Christoph Palm
Regensburg Medical Image Computing, Regensburg Center of Health Sciences and Technology
Video Dataset for Surgical Phase, Keypoint, and Instrument Recognition in Laparoscopic Surgery (PhaKIR)
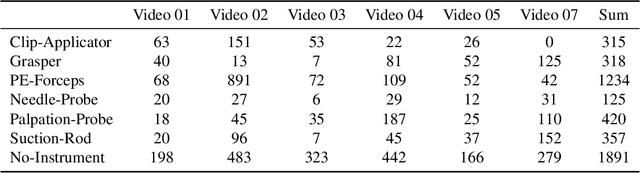
Nov 09, 2025Abstract:Robotic- and computer-assisted minimally invasive surgery (RAMIS) is increasingly relying on computer vision methods for reliable instrument recognition and surgical workflow understanding. Developing such systems often requires large, well-annotated datasets, but existing resources often address isolated tasks, neglect temporal dependencies, or lack multi-center variability. We present the Surgical Procedure Phase, Keypoint, and Instrument Recognition (PhaKIR) dataset, comprising eight complete laparoscopic cholecystectomy videos recorded at three medical centers. The dataset provides frame-level annotations for three interconnected tasks: surgical phase recognition (485,875 frames), instrument keypoint estimation (19,435 frames), and instrument instance segmentation (19,435 frames). PhaKIR is, to our knowledge, the first multi-institutional dataset to jointly provide phase labels, instrument pose information, and pixel-accurate instrument segmentations, while also enabling the exploitation of temporal context since full surgical procedure sequences are available. It served as the basis for the PhaKIR Challenge as part of the Endoscopic Vision (EndoVis) Challenge at MICCAI 2024 to benchmark methods in surgical scene understanding, thereby further validating the dataset's quality and relevance. The dataset is publicly available upon request via the Zenodo platform.
Comparative validation of surgical phase recognition, instrument keypoint estimation, and instrument instance segmentation in endoscopy: Results of the PhaKIR 2024 challenge
Jul 22, 2025Abstract:Reliable recognition and localization of surgical instruments in endoscopic video recordings are foundational for a wide range of applications in computer- and robot-assisted minimally invasive surgery (RAMIS), including surgical training, skill assessment, and autonomous assistance. However, robust performance under real-world conditions remains a significant challenge. Incorporating surgical context - such as the current procedural phase - has emerged as a promising strategy to improve robustness and interpretability. To address these challenges, we organized the Surgical Procedure Phase, Keypoint, and Instrument Recognition (PhaKIR) sub-challenge as part of the Endoscopic Vision (EndoVis) challenge at MICCAI 2024. We introduced a novel, multi-center dataset comprising thirteen full-length laparoscopic cholecystectomy videos collected from three distinct medical institutions, with unified annotations for three interrelated tasks: surgical phase recognition, instrument keypoint estimation, and instrument instance segmentation. Unlike existing datasets, ours enables joint investigation of instrument localization and procedural context within the same data while supporting the integration of temporal information across entire procedures. We report results and findings in accordance with the BIAS guidelines for biomedical image analysis challenges. The PhaKIR sub-challenge advances the field by providing a unique benchmark for developing temporally aware, context-driven methods in RAMIS and offers a high-quality resource to support future research in surgical scene understanding.
OpenMIBOOD: Open Medical Imaging Benchmarks for Out-Of-Distribution Detection
Mar 20, 2025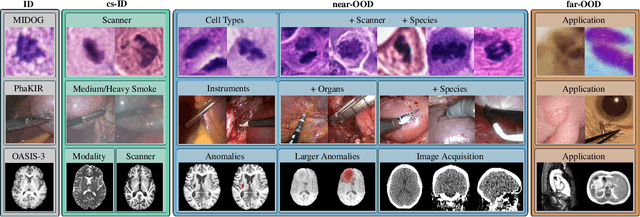
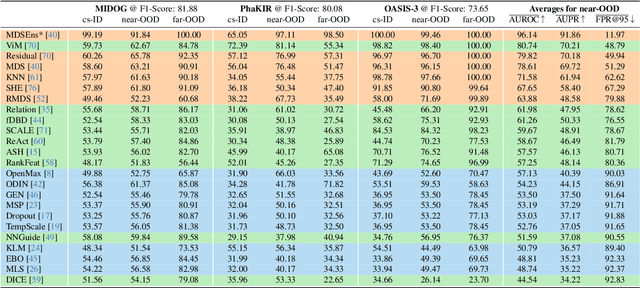
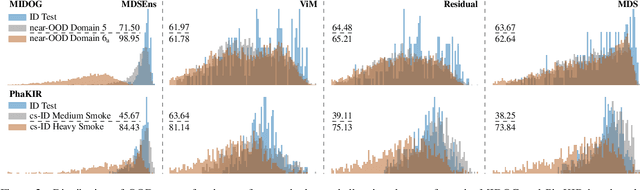
Abstract:The growing reliance on Artificial Intelligence (AI) in critical domains such as healthcare demands robust mechanisms to ensure the trustworthiness of these systems, especially when faced with unexpected or anomalous inputs. This paper introduces the Open Medical Imaging Benchmarks for Out-Of-Distribution Detection (OpenMIBOOD), a comprehensive framework for evaluating out-of-distribution (OOD) detection methods specifically in medical imaging contexts. OpenMIBOOD includes three benchmarks from diverse medical domains, encompassing 14 datasets divided into covariate-shifted in-distribution, near-OOD, and far-OOD categories. We evaluate 24 post-hoc methods across these benchmarks, providing a standardized reference to advance the development and fair comparison of OOD detection methods. Results reveal that findings from broad-scale OOD benchmarks in natural image domains do not translate to medical applications, underscoring the critical need for such benchmarks in the medical field. By mitigating the risk of exposing AI models to inputs outside their training distribution, OpenMIBOOD aims to support the advancement of reliable and trustworthy AI systems in healthcare. The repository is available at https://github.com/remic-othr/OpenMIBOOD.
iRBSM: A Deep Implicit 3D Breast Shape Model
Dec 17, 2024
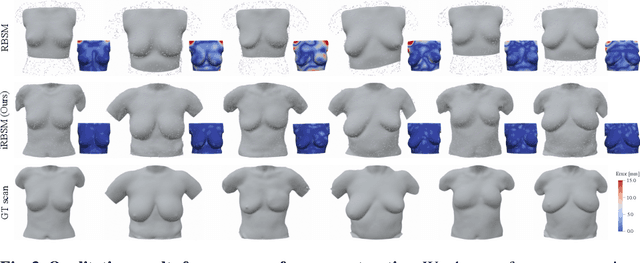
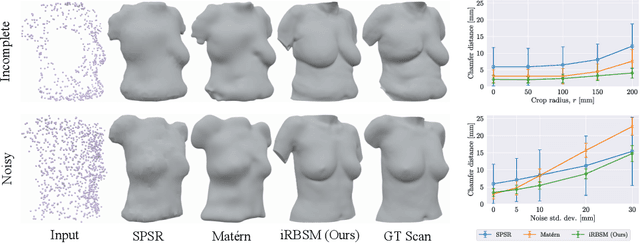
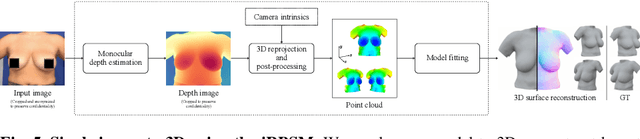
Abstract:We present the first deep implicit 3D shape model of the female breast, building upon and improving the recently proposed Regensburg Breast Shape Model (RBSM). Compared to its PCA-based predecessor, our model employs implicit neural representations; hence, it can be trained on raw 3D breast scans and eliminates the need for computationally demanding non-rigid registration -- a task that is particularly difficult for feature-less breast shapes. The resulting model, dubbed iRBSM, captures detailed surface geometry including fine structures such as nipples and belly buttons, is highly expressive, and outperforms the RBSM on different surface reconstruction tasks. Finally, leveraging the iRBSM, we present a prototype application to 3D reconstruct breast shapes from just a single image. Model and code publicly available at https://rbsm.re-mic.de/implicit.
Motion-Corrected Moving Average: Including Post-Hoc Temporal Information for Improved Video Segmentation
Mar 05, 2024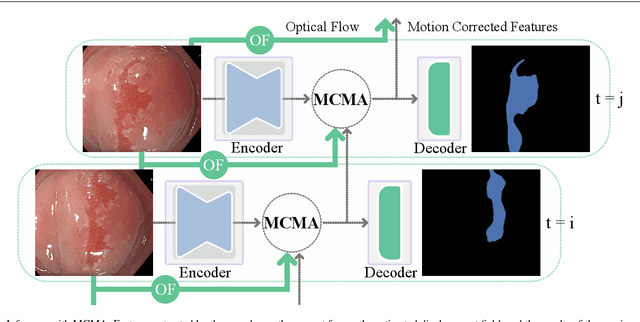
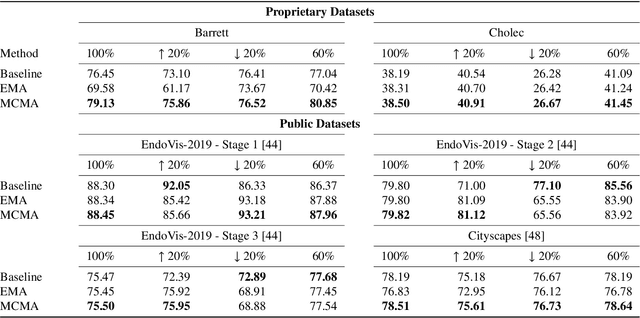
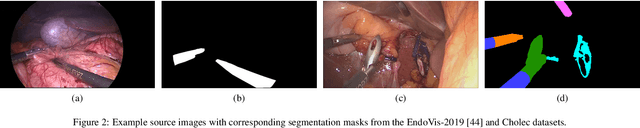
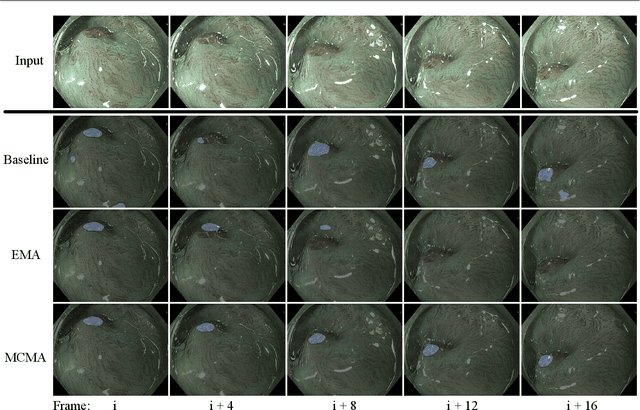
Abstract:Real-time computational speed and a high degree of precision are requirements for computer-assisted interventions. Applying a segmentation network to a medical video processing task can introduce significant inter-frame prediction noise. Existing approaches can reduce inconsistencies by including temporal information but often impose requirements on the architecture or dataset. This paper proposes a method to include temporal information in any segmentation model and, thus, a technique to improve video segmentation performance without alterations during training or additional labeling. With Motion-Corrected Moving Average, we refine the exponential moving average between the current and previous predictions. Using optical flow to estimate the movement between consecutive frames, we can shift the prior term in the moving-average calculation to align with the geometry of the current frame. The optical flow calculation does not require the output of the model and can therefore be performed in parallel, leading to no significant runtime penalty for our approach. We evaluate our approach on two publicly available segmentation datasets and two proprietary endoscopic datasets and show improvements over a baseline approach.
Methods and datasets for segmentation of minimally invasive surgical instruments in endoscopic images and videos: A review of the state of the art
Apr 25, 2023Abstract:In the field of computer- and robot-assisted minimally invasive surgery, enormous progress has been made in recent years based on the recognition of surgical instruments in endoscopic images. Especially the determination of the position and type of the instruments is of great interest here. Current work involves both spatial and temporal information with the idea, that the prediction of movement of surgical tools over time may improve the quality of final segmentations. The provision of publicly available datasets has recently encouraged the development of new methods, mainly based on deep learning. In this review, we identify datasets used for method development and evaluation, as well as quantify their frequency of use in the literature. We further present an overview of the current state of research regarding the segmentation and tracking of minimally invasive surgical instruments in endoscopic images. The paper focuses on methods that work purely visually without attached markers of any kind on the instruments, taking into account both single-frame segmentation approaches as well as those involving temporal information. A discussion of the reviewed literature is provided, highlighting existing shortcomings and emphasizing available potential for future developments. The publications considered were identified through the platforms Google Scholar, Web of Science, and PubMed. The search terms used were "instrument segmentation", "instrument tracking", "surgical tool segmentation", and "surgical tool tracking" and result in 408 articles published between 2015 and 2022 from which 109 were included using systematic selection criteria.
Learning the shape of female breasts: an open-access 3D statistical shape model of the female breast built from 110 breast scans
Jul 28, 2021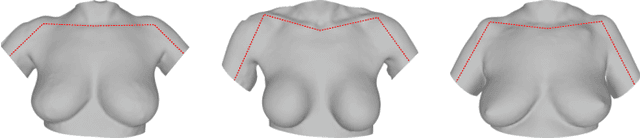
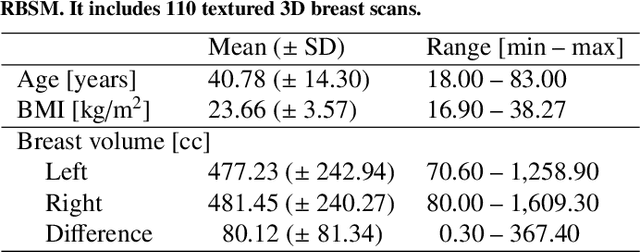
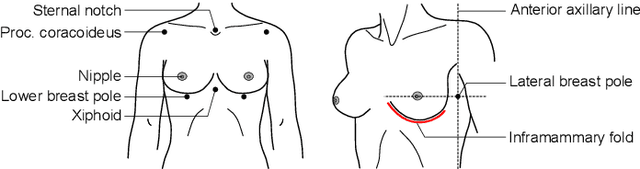
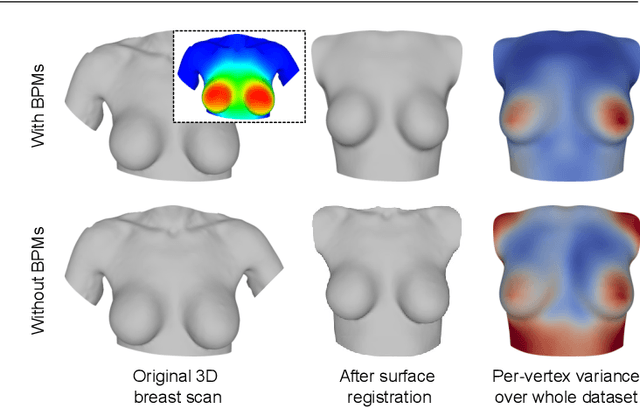
Abstract:We present the Regensburg Breast Shape Model (RBSM) - a 3D statistical shape model of the female breast built from 110 breast scans, and the first ever publicly available. Together with the model, a fully automated, pairwise surface registration pipeline used to establish correspondence among 3D breast scans is introduced. Our method is computationally efficient and requires only four landmarks to guide the registration process. In order to weaken the strong coupling between breast and thorax, we propose to minimize the variance outside the breast region as much as possible. To achieve this goal, a novel concept called breast probability masks (BPMs) is introduced. A BPM assigns probabilities to each point of a 3D breast scan, telling how likely it is that a particular point belongs to the breast area. During registration, we use BPMs to align the template to the target as accurately as possible inside the breast region and only roughly outside. This simple yet effective strategy significantly reduces the unwanted variance outside the breast region, leading to better statistical shape models in which breast shapes are quite well decoupled from the thorax. The RBSM is thus able to produce a variety of different breast shapes as independently as possible from the shape of the thorax. Our systematic experimental evaluation reveals a generalization ability of 0.17 mm and a specificity of 2.8 mm for the RBSM. Ultimately, our model is seen as a first step towards combining physically motivated deformable models of the breast and statistical approaches in order to enable more realistic surgical outcome simulation.
Learning Visual Representations with Optimum-Path Forest and its Applications to Barrett's Esophagus and Adenocarcinoma Diagnosis
Jan 19, 2021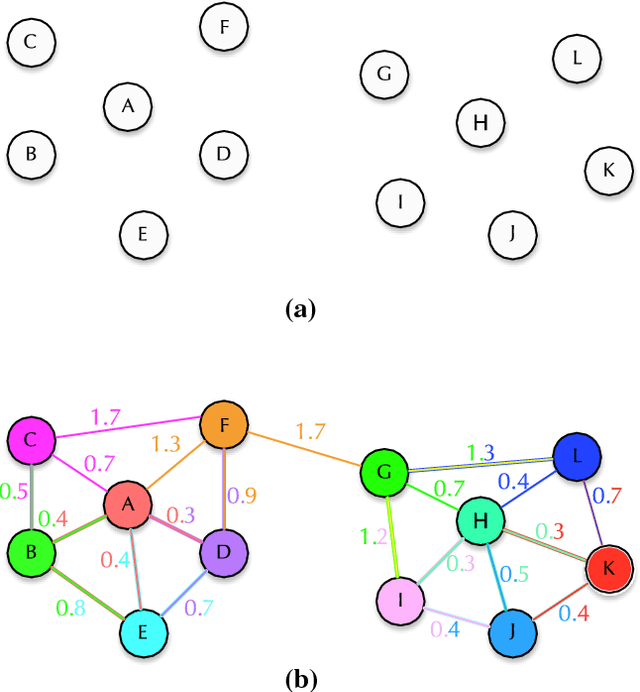
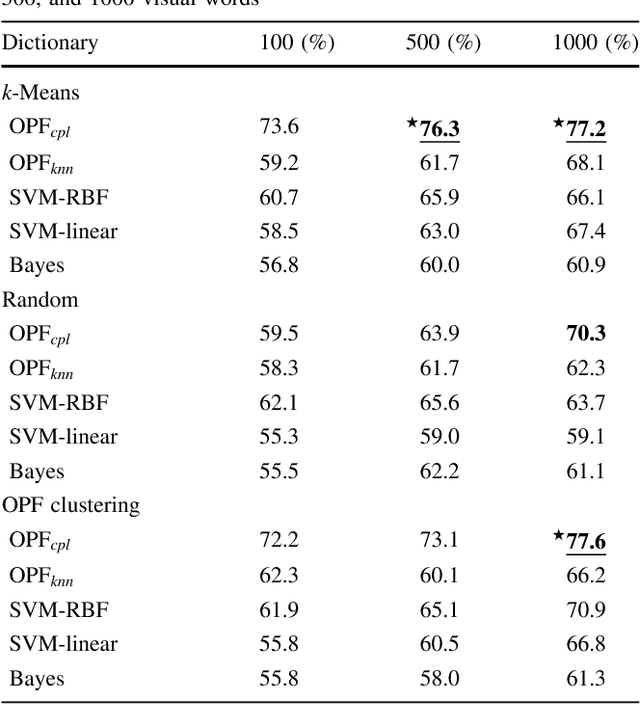
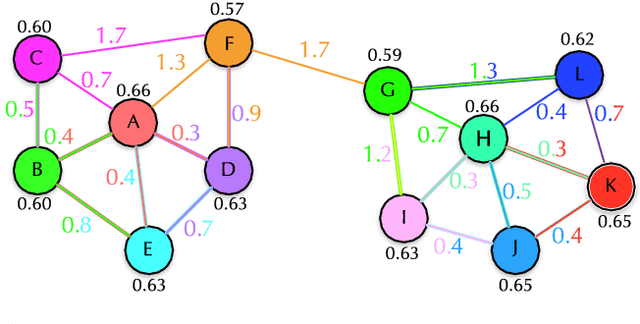
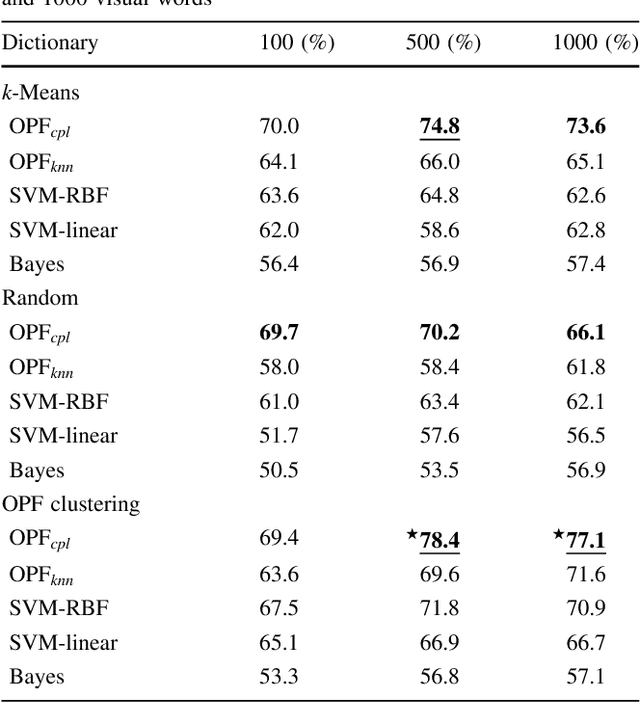
Abstract:In this work, we introduce the unsupervised Optimum-Path Forest (OPF) classifier for learning visual dictionaries in the context of Barrett's esophagus (BE) and automatic adenocarcinoma diagnosis. The proposed approach was validated in two datasets (MICCAI 2015 and Augsburg) using three different feature extractors (SIFT, SURF, and the not yet applied to the BE context A-KAZE), as well as five supervised classifiers, including two variants of the OPF, Support Vector Machines with Radial Basis Function and Linear kernels, and a Bayesian classifier. Concerning MICCAI 2015 dataset, the best results were obtained using unsupervised OPF for dictionary generation using supervised OPF for classification purposes and using SURF feature extractor with accuracy nearly to 78% for distinguishing BE patients from adenocarcinoma ones. Regarding the Augsburg dataset, the most accurate results were also obtained using both OPF classifiers but with A-KAZE as the feature extractor with accuracy close to 73%. The combination of feature extraction and bag-of-visual-words techniques showed results that outperformed others obtained recently in the literature, as well as we highlight new advances in the related research area. Reinforcing the significance of this work, to the best of our knowledge, this is the first one that aimed at addressing computer-aided BE identification using bag-of-visual-words and OPF classifiers, being this application of unsupervised technique in the BE feature calculation the major contribution of this work. It is also proposed a new BE and adenocarcinoma description using the A-KAZE features, not yet applied in the literature.
2018 Robotic Scene Segmentation Challenge
Feb 03, 2020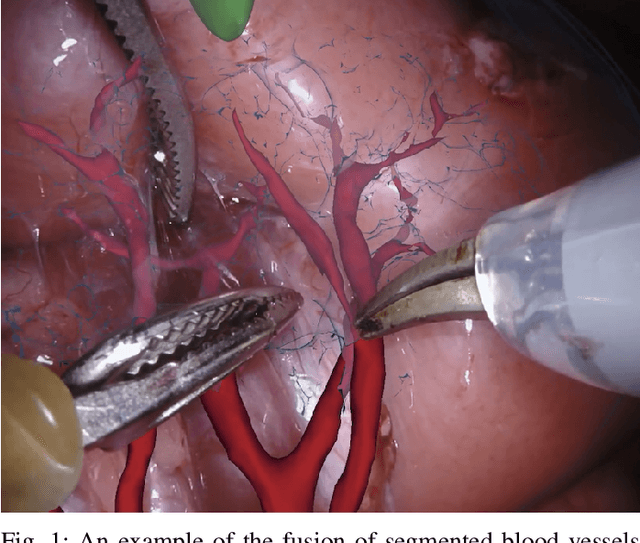
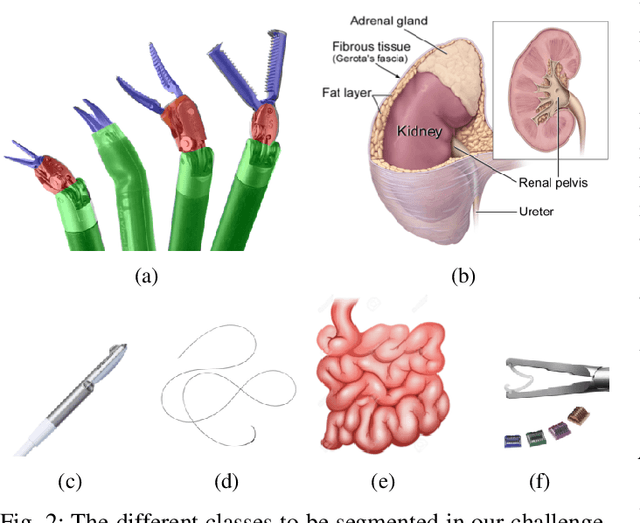
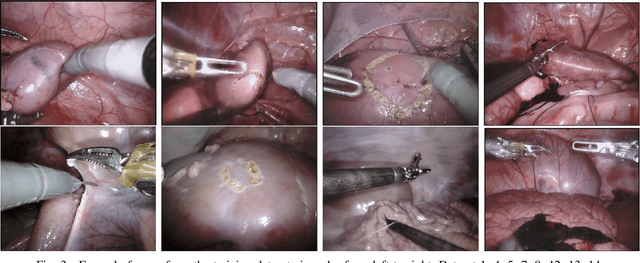
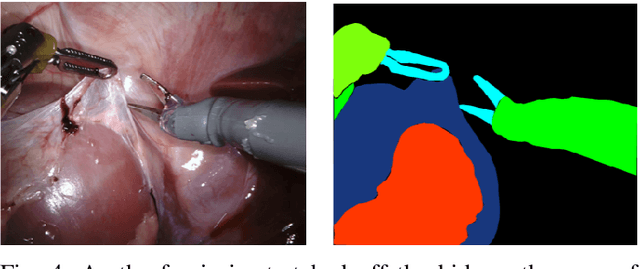
Abstract:In 2015 we began a sub-challenge at the EndoVis workshop at MICCAI in Munich using endoscope images of ex-vivo tissue with automatically generated annotations from robot forward kinematics and instrument CAD models. However, the limited background variation and simple motion rendered the dataset uninformative in learning about which techniques would be suitable for segmentation in real surgery. In 2017, at the same workshop in Quebec we introduced the robotic instrument segmentation dataset with 10 teams participating in the challenge to perform binary, articulating parts and type segmentation of da Vinci instruments. This challenge included realistic instrument motion and more complex porcine tissue as background and was widely addressed with modifications on U-Nets and other popular CNN architectures. In 2018 we added to the complexity by introducing a set of anatomical objects and medical devices to the segmented classes. To avoid over-complicating the challenge, we continued with porcine data which is dramatically simpler than human tissue due to the lack of fatty tissue occluding many organs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge