Zilong Bai
Extracting Post-Acute Sequelae of SARS-CoV-2 Infection Symptoms from Clinical Notes via Hybrid Natural Language Processing
Aug 17, 2025Abstract:Accurately and efficiently diagnosing Post-Acute Sequelae of COVID-19 (PASC) remains challenging due to its myriad symptoms that evolve over long- and variable-time intervals. To address this issue, we developed a hybrid natural language processing pipeline that integrates rule-based named entity recognition with BERT-based assertion detection modules for PASC-symptom extraction and assertion detection from clinical notes. We developed a comprehensive PASC lexicon with clinical specialists. From 11 health systems of the RECOVER initiative network across the U.S., we curated 160 intake progress notes for model development and evaluation, and collected 47,654 progress notes for a population-level prevalence study. We achieved an average F1 score of 0.82 in one-site internal validation and 0.76 in 10-site external validation for assertion detection. Our pipeline processed each note at $2.448\pm 0.812$ seconds on average. Spearman correlation tests showed $\rho >0.83$ for positive mentions and $\rho >0.72$ for negative ones, both with $P <0.0001$. These demonstrate the effectiveness and efficiency of our models and their potential for improving PASC diagnosis.
Streamlining Biomedical Research with Specialized LLMs
Apr 15, 2025Abstract:In this paper, we propose a novel system that integrates state-of-the-art, domain-specific large language models with advanced information retrieval techniques to deliver comprehensive and context-aware responses. Our approach facilitates seamless interaction among diverse components, enabling cross-validation of outputs to produce accurate, high-quality responses enriched with relevant data, images, tables, and other modalities. We demonstrate the system's capability to enhance response precision by leveraging a robust question-answering model, significantly improving the quality of dialogue generation. The system provides an accessible platform for real-time, high-fidelity interactions, allowing users to benefit from efficient human-computer interaction, precise retrieval, and simultaneous access to a wide range of literature and data. This dramatically improves the research efficiency of professionals in the biomedical and pharmaceutical domains and facilitates faster, more informed decision-making throughout the R\&D process. Furthermore, the system proposed in this paper is available at https://synapse-chat.patsnap.com.
PharmaGPT: Domain-Specific Large Language Models for Bio-Pharmaceutical and Chemistry
Jul 03, 2024



Abstract:Large language models (LLMs) have revolutionized Natural Language Processing (NLP) by by minimizing the need for complex feature engineering. However, the application of LLMs in specialized domains like biopharmaceuticals and chemistry remains largely unexplored. These fields are characterized by intricate terminologies, specialized knowledge, and a high demand for precision areas where general purpose LLMs often fall short. In this study, we introduce PharmGPT, a suite of multilingual LLMs with 13 billion and 70 billion parameters, specifically trained on a comprehensive corpus of hundreds of billions of tokens tailored to the Bio-Pharmaceutical and Chemical sectors. Our evaluation shows that PharmGPT matches or surpasses existing general models on key benchmarks, such as NAPLEX, demonstrating its exceptional capability in domain-specific tasks. This advancement establishes a new benchmark for LLMs in the Bio-Pharmaceutical and Chemical fields, addressing the existing gap in specialized language modeling. Furthermore, this suggests a promising path for enhanced research and development in these specialized areas, paving the way for more precise and effective applications of NLP in specialized domains.
PharmGPT: Domain-Specific Large Language Models for Bio-Pharmaceutical and Chemistry
Jun 26, 2024



Abstract:Large language models (LLMs) have revolutionized Natural Language Processing (NLP) by by minimizing the need for complex feature engineering. However, the application of LLMs in specialized domains like biopharmaceuticals and chemistry remains largely unexplored. These fields are characterized by intricate terminologies, specialized knowledge, and a high demand for precision areas where general purpose LLMs often fall short. In this study, we introduce PharmGPT, a suite of multilingual LLMs with 13 billion and 70 billion parameters, specifically trained on a comprehensive corpus of hundreds of billions of tokens tailored to the Bio-Pharmaceutical and Chemical sectors. Our evaluation shows that PharmGPT matches or surpasses existing general models on key benchmarks, such as NAPLEX, demonstrating its exceptional capability in domain-specific tasks. This advancement establishes a new benchmark for LLMs in the Bio-Pharmaceutical and Chemical fields, addressing the existing gap in specialized language modeling. Furthermore, this suggests a promising path for enhanced research and development in these specialized areas, paving the way for more precise and effective applications of NLP in specialized domains.
PatentGPT: A Large Language Model for Intellectual Property
Apr 30, 2024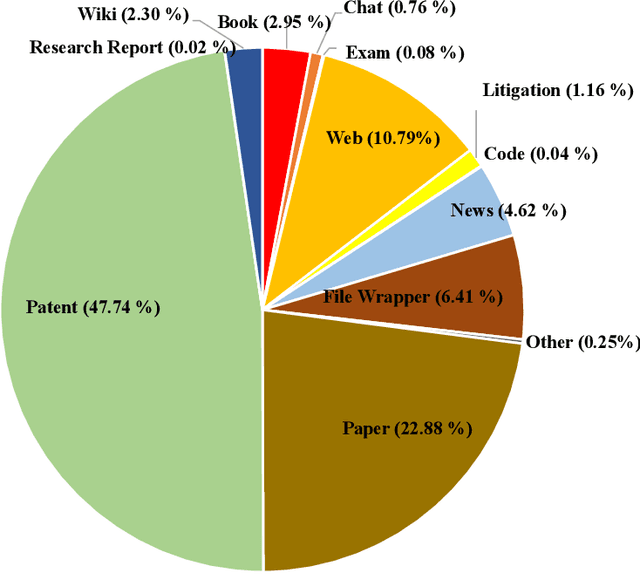

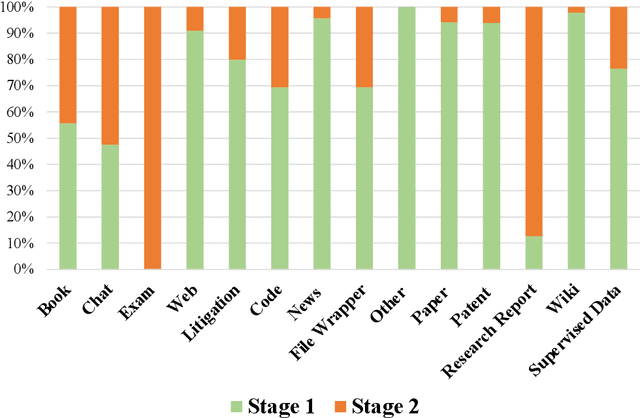
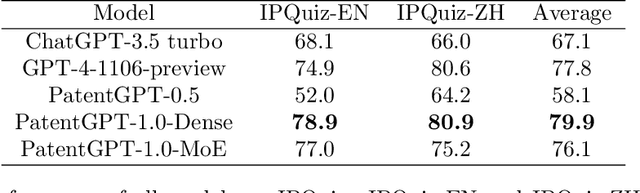
Abstract:In recent years, large language models have attracted significant attention due to their exceptional performance across a multitude of natural language process tasks, and have been widely applied in various fields. However, the application of large language models in the Intellectual Property (IP) space is challenging due to the strong need for specialized knowledge, privacy protection, processing of extremely long text in this field. In this technical report, we present for the first time a low-cost, standardized procedure for training IP-oriented LLMs, meeting the unique requirements of the IP domain. Using this standard process, we have trained the PatentGPT series models based on open-source pretrained models. By evaluating them on the open-source IP-oriented benchmark MOZIP, our domain-specific LLMs outperforms GPT-4, indicating the effectiveness of the proposed training procedure and the expertise of the PatentGPT models in the IP demain. What is impressive is that our model significantly outperformed GPT-4 on the 2019 China Patent Agent Qualification Examination by achieving a score of 65, reaching the level of human experts. Additionally, the PatentGPT model, which utilizes the SMoE architecture, achieves performance comparable to that of GPT-4 in the IP domain and demonstrates a better cost-performance ratio on long-text tasks, potentially serving as an alternative to GPT-4 within the IP domain.
Application of Deep Learning on Single-Cell RNA-sequencing Data Analysis: A Review
Oct 11, 2022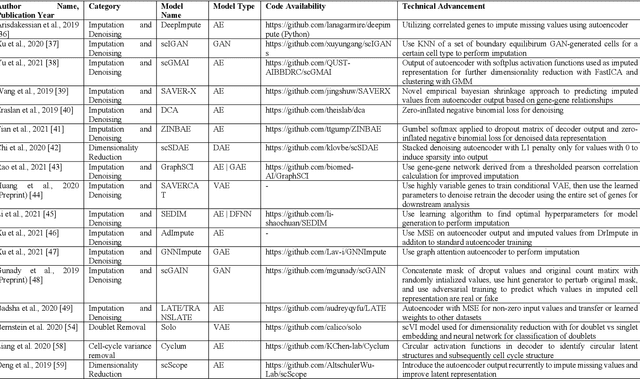

Abstract:Single-cell RNA-sequencing (scRNA-seq) has become a routinely used technique to quantify the gene expression profile of thousands of single cells simultaneously. Analysis of scRNA-seq data plays an important role in the study of cell states and phenotypes, and has helped elucidate biological processes, such as those occurring during development of complex organisms and improved our understanding of disease states, such as cancer, diabetes, and COVID, among others. Deep learning, a recent advance of artificial intelligence that has been used to address many problems involving large datasets, has also emerged as a promising tool for scRNA-seq data analysis, as it has a capacity to extract informative, compact features from noisy, heterogeneous, and high-dimensional scRNA-seq data to improve downstream analysis. The present review aims at surveying recently developed deep learning techniques in scRNA-seq data analysis, identifying key steps within the scRNA-seq data analysis pipeline that have been advanced by deep learning, and explaining the benefits of deep learning over more conventional analysis tools. Finally, we summarize the challenges in current deep learning approaches faced within scRNA-seq data and discuss potential directions for improvements in deep algorithms for scRNA-seq data analysis.
Block Model Guided Unsupervised Feature Selection
Jul 05, 2020
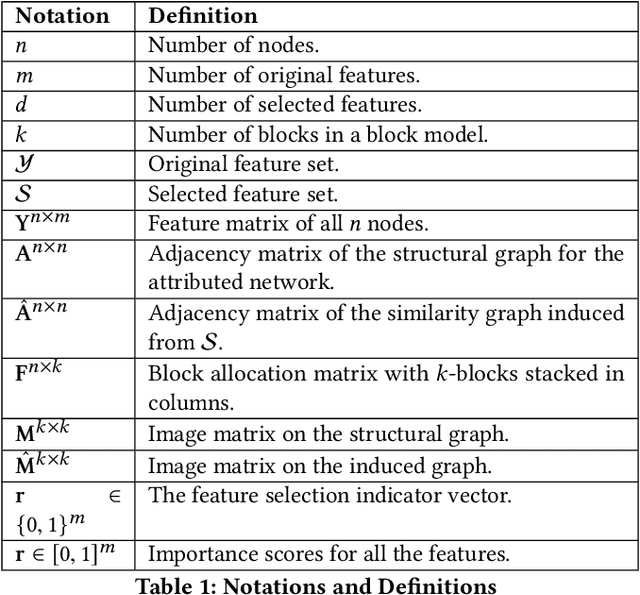
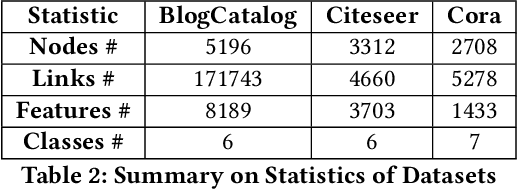

Abstract:Feature selection is a core area of data mining with a recent innovation of graph-driven unsupervised feature selection for linked data. In this setting we have a dataset $\mathbf{Y}$ consisting of $n$ instances each with $m$ features and a corresponding $n$ node graph (whose adjacency matrix is $\mathbf{A}$) with an edge indicating that the two instances are similar. Existing efforts for unsupervised feature selection on attributed networks have explored either directly regenerating the links by solving for $f$ such that $f(\mathbf{y}_i,\mathbf{y}_j) \approx \mathbf{A}_{i,j}$ or finding community structure in $\mathbf{A}$ and using the features in $\mathbf{Y}$ to predict these communities. However, graph-driven unsupervised feature selection remains an understudied area with respect to exploring more complex guidance. Here we take the novel approach of first building a block model on the graph and then using the block model for feature selection. That is, we discover $\mathbf{F}\mathbf{M}\mathbf{F}^T \approx \mathbf{A}$ and then find a subset of features $\mathcal{S}$ that induces another graph to preserve both $\mathbf{F}$ and $\mathbf{M}$. We call our approach Block Model Guided Unsupervised Feature Selection (BMGUFS). Experimental results show that our method outperforms the state of the art on several real-world public datasets in finding high-quality features for clustering.
* Published at KDD2020
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge