Dominik Müller
Classifying Mitotic Figures in the MIDOG25 Challenge with Deep Ensemble Learning and Rule Based Refinement
Aug 28, 2025


Abstract:Mitotic figures (MFs) are relevant biomarkers in tumor grading. Differentiating atypical MFs (AMFs) from normal MFs (NMFs) remains difficult, as manual annotation is time-consuming and subjective. In this work an ensemble of ConvNeXtBase models was trained with AUCMEDI and extend with a rule-based refinement (RBR) module. On the MIDOG25 preliminary test set, the ensemble achieved a balanced accuracy of 84.02%. While the RBR increased specificity, it reduced sensitivity and overall performance. The results show that deep ensembles perform well for AMF classification. RBR can increase specific metrics but requires further research.
Assessing the Performance of Deep Learning for Automated Gleason Grading in Prostate Cancer
Mar 25, 2024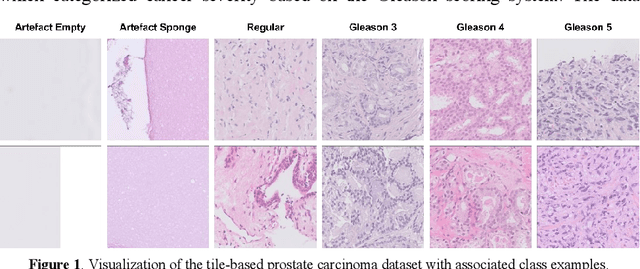
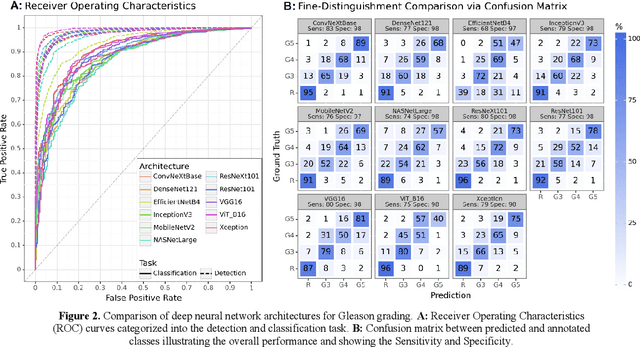
Abstract:Prostate cancer is a dominant health concern calling for advanced diagnostic tools. Utilizing digital pathology and artificial intelligence, this study explores the potential of 11 deep neural network architectures for automated Gleason grading in prostate carcinoma focusing on comparing traditional and recent architectures. A standardized image classification pipeline, based on the AUCMEDI framework, facilitated robust evaluation using an in-house dataset consisting of 34,264 annotated tissue tiles. The results indicated varying sensitivity across architectures, with ConvNeXt demonstrating the strongest performance. Notably, newer architectures achieved superior performance, even though with challenges in differentiating closely related Gleason grades. The ConvNeXt model was capable of learning a balance between complexity and generalizability. Overall, this study lays the groundwork for enhanced Gleason grading systems, potentially improving diagnostic efficiency for prostate cancer.
DeepGleason: a System for Automated Gleason Grading of Prostate Cancer using Deep Neural Networks
Mar 25, 2024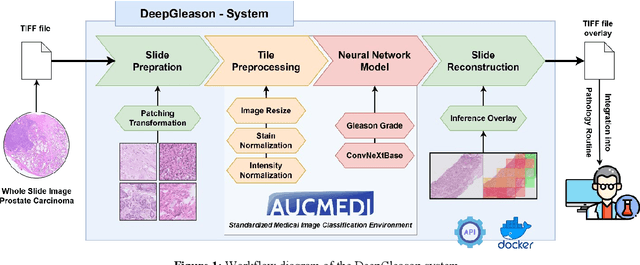
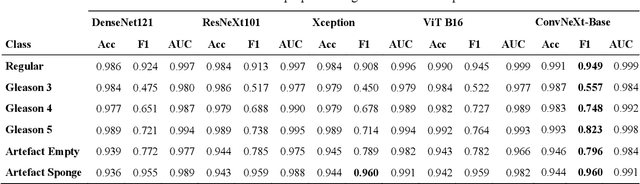
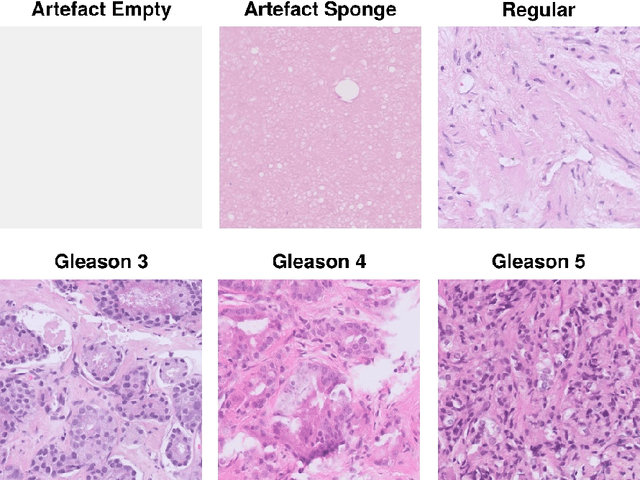
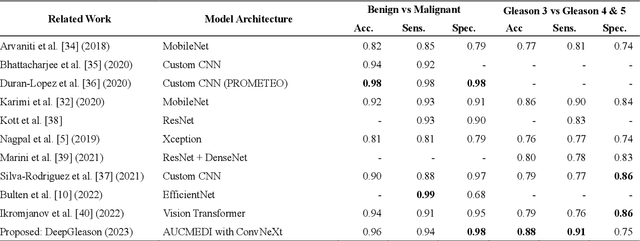
Abstract:Advances in digital pathology and artificial intelligence (AI) offer promising opportunities for clinical decision support and enhancing diagnostic workflows. Previous studies already demonstrated AI's potential for automated Gleason grading, but lack state-of-the-art methodology and model reusability. To address this issue, we propose DeepGleason: an open-source deep neural network based image classification system for automated Gleason grading using whole-slide histopathology images from prostate tissue sections. Implemented with the standardized AUCMEDI framework, our tool employs a tile-wise classification approach utilizing fine-tuned image preprocessing techniques in combination with a ConvNeXt architecture which was compared to various state-of-the-art architectures. The neural network model was trained and validated on an in-house dataset of 34,264 annotated tiles from 369 prostate carcinoma slides. We demonstrated that DeepGleason is capable of highly accurate and reliable Gleason grading with a macro-averaged F1-score of 0.806, AUC of 0.991, and Accuracy of 0.974. The internal architecture comparison revealed that the ConvNeXt model was superior performance-wise on our dataset to established and other modern architectures like transformers. Furthermore, we were able to outperform the current state-of-the-art in tile-wise fine-classification with a sensitivity and specificity of 0.94 and 0.98 for benign vs malignant detection as well as of 0.91 and 0.75 for Gleason 3 vs Gleason 4 & 5 classification, respectively. Our tool contributes to the wider adoption of AI-based Gleason grading within the research community and paves the way for broader clinical application of deep learning models in digital pathology. DeepGleason is open-source and publicly available for research application in the following Git repository: https://github.com/frankkramer-lab/DeepGleason.
Towards Automated COVID-19 Presence and Severity Classification
May 15, 2023Abstract:COVID-19 presence classification and severity prediction via (3D) thorax computed tomography scans have become important tasks in recent times. Especially for capacity planning of intensive care units, predicting the future severity of a COVID-19 patient is crucial. The presented approach follows state-of-theart techniques to aid medical professionals in these situations. It comprises an ensemble learning strategy via 5-fold cross-validation that includes transfer learning and combines pre-trained 3D-versions of ResNet34 and DenseNet121 for COVID19 classification and severity prediction respectively. Further, domain-specific preprocessing was applied to optimize model performance. In addition, medical information like the infection-lung-ratio, patient age, and sex were included. The presented model achieves an AUC of 79.0% to predict COVID-19 severity, and 83.7% AUC to classify the presence of an infection, which is comparable with other currently popular methods. This approach is implemented using the AUCMEDI framework and relies on well-known network architectures to ensure robustness and reproducibility.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
MISm: A Medical Image Segmentation Metric for Evaluation of weak labeled Data
Oct 24, 2022

Abstract:Performance measures are an important tool for assessing and comparing different medical image segmentation algorithms. Unfortunately, the current measures have their weaknesses when it comes to assessing certain edge cases. These limitations arouse when images with a very small region of interest or without a region of interest at all are assessed. As a solution for these limitations, we propose a new medical image segmentation metric: MISm. To evaluate MISm, the popular metrics in the medical image segmentation and MISm were compared using images of magnet resonance tomography from several scenarios. In order to allow application in the community and reproducibility of experimental results, we included MISm in the publicly available evaluation framework MISeval: https://github.com/frankkramer-lab/miseval/tree/master/miseval
Nucleus Segmentation and Analysis in Breast Cancer with the MIScnn Framework
Jun 16, 2022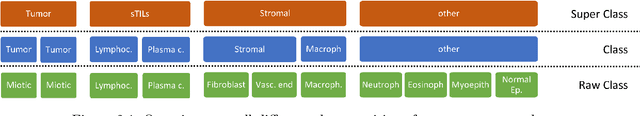
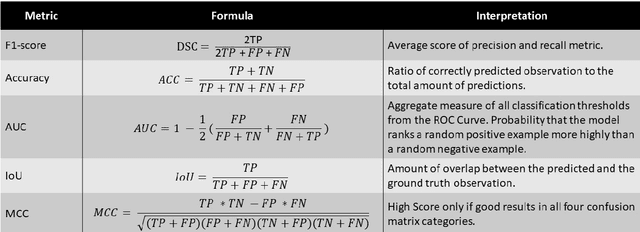

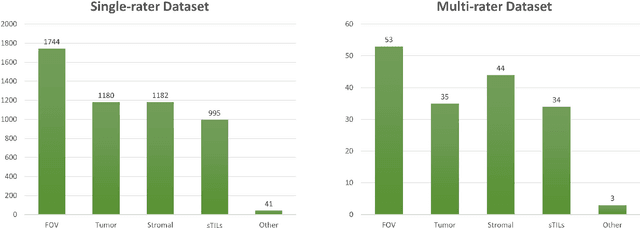

Abstract:The NuCLS dataset contains over 220.000 annotations of cell nuclei in breast cancers. We show how to use these data to create a multi-rater model with the MIScnn Framework to automate the analysis of cell nuclei. For the model creation, we use the widespread U-Net approach embedded in a pipeline. This pipeline provides besides the high performance convolution neural network, several preprocessor techniques and a extended data exploration. The final model is tested in the evaluation phase using a wide variety of metrics with a subsequent visualization. Finally, the results are compared and interpreted with the results of the NuCLS study. As an outlook, indications are given which are important for the future development of models in the context of cell nuclei.
Towards a Guideline for Evaluation Metrics in Medical Image Segmentation
Feb 10, 2022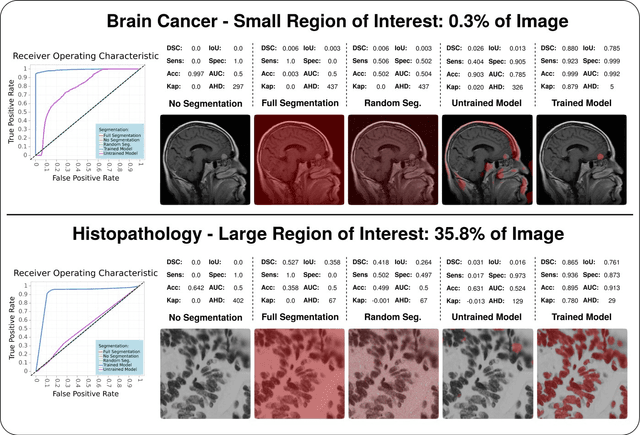
Abstract:In the last decade, research on artificial intelligence has seen rapid growth with deep learning models, especially in the field of medical image segmentation. Various studies demonstrated that these models have powerful prediction capabilities and achieved similar results as clinicians. However, recent studies revealed that the evaluation in image segmentation studies lacks reliable model performance assessment and showed statistical bias by incorrect metric implementation or usage. Thus, this work provides an overview and interpretation guide on the following metrics for medical image segmentation evaluation in binary as well as multi-class problems: Dice similarity coefficient, Jaccard, Sensitivity, Specificity, Rand index, ROC curves, Cohen's Kappa, and Hausdorff distance. As a summary, we propose a guideline for standardized medical image segmentation evaluation to improve evaluation quality, reproducibility, and comparability in the research field.
An Analysis on Ensemble Learning optimized Medical Image Classification with Deep Convolutional Neural Networks
Jan 27, 2022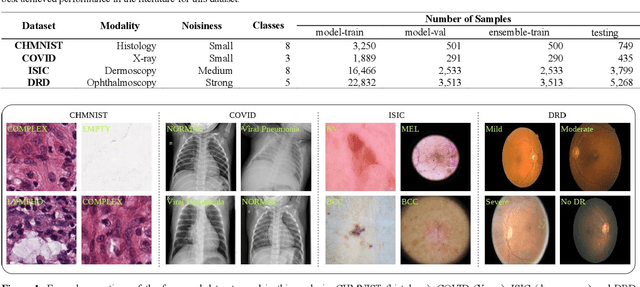
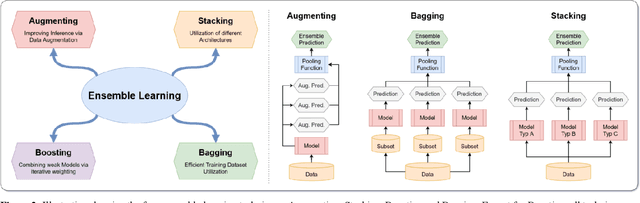
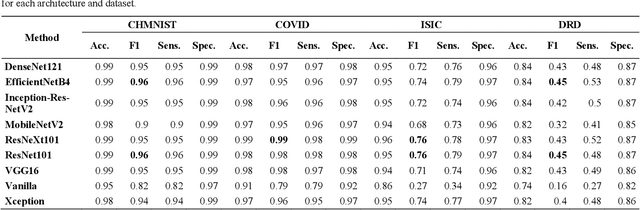
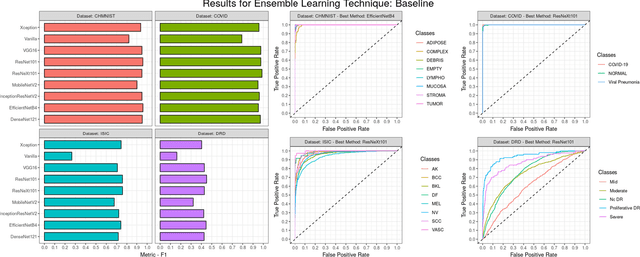
Abstract:Novel and high-performance medical image classification pipelines are heavily utilizing ensemble learning strategies. The idea of ensemble learning is to assemble diverse models or multiple predictions and, thus, boost prediction performance. However, it is still an open question to what extent as well as which ensemble learning strategies are beneficial in deep learning based medical image classification pipelines. In this work, we proposed a reproducible medical image classification pipeline for analyzing the performance impact of the following ensemble learning techniques: Augmenting, Stacking, and Bagging. The pipeline consists of state-of-the-art preprocessing and image augmentation methods as well as 9 deep convolution neural network architectures. It was applied on four popular medical imaging datasets with varying complexity. Furthermore, 12 pooling functions for combining multiple predictions were analyzed, ranging from simple statistical functions like unweighted averaging up to more complex learning-based functions like support vector machines. Our results revealed that Stacking achieved the largest performance gain of up to 13% F1-score increase. Augmenting showed consistent improvement capabilities by up to 4% and is also applicable to single model based pipelines. Cross-validation based Bagging demonstrated to be the most complex ensemble learning method, which resulted in an F1-score decrease in all analyzed datasets (up to -10%). Furthermore, we demonstrated that simple statistical pooling functions are equal or often even better than more complex pooling functions. We concluded that the integration of Stacking and Augmentation ensemble learning techniques is a powerful method for any medical image classification pipeline to improve robustness and boost performance.
MISeval: a Metric Library for Medical Image Segmentation Evaluation
Jan 23, 2022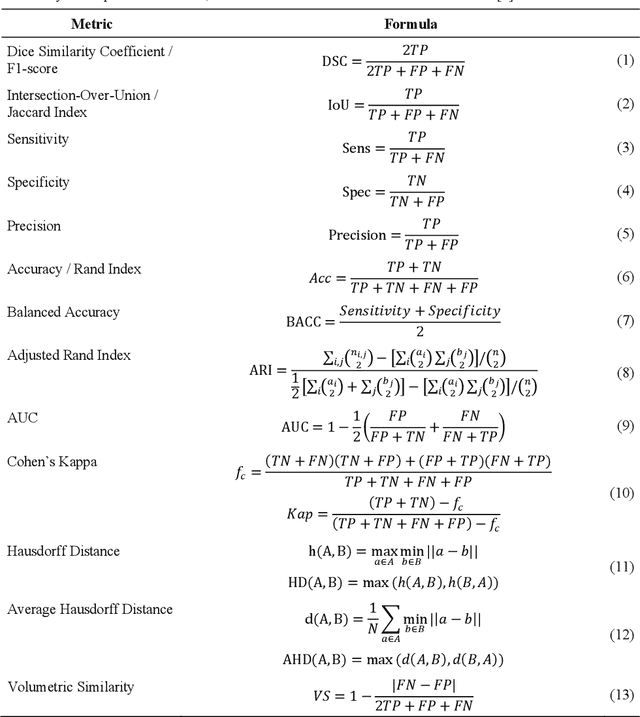
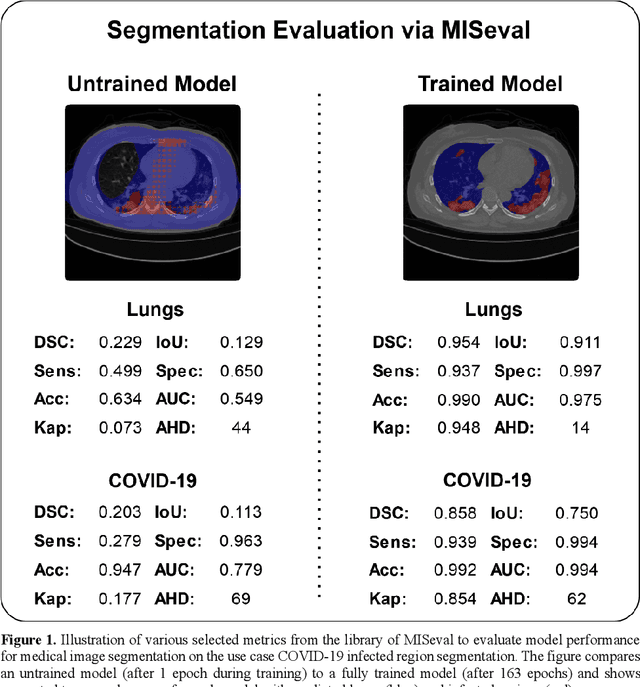
Abstract:Correct performance assessment is crucial for evaluating modern artificial intelligence algorithms in medicine like deep-learning based medical image segmentation models. However, there is no universal metric library in Python for standardized and reproducible evaluation. Thus, we propose our open-source publicly available Python package MISeval: a metric library for Medical Image Segmentation Evaluation. The implemented metrics can be intuitively used and easily integrated into any performance assessment pipeline. The package utilizes modern CI/CD strategies to ensure functionality and stability. MISeval is available from PyPI (miseval) and GitHub: https://github.com/frankkramer-lab/miseval.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge