. Objectivity, material symmetry, and normalization are embedded directly into the architecture through invariant-based representations and zero-anchored formulations. We demonstrate the performance of the proposed framework on synthetic and experimental datasets, including purely thermal problems and fully coupled thermomechanical responses of soft tissues and filled rubbers. The results show that the learned models accurately capture the underlying constitutive behavior. All code, data, and trained models are made publicly available via https://doi.org/10.5281/zenodo.19248596.
Atrial constitutive neural networks
Apr 03, 2025


Abstract:This work presents a novel approach for characterizing the mechanical behavior of atrial tissue using constitutive neural networks. Based on experimental biaxial tensile test data of healthy human atria, we automatically discover the most appropriate constitutive material model, thereby overcoming the limitations of traditional, pre-defined models. This approach offers a new perspective on modeling atrial mechanics and is a significant step towards improved simulation and prediction of cardiac health.
Discovering uncertainty: Gaussian constitutive neural networks with correlated weights
Mar 16, 2025Abstract:When characterizing materials, it can be important to not only predict their mechanical properties, but also to estimate the probability distribution of these properties across a set of samples. Constitutive neural networks allow for the automated discovery of constitutive models that exactly satisfy physical laws given experimental testing data, but are only capable of predicting the mean stress response. Stochastic methods treat each weight as a random variable and are capable of learning their probability distributions. Bayesian constitutive neural networks combine both methods, but their weights lack physical interpretability and we must sample each weight from a probability distribution to train or evaluate the model. Here we introduce a more interpretable network with fewer parameters, simpler training, and the potential to discover correlated weights: Gaussian constitutive neural networks. We demonstrate the performance of our new Gaussian network on biaxial testing data, and discover a sparse and interpretable four-term model with correlated weights. Importantly, the discovered distributions of material parameters across a set of samples can serve as priors to discover better constitutive models for new samples with limited data. We anticipate that Gaussian constitutive neural networks are a natural first step towards generative constitutive models informed by physical laws and parameter uncertainty.
A generalized dual potential for inelastic Constitutive Artificial Neural Networks: A JAX implementation at finite strains
Feb 19, 2025



Abstract:We present a methodology for designing a generalized dual potential, or pseudo potential, for inelastic Constitutive Artificial Neural Networks (iCANNs). This potential, expressed in terms of stress invariants, inherently satisfies thermodynamic consistency for large deformations. In comparison to our previous work, the new potential captures a broader spectrum of material behaviors, including pressure-sensitive inelasticity. To this end, we revisit the underlying thermodynamic framework of iCANNs for finite strain inelasticity and derive conditions for constructing a convex, zero-valued, and non-negative dual potential. To embed these principles in a neural network, we detail the architecture's design, ensuring a priori compliance with thermodynamics. To evaluate the proposed architecture, we study its performance and limitations discovering visco-elastic material behavior, though the method is not limited to visco-elasticity. In this context, we investigate different aspects in the strategy of discovering inelastic materials. Our results indicate that the novel architecture robustly discovers interpretable models and parameters, while autonomously revealing the degree of inelasticity. The iCANN framework, implemented in JAX, is publicly accessible at https://doi.org/10.5281/zenodo.14894687.
Automated Model Discovery for Tensional Homeostasis: Constitutive Machine Learning in Growth and Remodeling
Oct 17, 2024



Abstract:Soft biological tissues exhibit a tendency to maintain a preferred state of tensile stress, known as tensional homeostasis, which is restored even after external mechanical stimuli. This macroscopic behavior can be described using the theory of kinematic growth, where the deformation gradient is multiplicatively decomposed into an elastic part and a part related to growth and remodeling. Recently, the concept of homeostatic surfaces was introduced to define the state of homeostasis and the evolution equations for inelastic deformations. However, identifying the optimal model and material parameters to accurately capture the macroscopic behavior of inelastic materials can only be accomplished with significant expertise, is often time-consuming, and prone to error, regardless of the specific inelastic phenomenon. To address this challenge, built-in physics machine learning algorithms offer significant potential. In this work, we extend our inelastic Constitutive Artificial Neural Networks (iCANNs) by incorporating kinematic growth and homeostatic surfaces to discover the scalar model equations, namely the Helmholtz free energy and the pseudo potential. The latter describes the state of homeostasis in a smeared sense. We evaluate the ability of the proposed network to learn from experimentally obtained tissue equivalent data at the material point level, assess its predictive accuracy beyond the training regime, and discuss its current limitations when applied at the structural level. Our source code, data, examples, and an implementation of the corresponding material subroutine are made accessible to the public at https://doi.org/10.5281/zenodo.13946282.
Theory and implementation of inelastic Constitutive Artificial Neural Networks
Nov 10, 2023



Abstract:Nature has always been our inspiration in the research, design and development of materials and has driven us to gain a deep understanding of the mechanisms that characterize anisotropy and inelastic behavior. All this knowledge has been accumulated in the principles of thermodynamics. Deduced from these principles, the multiplicative decomposition combined with pseudo potentials are powerful and universal concepts. Simultaneously, the tremendous increase in computational performance enabled us to investigate and rethink our history-dependent material models to make the most of our predictions. Today, we have reached a point where materials and their models are becoming increasingly sophisticated. This raises the question: How do we find the best model that includes all inelastic effects to explain our complex data? Constitutive Artificial Neural Networks (CANN) may answer this question. Here, we extend the CANNs to inelastic materials (iCANN). Rigorous considerations of objectivity, rigid motion of the reference configuration, multiplicative decomposition and its inherent non-uniqueness, restrictions of energy and pseudo potential, and consistent evolution guide us towards the architecture of the iCANN satisfying thermodynamics per design. We combine feed-forward networks of the free energy and pseudo potential with a recurrent neural network approach to take time dependencies into account. We demonstrate that the iCANN is capable of autonomously discovering models for artificially generated data, the response of polymers for cyclic loading and the relaxation behavior of muscle data. As the design of the network is not limited to visco-elasticity, our vision is that the iCANN will reveal to us new ways to find the various inelastic phenomena hidden in the data and to understand their interaction. Our source code, data, and examples are available at doi.org/10.5281/zenodo.10066805
On sparse regression, Lp-regularization, and automated model discovery
Oct 09, 2023Abstract:Sparse regression and feature extraction are the cornerstones of knowledge discovery from massive data. Their goal is to discover interpretable and predictive models that provide simple relationships among scientific variables. While the statistical tools for model discovery are well established in the context of linear regression, their generalization to nonlinear regression in material modeling is highly problem-specific and insufficiently understood. Here we explore the potential of neural networks for automatic model discovery and induce sparsity by a hybrid approach that combines two strategies: regularization and physical constraints. We integrate the concept of Lp regularization for subset selection with constitutive neural networks that leverage our domain knowledge in kinematics and thermodynamics. We train our networks with both, synthetic and real data, and perform several thousand discovery runs to infer common guidelines and trends: L2 regularization or ridge regression is unsuitable for model discovery; L1 regularization or lasso promotes sparsity, but induces strong bias; only L0 regularization allows us to transparently fine-tune the trade-off between interpretability and predictability, simplicity and accuracy, and bias and variance. With these insights, we demonstrate that Lp regularized constitutive neural networks can simultaneously discover both, interpretable models and physically meaningful parameters. We anticipate that our findings will generalize to alternative discovery techniques such as sparse and symbolic regression, and to other domains such as biology, chemistry, or medicine. Our ability to automatically discover material models from data could have tremendous applications in generative material design and open new opportunities to manipulate matter, alter properties of existing materials, and discover new materials with user-defined properties.
Discovering a reaction-diffusion model for Alzheimer's disease by combining PINNs with symbolic regression
Jul 16, 2023Abstract:Misfolded tau proteins play a critical role in the progression and pathology of Alzheimer's disease. Recent studies suggest that the spatio-temporal pattern of misfolded tau follows a reaction-diffusion type equation. However, the precise mathematical model and parameters that characterize the progression of misfolded protein across the brain remain incompletely understood. Here, we use deep learning and artificial intelligence to discover a mathematical model for the progression of Alzheimer's disease using longitudinal tau positron emission tomography from the Alzheimer's Disease Neuroimaging Initiative database. Specifically, we integrate physics informed neural networks (PINNs) and symbolic regression to discover a reaction-diffusion type partial differential equation for tau protein misfolding and spreading. First, we demonstrate the potential of our model and parameter discovery on synthetic data. Then, we apply our method to discover the best model and parameters to explain tau imaging data from 46 individuals who are likely to develop Alzheimer's disease and 30 healthy controls. Our symbolic regression discovers different misfolding models $f(c)$ for two groups, with a faster misfolding for the Alzheimer's group, $f(c) = 0.23c^3 - 1.34c^2 + 1.11c$, than for the healthy control group, $f(c) = -c^3 +0.62c^2 + 0.39c$. Our results suggest that PINNs, supplemented by symbolic regression, can discover a reaction-diffusion type model to explain misfolded tau protein concentrations in Alzheimer's disease. We expect our study to be the starting point for a more holistic analysis to provide image-based technologies for early diagnosis, and ideally early treatment of neurodegeneration in Alzheimer's disease and possibly other misfolding-protein based neurodegenerative disorders.
Physics-guided deep learning for data scarcity
Nov 24, 2022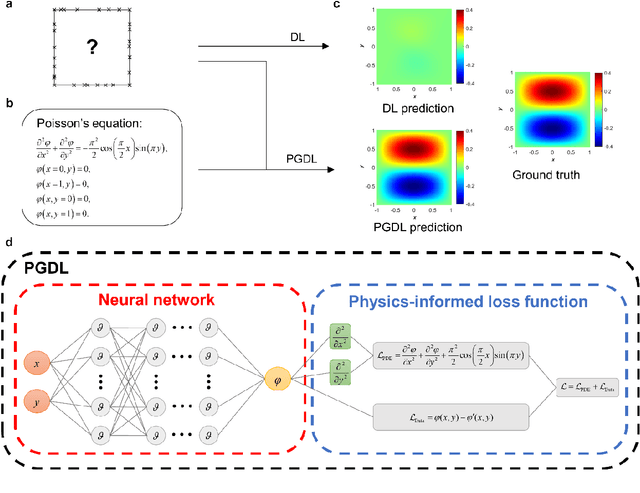
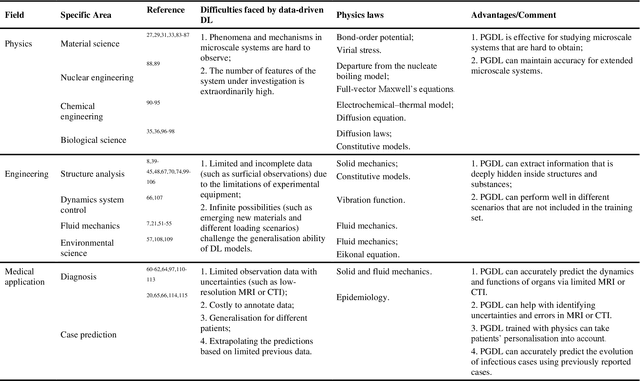
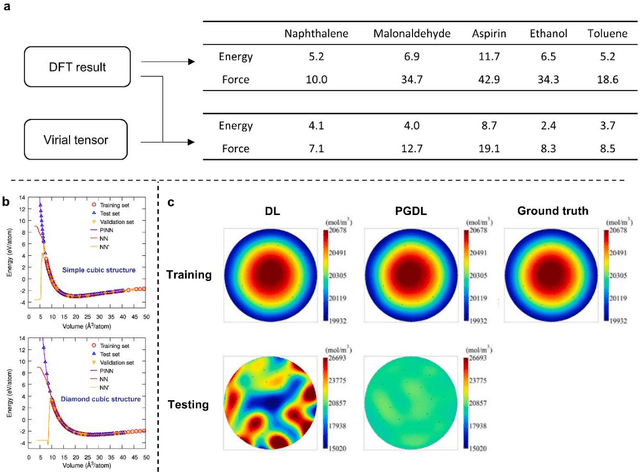
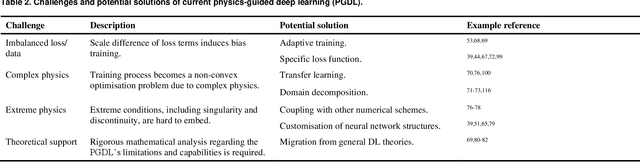
Abstract:Data are the core of deep learning (DL), and the quality of data significantly affects the performance of DL models. However, high-quality and well-annotated databases are hard or even impossible to acquire for use in many applications, such as structural risk estimation and medical diagnosis, which is an essential barrier that blocks the applications of DL in real life. Physics-guided deep learning (PGDL) is a novel type of DL that can integrate physics laws to train neural networks. It can be used for any systems that are controlled or governed by physics laws, such as mechanics, finance and medical applications. It has been shown that, with the additional information provided by physics laws, PGDL achieves great accuracy and generalisation when facing data scarcity. In this review, the details of PGDL are elucidated, and a structured overview of PGDL with respect to data scarcity in various applications is presented, including physics, engineering and medical applications. Moreover, the limitations and opportunities for current PGDL in terms of data scarcity are identified, and the future outlook for PGDL is discussed in depth.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge