the Alzheimer's Disease Neuroimaging Initiative
EPAD
A Weighted Prognostic Covariate Adjustment Method for Efficient and Powerful Treatment Effect Inferences in Randomized Controlled Trials
Sep 25, 2023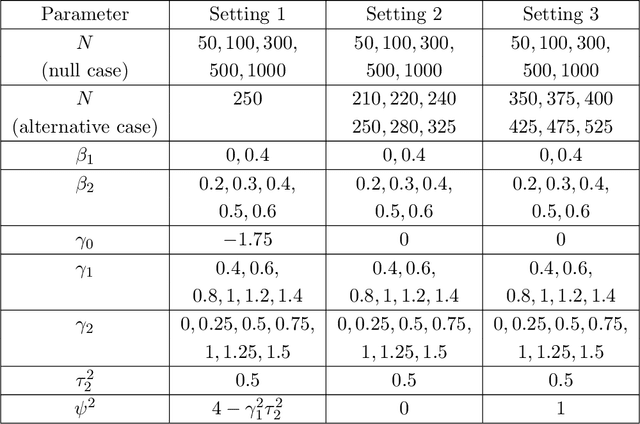
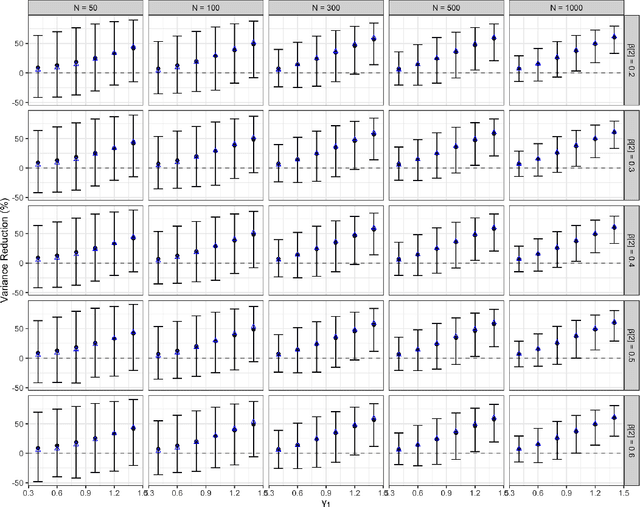
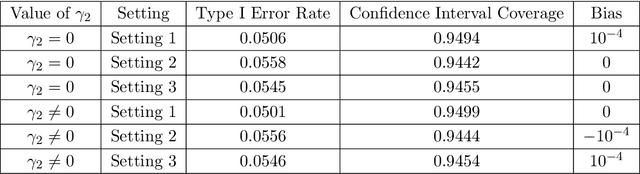

Abstract:A crucial task for a randomized controlled trial (RCT) is to specify a statistical method that can yield an efficient estimator and powerful test for the treatment effect. A novel and effective strategy to obtain efficient and powerful treatment effect inferences is to incorporate predictions from generative artificial intelligence (AI) algorithms into covariate adjustment for the regression analysis of a RCT. Training a generative AI algorithm on historical control data enables one to construct a digital twin generator (DTG) for RCT participants, which utilizes a participant's baseline covariates to generate a probability distribution for their potential control outcome. Summaries of the probability distribution from the DTG are highly predictive of the trial outcome, and adjusting for these features via regression can thus improve the quality of treatment effect inferences, while satisfying regulatory guidelines on statistical analyses, for a RCT. However, a critical assumption in this strategy is homoskedasticity, or constant variance of the outcome conditional on the covariates. In the case of heteroskedasticity, existing covariate adjustment methods yield inefficient estimators and underpowered tests. We propose to address heteroskedasticity via a weighted prognostic covariate adjustment methodology (Weighted PROCOVA) that adjusts for both the mean and variance of the regression model using information obtained from the DTG. We prove that our method yields unbiased treatment effect estimators, and demonstrate via comprehensive simulation studies and case studies from Alzheimer's disease that it can reduce the variance of the treatment effect estimator, maintain the Type I error rate, and increase the power of the test for the treatment effect from 80% to 85%~90% when the variances from the DTG can explain 5%~10% of the variation in the RCT participants' outcomes.
Predicting Time-to-conversion for Dementia of Alzheimer's Type using Multi-modal Deep Survival Analysis
May 02, 2022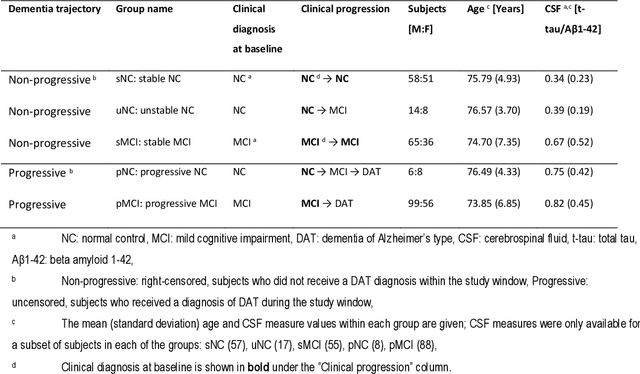
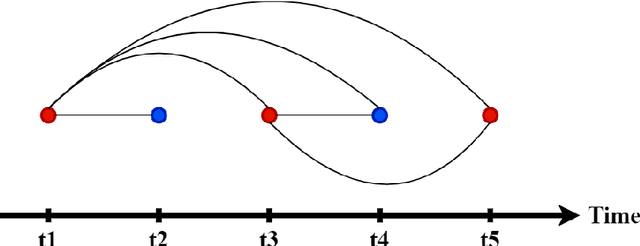

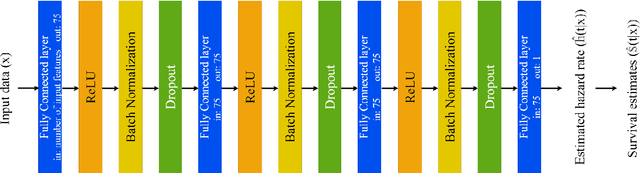
Abstract:Dementia of Alzheimer's Type (DAT) is a complex disorder influenced by numerous factors, but it is unclear how each factor contributes to disease progression. An in-depth examination of these factors may yield an accurate estimate of time-to-conversion to DAT for patients at various disease stages. We used 401 subjects with 63 features from MRI, genetic, and CDC (Cognitive tests, Demographic, and CSF) data modalities in the Alzheimer's Disease Neuroimaging Initiative (ADNI) database. We used a deep learning-based survival analysis model that extends the classic Cox regression model to predict time-to-conversion to DAT. Our findings showed that genetic features contributed the least to survival analysis, while CDC features contributed the most. Combining MRI and genetic features improved survival prediction over using either modality alone, but adding CDC to any combination of features only worked as well as using only CDC features. Consequently, our study demonstrated that using the current clinical procedure, which includes gathering cognitive test results, can outperform survival analysis results produced using costly genetic or CSF data.
Machine Learning Based Multimodal Neuroimaging Genomics Dementia Score for Predicting Future Conversion to Alzheimer's Disease
Mar 11, 2022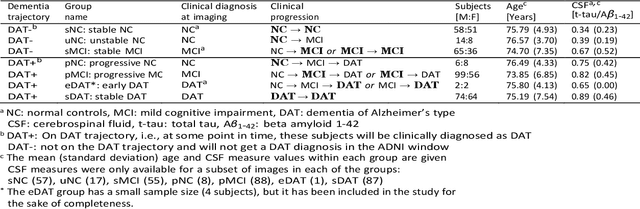
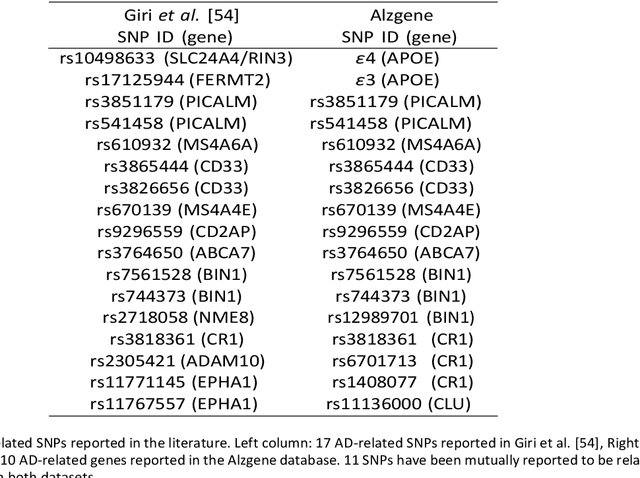
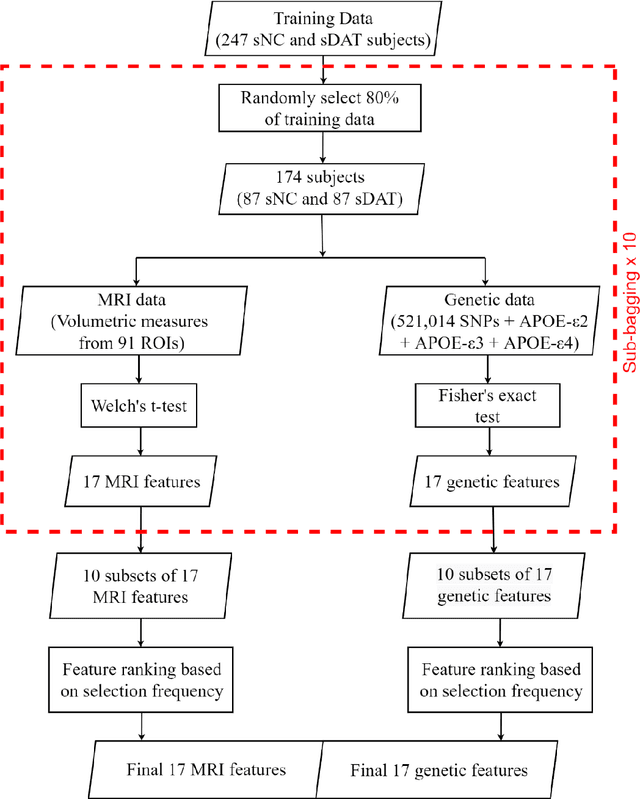

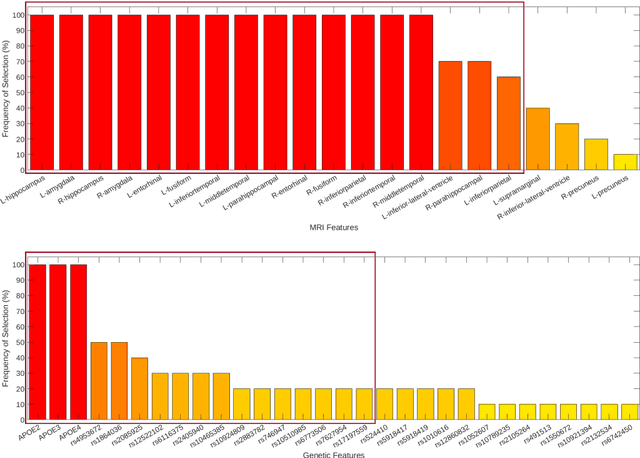
Abstract:Background: The increasing availability of databases containing both magnetic resonance imaging (MRI) and genetic data allows researchers to utilize multimodal data to better understand the characteristics of dementia of Alzheimer's type (DAT). Objective: The goal of this study was to develop and analyze novel biomarkers that can help predict the development and progression of DAT. Methods: We used feature selection and ensemble learning classifier to develop an image/genotype-based DAT score that represents a subject's likelihood of developing DAT in the future. Three feature types were used: MRI only, genetic only, and combined multimodal data. We used a novel data stratification method to better represent different stages of DAT. Using a pre-defined 0.5 threshold on DAT scores, we predicted whether or not a subject would develop DAT in the future. Results: Our results on Alzheimer's Disease Neuroimaging Initiative (ADNI) database showed that dementia scores using genetic data could better predict future DAT progression for currently normal control subjects (Accuracy=0.857) compared to MRI (Accuracy=0.143), while MRI can better characterize subjects with stable mild cognitive impairment (Accuracy=0.614) compared to genetics (Accuracy=0.356). Combining MRI and genetic data showed improved classification performance in the remaining stratified groups. Conclusion: MRI and genetic data can contribute to DAT prediction in different ways. MRI data reflects anatomical changes in the brain, while genetic data can detect the risk of DAT progression prior to the symptomatic onset. Combining information from multimodal data in the right way can improve prediction performance.
Diagnosis of Alzheimer's Disease via Multi-modality 3D Convolutional Neural Network
Feb 26, 2019
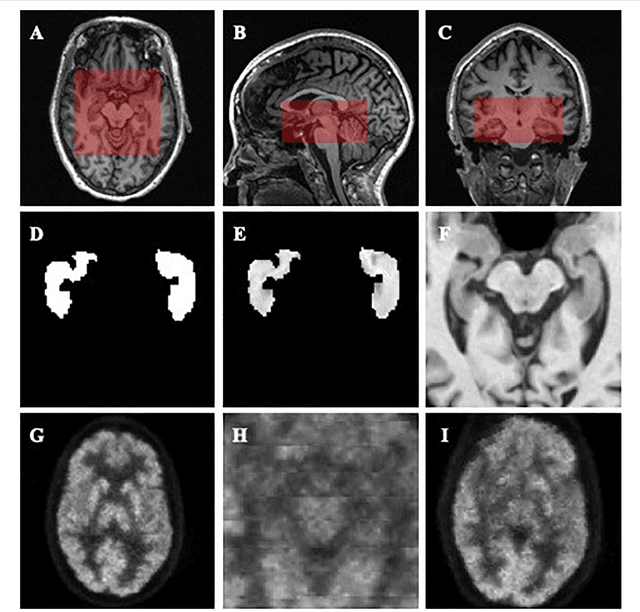

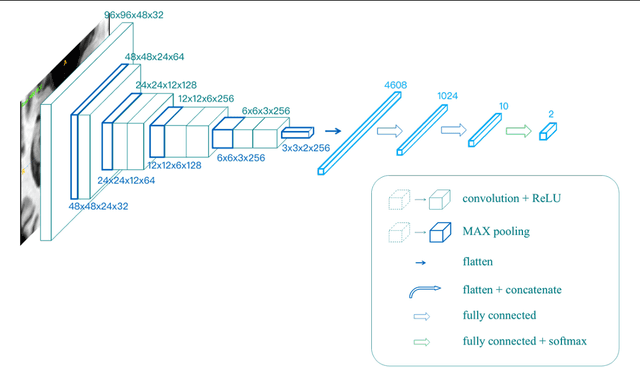
Abstract:Alzheimer's Disease (AD) is one of the most concerned neurodegenerative diseases. In the last decade, studies on AD diagnosis attached great significance to artificial intelligence (AI)-based diagnostic algorithms. Among the diverse modality imaging data, T1-weighted MRI and 18F-FDGPET are widely researched for this task. In this paper, we propose a novel convolutional neural network (CNN) to fuse the multi-modality information including T1-MRI and FDG-PDT images around the hippocampal area for the diagnosis of AD. Different from the traditional machine learning algorithms, this method does not require manually extracted features, and utilizes the stateof-art 3D image-processing CNNs to learn features for the diagnosis and prognosis of AD. To validate the performance of the proposed network, we trained the classifier with paired T1-MRI and FDG-PET images using the ADNI datasets, including 731 Normal (NL) subjects, 647 AD subjects, 441 stable MCI (sMCI) subjects and 326 progressive MCI (pMCI) subjects. We obtained the maximal accuracies of 90.10% for NL/AD task, 87.46% for NL/pMCI task, and 76.90% for sMCI/pMCI task. The proposed framework yields comparative results against state-of-the-art approaches. Moreover, the experimental results have demonstrated that (1) segmentation is not a prerequisite by using CNN, (2) the hippocampal area provides enough information to give a reference to AD diagnosis. Keywords: Alzheimer's Disease, Multi-modality, Image Classification, CNN, Deep Learning, Hippocampal
Neuroimaging Modality Fusion in Alzheimer's Classification Using Convolutional Neural Networks
Nov 13, 2018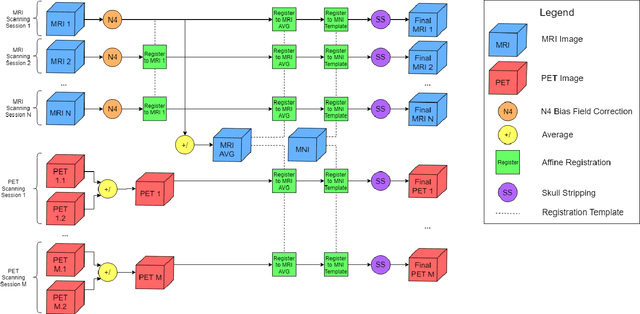
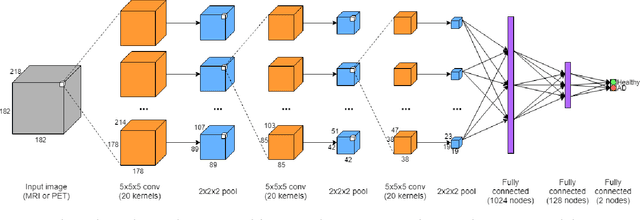
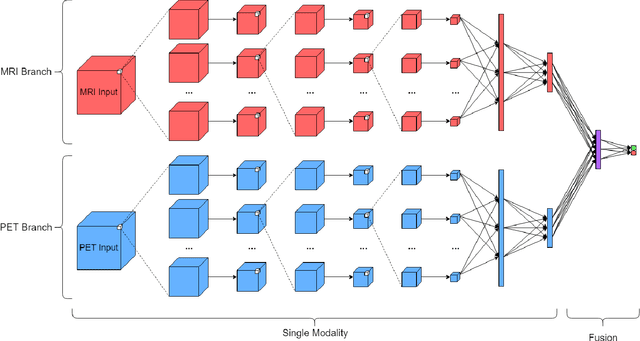
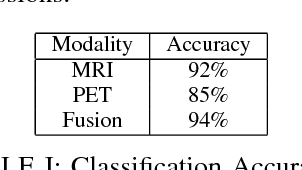
Abstract:Automated methods for Alzheimer's disease (AD) classification have the potential for great clinical benefits and may provide insight for combating the disease. Machine learning, and more specifically deep neural networks, have been shown to have great efficacy in this domain. These algorithms often use neurological imaging data such as MRI and PET, but a comprehensive and balanced comparison of these modalities has not been performed. In order to accurately determine the relative strength of each imaging variant, this work performs a comparison study in the context of Alzheimer's dementia classification using the Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset. Furthermore, this work analyzes the benefits of using both modalities in a fusion setting and discusses how these data types may be leveraged in future AD studies using deep learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge