Bogdan Matuszewski
FCB-SwinV2 Transformer for Polyp Segmentation
Feb 02, 2023Abstract:Polyp segmentation within colonoscopy video frames using deep learning models has the potential to automate the workflow of clinicians. This could help improve the early detection rate and characterization of polyps which could progress to colorectal cancer. Recent state-of-the-art deep learning polyp segmentation models have combined the outputs of Fully Convolutional Network architectures and Transformer Network architectures which work in parallel. In this paper we propose modifications to the current state-of-the-art polyp segmentation model FCBFormer. The transformer architecture of the FCBFormer is replaced with a SwinV2 Transformer-UNET and minor changes to the Fully Convolutional Network architecture are made to create the FCB-SwinV2 Transformer. The performance of the FCB-SwinV2 Transformer is evaluated on the popular colonoscopy segmentation bench-marking datasets Kvasir-SEG and CVC-ClinicDB. Generalizability tests are also conducted. The FCB-SwinV2 Transformer is able to consistently achieve higher mDice scores across all tests conducted and therefore represents new state-of-the-art performance. Issues found with how colonoscopy segmentation model performance is evaluated within literature are also re-ported and discussed. One of the most important issues identified is that when evaluating performance on the CVC-ClinicDB dataset it would be preferable to ensure no data leakage from video sequences occurs during the training/validation/test data partition.
A translational pathway of deep learning methods in GastroIntestinal Endoscopy
Oct 12, 2020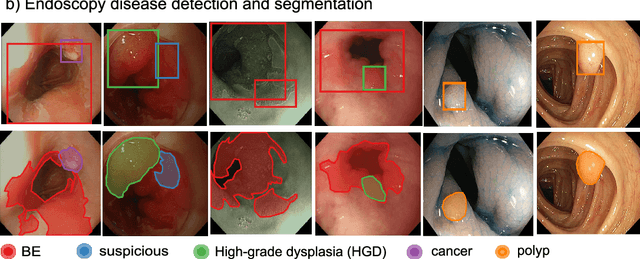
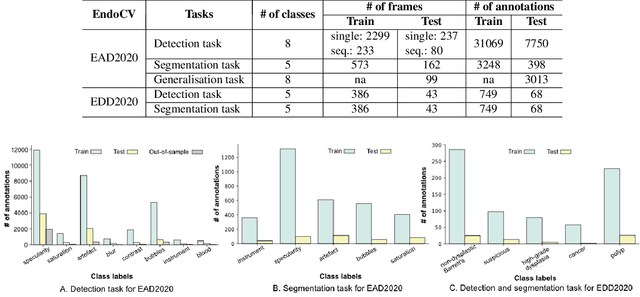
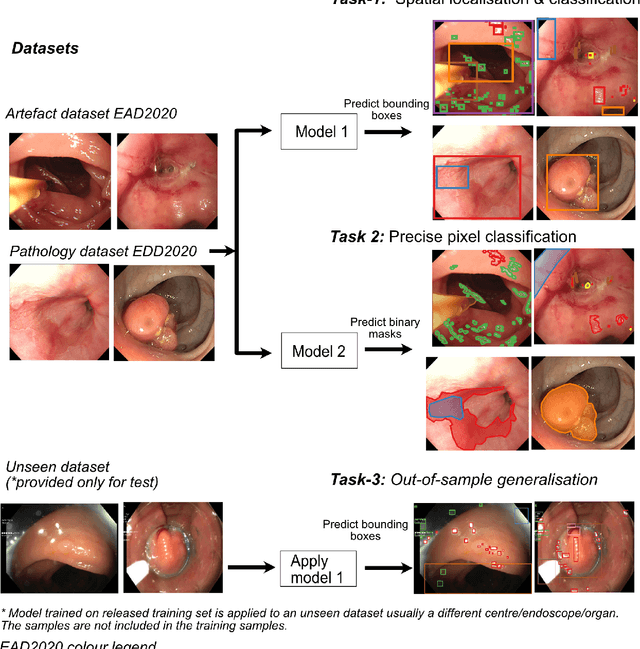
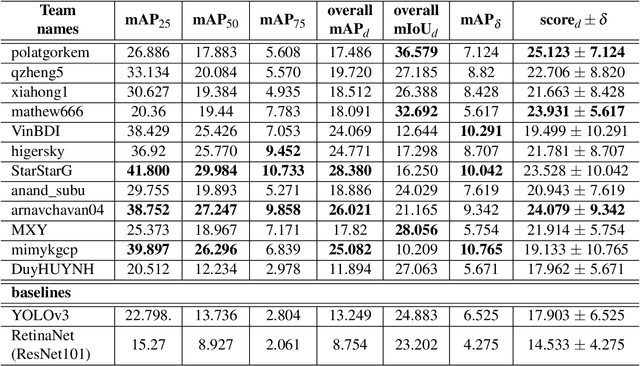
Abstract:The Endoscopy Computer Vision Challenge (EndoCV) is a crowd-sourcing initiative to address eminent problems in developing reliable computer aided detection and diagnosis endoscopy systems and suggest a pathway for clinical translation of technologies. Whilst endoscopy is a widely used diagnostic and treatment tool for hollow-organs, there are several core challenges often faced by endoscopists, mainly: 1) presence of multi-class artefacts that hinder their visual interpretation, and 2) difficulty in identifying subtle precancerous precursors and cancer abnormalities. Artefacts often affect the robustness of deep learning methods applied to the gastrointestinal tract organs as they can be confused with tissue of interest. EndoCV2020 challenges are designed to address research questions in these remits. In this paper, we present a summary of methods developed by the top 17 teams and provide an objective comparison of state-of-the-art methods and methods designed by the participants for two sub-challenges: i) artefact detection and segmentation (EAD2020), and ii) disease detection and segmentation (EDD2020). Multi-center, multi-organ, multi-class, and multi-modal clinical endoscopy datasets were compiled for both EAD2020 and EDD2020 sub-challenges. An out-of-sample generalisation ability of detection algorithms was also evaluated. Whilst most teams focused on accuracy improvements, only a few methods hold credibility for clinical usability. The best performing teams provided solutions to tackle class imbalance, and variabilities in size, origin, modality and occurrences by exploring data augmentation, data fusion, and optimal class thresholding techniques.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge