Dat Q. Tran
VinDr-CXR: An open dataset of chest X-rays with radiologist's annotations
Jan 03, 2021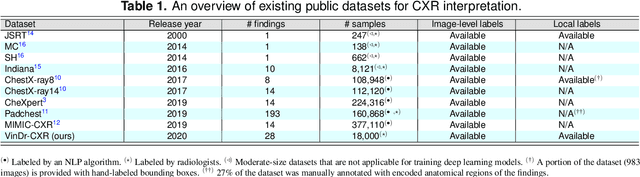
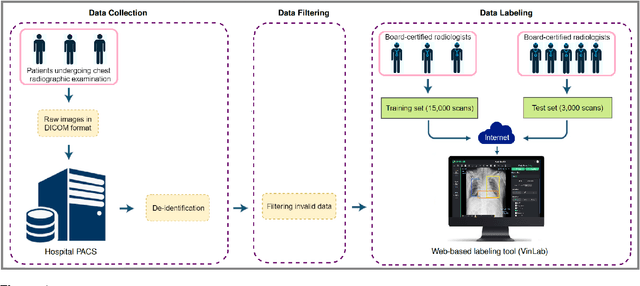
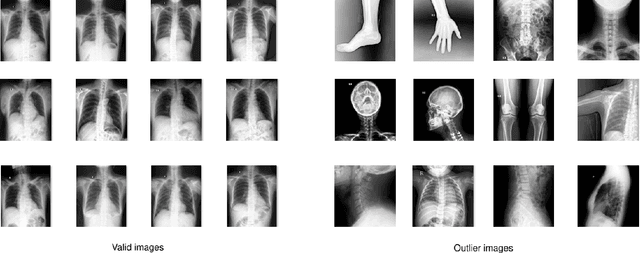
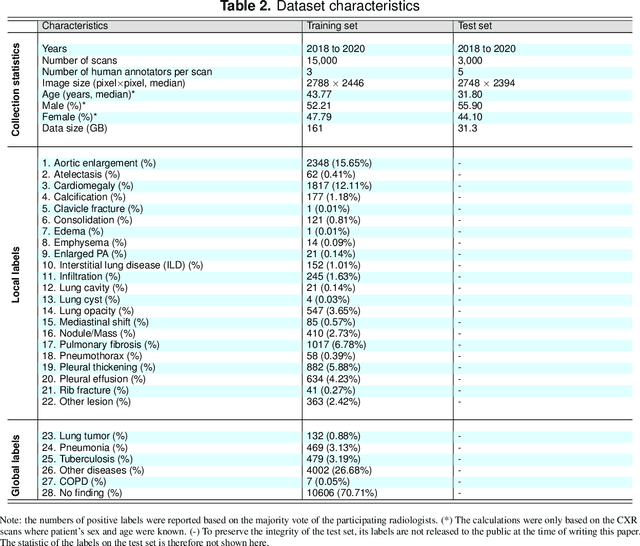
Abstract:Most of the existing chest X-ray datasets include labels from a list of findings without specifying their locations on the radiographs. This limits the development of machine learning algorithms for the detection and localization of chest abnormalities. In this work, we describe a dataset of more than 100,000 chest X-ray scans that were retrospectively collected from two major hospitals in Vietnam. Out of this raw data, we release 18,000 images that were manually annotated by a total of 17 experienced radiologists with 22 local labels of rectangles surrounding abnormalities and 6 global labels of suspected diseases. The released dataset is divided into a training set of 15,000 and a test set of 3,000. Each scan in the training set was independently labeled by 3 radiologists, while each scan in the test set was labeled by the consensus of 5 radiologists. We designed and built a labeling platform for DICOM images to facilitate these annotation procedures. All images are made publicly available in DICOM format in company with the labels of the training set. The labels of the test set are hidden at the time of writing this paper as they will be used for benchmarking machine learning algorithms on an open platform.
A translational pathway of deep learning methods in GastroIntestinal Endoscopy
Oct 12, 2020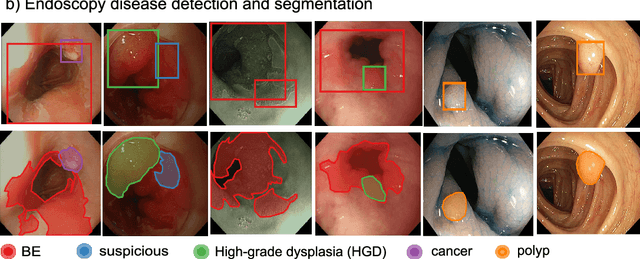
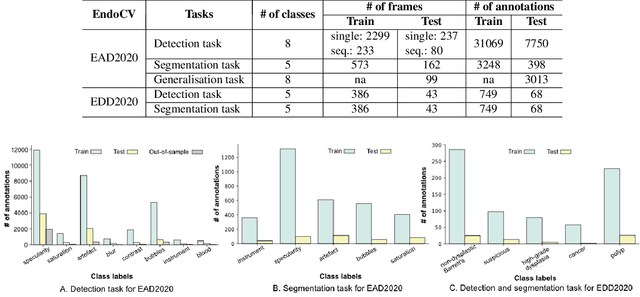
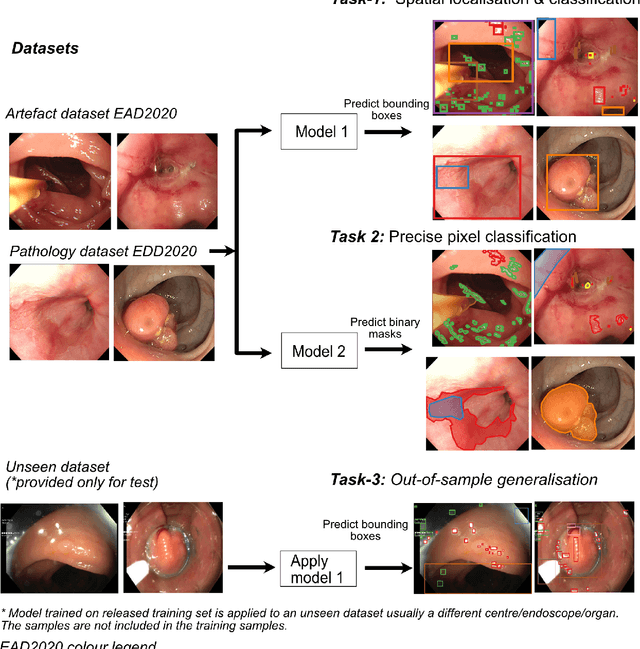
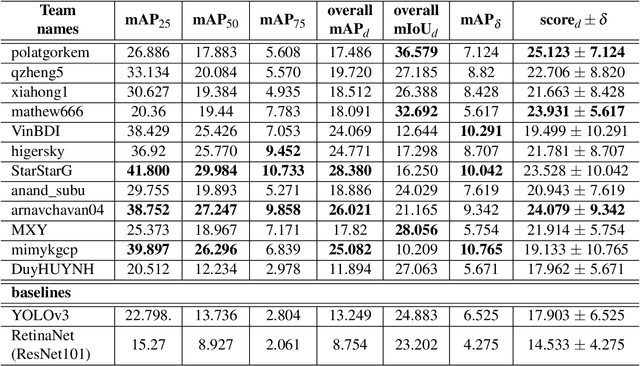
Abstract:The Endoscopy Computer Vision Challenge (EndoCV) is a crowd-sourcing initiative to address eminent problems in developing reliable computer aided detection and diagnosis endoscopy systems and suggest a pathway for clinical translation of technologies. Whilst endoscopy is a widely used diagnostic and treatment tool for hollow-organs, there are several core challenges often faced by endoscopists, mainly: 1) presence of multi-class artefacts that hinder their visual interpretation, and 2) difficulty in identifying subtle precancerous precursors and cancer abnormalities. Artefacts often affect the robustness of deep learning methods applied to the gastrointestinal tract organs as they can be confused with tissue of interest. EndoCV2020 challenges are designed to address research questions in these remits. In this paper, we present a summary of methods developed by the top 17 teams and provide an objective comparison of state-of-the-art methods and methods designed by the participants for two sub-challenges: i) artefact detection and segmentation (EAD2020), and ii) disease detection and segmentation (EDD2020). Multi-center, multi-organ, multi-class, and multi-modal clinical endoscopy datasets were compiled for both EAD2020 and EDD2020 sub-challenges. An out-of-sample generalisation ability of detection algorithms was also evaluated. Whilst most teams focused on accuracy improvements, only a few methods hold credibility for clinical usability. The best performing teams provided solutions to tackle class imbalance, and variabilities in size, origin, modality and occurrences by exploring data augmentation, data fusion, and optimal class thresholding techniques.
Interpreting Chest X-rays via CNNs that Exploit Hierarchical Disease Dependencies and Uncertainty Labels
May 25, 2020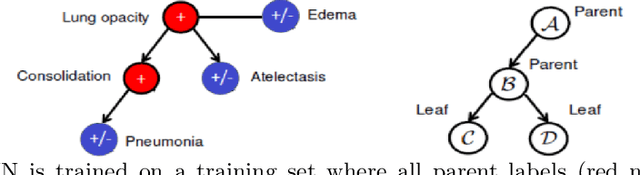
Abstract:The chest X-rays (CXRs) is one of the views most commonly ordered by radiologists (NHS),which is critical for diagnosis of many different thoracic diseases. Accurately detecting thepresence of multiple diseases from CXRs is still a challenging task. We present a multi-labelclassification framework based on deep convolutional neural networks (CNNs) for diagnos-ing the presence of 14 common thoracic diseases and observations. Specifically, we trained astrong set of CNNs that exploit dependencies among abnormality labels and used the labelsmoothing regularization (LSR) for a better handling of uncertain samples. Our deep net-works were trained on over 200,000 CXRs of the recently released CheXpert dataset (Irvinandal., 2019) and the final model, which was an ensemble of the best performing networks,achieved a mean area under the curve (AUC) of 0.940 in predicting 5 selected pathologiesfrom the validation set. To the best of our knowledge, this is the highest AUC score yetreported to date. More importantly, the proposed method was also evaluated on an inde-pendent test set of the CheXpert competition, containing 500 CXR studies annotated by apanel of 5 experienced radiologists. The reported performance was on average better than2.6 out of 3 other individual radiologists with a mean AUC of 0.930, which had led to thecurrent state-of-the-art performance on the CheXpert test set.
A CNN-LSTM Architecture for Detection of Intracranial Hemorrhage on CT scans
May 25, 2020Abstract:We propose a novel method that combines a convolutional neural network (CNN) with a long short-term memory (LSTM) mechanism for accurate prediction of intracranial hemorrhage on computed tomography (CT) scans. The CNN plays the role of a slice-wise feature extractor while the LSTM is responsible for linking the features across slices. The whole architecture is trained end-to-end with input being an RGB-like image formed by stacking 3 different viewing windows of a single slice. We validate the method on the recent RSNA Intracranial Hemorrhage Detection challenge and on the CQ500 dataset. For the RSNA challenge, our best single model achieves a weighted log loss of 0.0522 on the leaderboard, which is comparable to the top 3% performances, almost all of which make use of ensemble learning. Importantly, our method generalizes very well: the model trained on the RSNA dataset significantly outperforms the 2D model, which does not take into account the relationship between slices, on CQ500. Our codes and models is publicly avaiable at https://github.com/nhannguyen2709/RSNA.
Interpreting chest X-rays via CNNs that exploit disease dependencies and uncertainty labels
Nov 18, 2019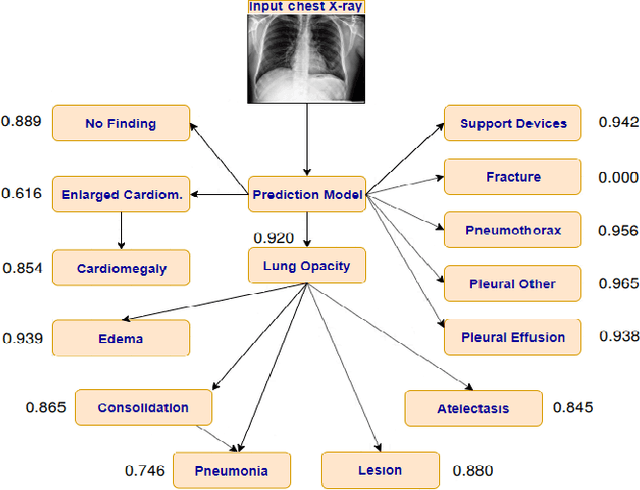
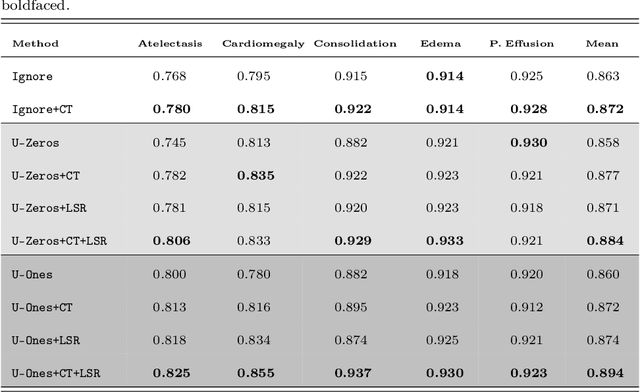

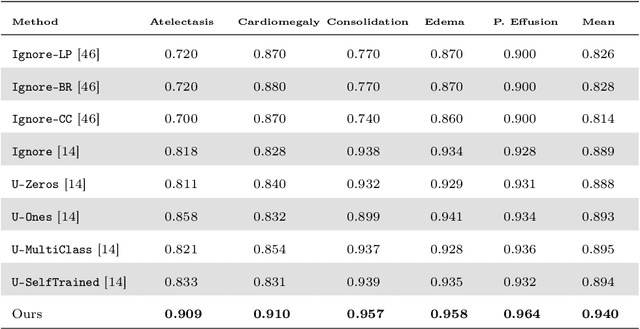
Abstract:Chest radiography is one of the most common types of diagnostic radiology exams, which is critical for screening and diagnosis of many different thoracic diseases. Specialized algorithms have been developed to detect several specific pathologies such as lung nodule or lung cancer. However, accurately detecting the presence of multiple diseases from chest X-rays (CXRs) is still a challenging task. This paper presents a supervised multi-label classification framework based on deep convolutional neural networks (CNNs) for predicting the risk of 14 common thoracic diseases. We tackle this problem by training state-of-the-art CNNs that exploit dependencies among abnormality labels. We also propose to use the label smoothing technique for a better handling of uncertain samples, which occupy a significant portion of almost every CXR dataset. Our model is trained on over 200,000 CXRs of the recently released CheXpert dataset and achieves a mean area under the curve (AUC) of 0.940 in predicting 5 selected pathologies from the validation set. This is the highest AUC score yet reported to date. The proposed method is also evaluated on the independent test set of the CheXpert competition, which is composed of 500 CXR studies annotated by a panel of 5 experienced radiologists. The performance is on average better than 2.6 out of 3 other individual radiologists with a mean AUC of 0.930, which ranks first on the CheXpert leaderboard at the time of writing this paper.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge