Arlene Casey
Infusing clinical knowledge into tokenisers for language models
Jun 20, 2024



Abstract:This study introduces a novel knowledge enhanced tokenisation mechanism, K-Tokeniser, for clinical text processing. Technically, at initialisation stage, K-Tokeniser populates global representations of tokens based on semantic types of domain concepts (such as drugs or diseases) from either a domain ontology like Unified Medical Language System or the training data of the task related corpus. At training or inference stage, sentence level localised context will be utilised for choosing the optimal global token representation to realise the semantic-based tokenisation. To avoid pretraining using the new tokeniser, an embedding initialisation approach is proposed to generate representations for new tokens. Using three transformer-based language models, a comprehensive set of experiments are conducted on four real-world datasets for evaluating K-Tokeniser in a wide range of clinical text analytics tasks including clinical concept and relation extraction, automated clinical coding, clinical phenotype identification, and clinical research article classification. Overall, our models demonstrate consistent improvements over their counterparts in all tasks. In particular, substantial improvements are observed in the automated clinical coding task with 13\% increase on Micro $F_1$ score. Furthermore, K-Tokeniser also shows significant capacities in facilitating quicker converge of language models. Specifically, using K-Tokeniser, the language models would only require 50\% of the training data to achieve the best performance of the baseline tokeniser using all training data in the concept extraction task and less than 20\% of the data for the automated coding task. It is worth mentioning that all these improvements require no pre-training process, making the approach generalisable.
Ontology-Based and Weakly Supervised Rare Disease Phenotyping from Clinical Notes
May 11, 2022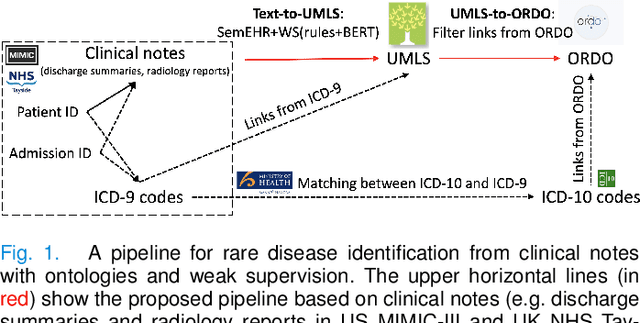
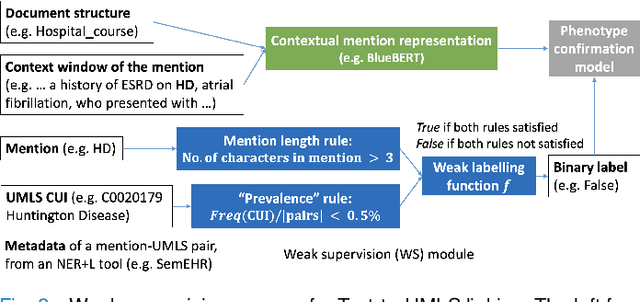
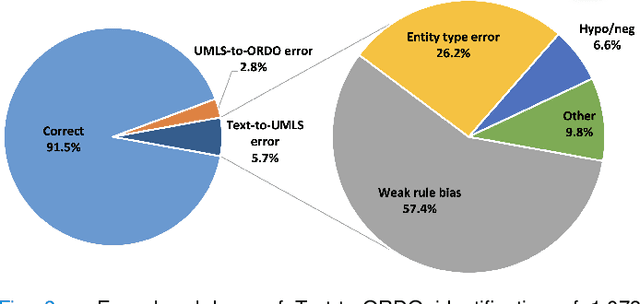
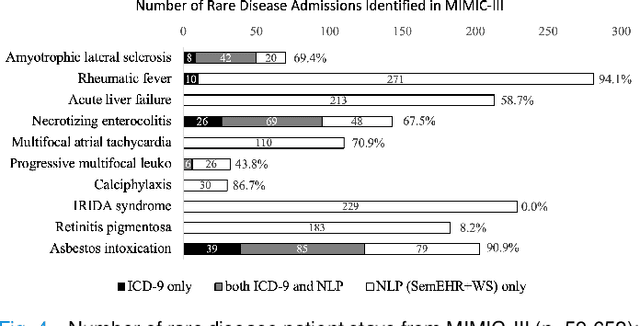
Abstract:Computational text phenotyping is the practice of identifying patients with certain disorders and traits from clinical notes. Rare diseases are challenging to be identified due to few cases available for machine learning and the need for data annotation from domain experts. We propose a method using ontologies and weak supervision, with recent pre-trained contextual representations from Bi-directional Transformers (e.g. BERT). The ontology-based framework includes two steps: (i) Text-to-UMLS, extracting phenotypes by contextually linking mentions to concepts in Unified Medical Language System (UMLS), with a Named Entity Recognition and Linking (NER+L) tool, SemEHR, and weak supervision with customised rules and contextual mention representation; (ii) UMLS-to-ORDO, matching UMLS concepts to rare diseases in Orphanet Rare Disease Ontology (ORDO). The weakly supervised approach is proposed to learn a phenotype confirmation model to improve Text-to-UMLS linking, without annotated data from domain experts. We evaluated the approach on three clinical datasets of discharge summaries and radiology reports from two institutions in the US and the UK. Our best weakly supervised method achieved 81.4% precision and 91.4% recall on extracting rare disease UMLS phenotypes from MIMIC-III discharge summaries. The overall pipeline processing clinical notes can surface rare disease cases, mostly uncaptured in structured data (manually assigned ICD codes). Results on radiology reports from MIMIC-III and NHS Tayside were consistent with the discharge summaries. We discuss the usefulness of the weak supervision approach and propose directions for future studies.
A Systematic Review of Natural Language Processing Applied to Radiology Reports
Feb 18, 2021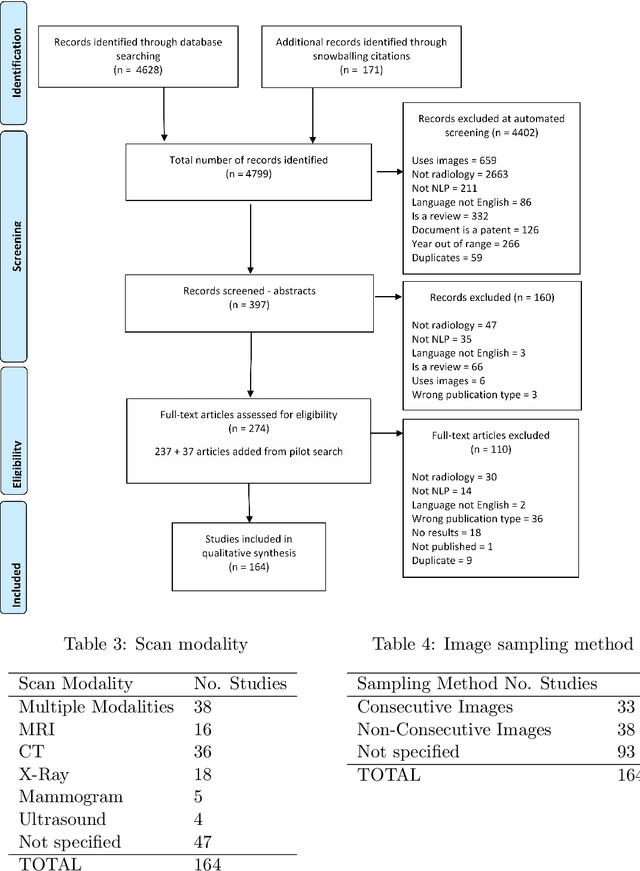
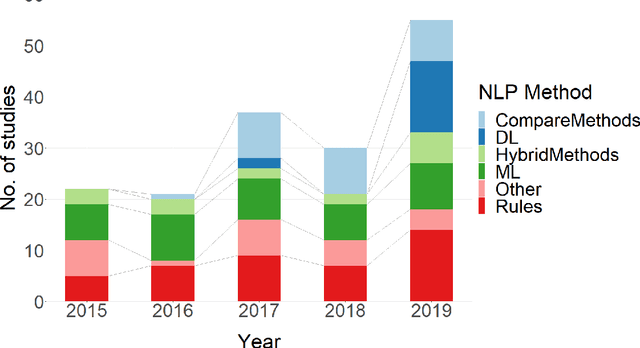
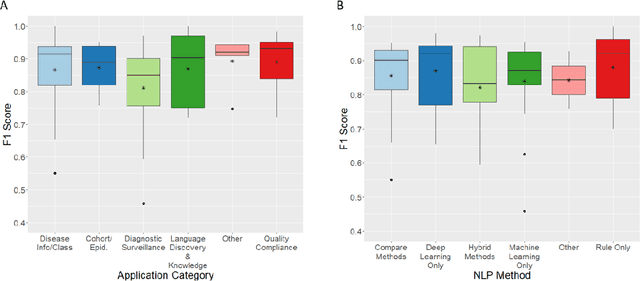
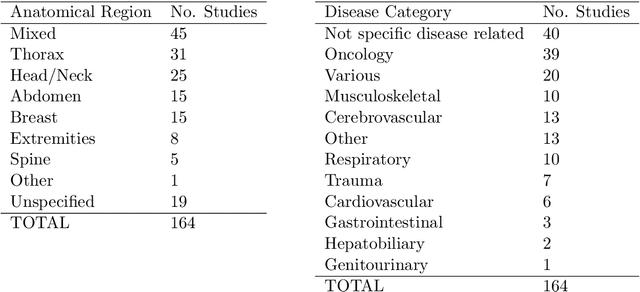
Abstract:NLP has a significant role in advancing healthcare and has been found to be key in extracting structured information from radiology reports. Understanding recent developments in NLP application to radiology is of significance but recent reviews on this are limited. This study systematically assesses recent literature in NLP applied to radiology reports. Our automated literature search yields 4,799 results using automated filtering, metadata enriching steps and citation search combined with manual review. Our analysis is based on 21 variables including radiology characteristics, NLP methodology, performance, study, and clinical application characteristics. We present a comprehensive analysis of the 164 publications retrieved with each categorised into one of 6 clinical application categories. Deep learning use increases but conventional machine learning approaches are still prevalent. Deep learning remains challenged when data is scarce and there is little evidence of adoption into clinical practice. Despite 17% of studies reporting greater than 0.85 F1 scores, it is hard to comparatively evaluate these approaches given that most of them use different datasets. Only 14 studies made their data and 15 their code available with 10 externally validating results. Automated understanding of clinical narratives of the radiology reports has the potential to enhance the healthcare process but reproducibility and explainability of models are important if the domain is to move applications into clinical use. More could be done to share code enabling validation of methods on different institutional data and to reduce heterogeneity in reporting of study properties allowing inter-study comparisons. Our results have significance for researchers providing a systematic synthesis of existing work to build on, identify gaps, opportunities for collaboration and avoid duplication.
Plague Dot Text: Text mining and annotation of outbreak reports of the Third Plague Pandemic
Feb 04, 2020
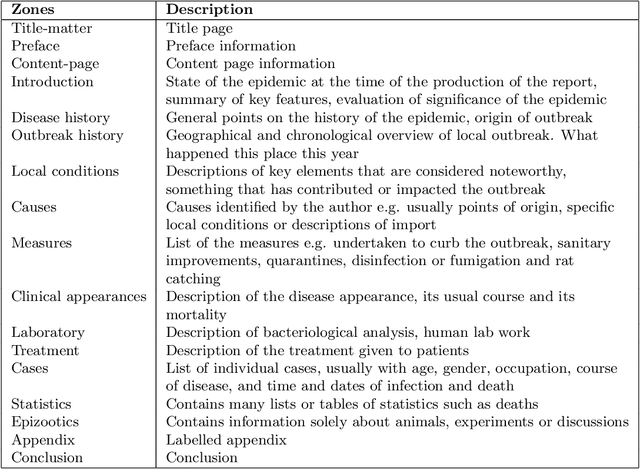
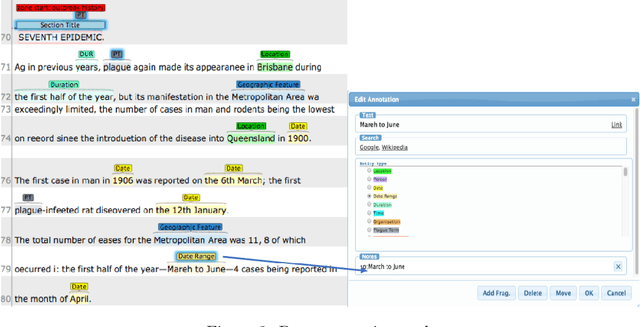
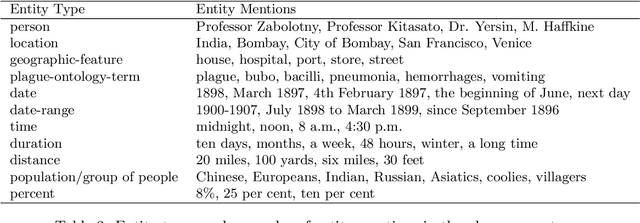
Abstract:The design of models that govern diseases in population is commonly built on information and data gathered from past outbreaks. However, epidemic outbreaks are never captured in statistical data alone but are communicated by narratives, supported by empirical observations. Outbreak reports discuss correlations between populations, locations and the disease to infer insights into causes, vectors and potential interventions. The problem with these narratives is usually the lack of consistent structure or strong conventions, which prohibit their formal analysis in larger corpora. Our interdisciplinary research investigates more than 100 reports from the third plague pandemic (1894-1952) evaluating ways of building a corpus to extract and structure this narrative information through text mining and manual annotation. In this paper we discuss the progress of our ongoing exploratory project, how we enhance optical character recognition (OCR) methods to improve text capture, our approach to structure the narratives and identify relevant entities in the reports. The structured corpus is made available via Solr enabling search and analysis across the whole collection for future research dedicated, for example, to the identification of concepts. We show preliminary visualisations of the characteristics of causation and differences with respect to gender as a result of syntactic-category-dependent corpus statistics. Our goal is to develop structured accounts of some of the most significant concepts that were used to understand the epidemiology of the third plague pandemic around the globe. The corpus enables researchers to analyse the reports collectively allowing for deep insights into the global epidemiological consideration of plague in the early twentieth century.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge